Abstracts
Bacillus species constitute a diverse group of bacteria widely distributed in soil and the aquatic environment. In this study, Bacillus strains isolated from the coastal environment of Cochin, India were identified by detailed conventional biochemical methods, fatty acid methyl ester (FAME) analysis and partial 16S rDNA sequencing. Analysis of the data revealed that Bacillus pumilus was the most predominant species in the region under study followed by B. cereus and B. sphaericus. The B. pumilus isolates were further characterized by arbitrarily primed PCR (AP-PCR), antibiotic sensitivity profiling and PCR screening for known toxin genes associated with Bacillus spp. All B. pumilus isolates were biochemically identical, exhibited high protease and lipase activity and uniformly sensitive to antibiotics tested in this study. One strain of B. pumilus harboured cereulide synthetase gene cesB of B. cereus which was indistinguishable from rest of the isolates biochemically and by AP-PCR. This study reports, for the first time, the presence of the emetic toxin gene cesB in B. pumilus.
Bacillus pumilus; FAME; 16S rDNA; cesB; AP-PCR
As espécies de Bacillus constituem um grupo diversificado de bactérias amplamente distribuídas no solo e no ambiente aquático. Neste estudo, cepas de Bacillus isoladas do ambiente costeiro de Cochin, Índia, foram identificadas através de métodos bioquímicos convencionais, análise de ésteres metílicos de ácidos graxos (FAME) e sequenciamento de 16S rDNA. A análise dos dados revelou que Bacillus pumilus foi a espécie predominante na região estudada, seguido de B. cereus e B. sphaericus. Os isolados de B. pumilus foram caracterizados através da reação em cadeia da polimerase com primers arbitrários (AP-PCR), perfil de sensibilidade a antibióticos e triagem por PCR de genes de toxinas associadas com Bacillus spp. Todos os isolados de B. pumilus foram bioquimicamente idênticos, apresentaram elevada atividade de protease e lipase e foram uniformemente sensíveis aos antibióticos estudados. Um dos isolados de B. pumilus apresentou o gene cesB de B. cereus, que não foinão distinguível dos demais isolados por testes bioquímicos nem por AP-PCR. Este é o primeiro relato da presença do gene cesB da toxina eméticaem B. pumilus.
Aspergillus flavus; pimenta em pó; alterações nutricionais; aflatoxina
ENVIRONMETAL MICROBIOLOGY
Biochemical and molecular characterization of Bacillus pumilus isolated from coastal environment in Cochin, India
Caracterização bioquímica e molecular de Bacillus pumilus isolado do ambiente costeiro de Cochin, Índia
Ammini ParvathiI; Kiran KrishnaI; Jiya JoseI; Neetha JosephI; Santha NairI,II
IDepartment of Biological Oceanography, National Institute of Oceanography, Regional Center, Cochin - 682 018, India
IINational Institute of Oceanography, Dona Paula, Goa - 403 004, India
Corresponding Author Corresponding Author Ammini Parvathi Department of Biological Oceanography, National Institute of Oceanography, Regional Centre Cochin 682 018, India Tel.: 91-484-2390814; Fax: 91-484-2390618 E-mail: parubfsc@yahoo.co.in
ABSTRACT
Bacillus species constitute a diverse group of bacteria widely distributed in soil and the aquatic environment. In this study, Bacillus strains isolated from the coastal environment of Cochin, India were identified by detailed conventional biochemical methods, fatty acid methyl ester (FAME) analysis and partial 16S rDNA sequencing. Analysis of the data revealed that Bacillus pumilus was the most predominant species in the region under study followed by B. cereus and B. sphaericus. The B. pumilus isolates were further characterized by arbitrarily primed PCR (AP-PCR), antibiotic sensitivity profiling and PCR screening for known toxin genes associated with Bacillus spp. All B. pumilus isolates were biochemically identical, exhibited high protease and lipase activity and uniformly sensitive to antibiotics tested in this study. One strain of B. pumilus harboured cereulide synthetase gene cesB of B. cereus which was indistinguishable from rest of the isolates biochemically and by AP-PCR. This study reports, for the first time, the presence of the emetic toxin gene cesB in B. pumilus.
Key-words:Bacillus pumilus; FAME; 16S rDNA; cesB; AP-PCR.
RESUMO
As espécies de Bacillus constituem um grupo diversificado de bactérias amplamente distribuídas no solo e no ambiente aquático. Neste estudo, cepas de Bacillus isoladas do ambiente costeiro de Cochin, Índia, foram identificadas através de métodos bioquímicos convencionais, análise de ésteres metílicos de ácidos graxos (FAME) e sequenciamento de 16S rDNA. A análise dos dados revelou que Bacillus pumilus foi a espécie predominante na região estudada, seguido de B. cereus e B. sphaericus. Os isolados de B. pumilus foram caracterizados através da reação em cadeia da polimerase com primers arbitrários (AP-PCR), perfil de sensibilidade a antibióticos e triagem por PCR de genes de toxinas associadas com Bacillus spp. Todos os isolados de B. pumilus foram bioquimicamente idênticos, apresentaram elevada atividade de protease e lipase e foram uniformemente sensíveis aos antibióticos estudados. Um dos isolados de B. pumilus apresentou o gene cesB de B. cereus, que não foinão distinguível dos demais isolados por testes bioquímicos nem por AP-PCR. Este é o primeiro relato da presença do gene cesB da toxina eméticaem B. pumilus.
Palavras-chave:Aspergillus flavus, pimenta em pó, alterações nutricionais, aflatoxina.
INTRODUCTION
The Gram-positive, aerobic, rod-shaped endospore-forming bacteria of the Genus Bacillus are the most widely represented organisms in the soil. Due to their ability to form spores and withstand a range of variable environmental conditions, Bacillus spp. adapt easily to diverse habitats (25). The diverse physiology of Bacillus spp. requires elaborate biochemical tests for their identification (30). Advances in chromatographic analysis of whole cell fatty acid methyl ester (FAME) profiles have made this technique sufficiently sensitive and reliable for grouping of Bacillus at species level (31). Further, nucleic acid based techniques such as 16S rDNA (3,34) and gyrase B (gyrB) sequence analysis have proved to be of immense value for phylogenetic analysis of bacteria (35). Based on the 16S rDNA sequence analysis, 5 groups have been identified within the genus Bacillus, of which the group 1 (B. subitilis group) comprises of B. amyloliquefaciens, B. subtilis and B. pumilus (3,12).
Several species of Bacillus inhabit coastal and marine environments, though it is hard to strictly classify them as indigenous to these habitats. Together with B. cereus and B. subtilis, B. pumilus is considered as a major component of marine bacterial communities (8,15,22,27). Recently, B. pumilus has also been reported to be the second most predominant Bacillus species in spacecrafts (17). This bacterium is highly resistant to extreme environmental conditions such as low or no nutrient availability, desiccation, irradiation, H2O2 and chemical disinfections (19). The ecological role of B. pumilus is emphasized by the fact that they do produce compounds antagonist to fungal and bacterial pathogens (4,6). Thus, B. pumilus is of considerable research interest to understand its physiological diversity, genetic relatedness with other Bacillus spp. and the possible presence of toxigenic factors. In the study reported here, we describe i) isolation and identification of Bacillus spp. from environmental samples by conventional methods, FAME and 16S rDNA sequencing and ii) further phenotypic and genetic characterization of B. pumilus, the predominant Bacillus group of bacteria in the coastal region under study.
MATERIALS AND METHODS
Isolation and biochemical characterization of Bacillus spp.
Sea water, sediment, fish and shellfish were collected off Cochin, West coast of India and processed for the isolation of Bacillus spp. Fish, shellfish or sediment samples were homogenized in phosphate buffered saline (PBS 0.05 M, pH 7.2), serially diluted in the same medium and spread plated on nutrient agar prepared in 50% seawater. One hundred micro liters of seawater samples were directly spread plated on the same medium and incubated at 30ºC for 24-48 h. The colonies that came up on agar plates were purified and stored at -80ºC in nutrient broth containing 30% glycerol. For taxonomic identification, the isolates were subjected to a series of biochemical tests (11), which included nitrate reduction, anaerobic growth, gas production from glucose, Voges-Proskauer (VP), growth at different NaCl concentrations, temperature and pH ranges, degradation of starch, casein, urea, tween 20, gelatin, chitin, acid production from arabinose, mannitol, xylose, glucose, lactose, citrate utilization and production of DNAse. The production of extracellular enzymes namely caseinase, chitinase, protease, alkaline phosphatase, gelatinase and lipase was studied following the protocol described by Smibert and Krieg (29)
Fatty acid methyl ester (FAME) analysis
Gas chromatographic analysis of whole cell fatty acid methyl ester (FAME) was performed for further identification and grouping of isolates. Fatty acid methyl ester extraction was performed using standard procedures (28). The fatty acid profiles generated were compared against an inbuilt Sherlock TSBA Library version 3.9 (MIDI Inc., DE, USA). A similarity index of >60% was used for clustering of isolates at species level.
Antimicrobial susceptibility assay
The inhibition of B. pumilus strains by various antibiotics was tested by standard disc diffusion technique (7). The cultures were grown in nutrient broth overnight and plated on Muller Hinton agar (Hi-Media, Mumbai). The following antibiotic discs with their concentrations indicated in parenthesis were used; amoxicillin (25 mcg), penicillin (10 mcg), ciprofloxacin (5 mcg), gentamycin (10 mcg), cotrimaxazole (25 mcg), chloramphenicol (30 mcg), bacitracin (8 mcg), tetracycline (30 mcg), kanamycin (30 mcg), erythromycin (15 mcg), vancomycin (30 mcg).
DNA isolation and purification
Pure genomic DNA was isolated following the method of Ausubel et al. (5). Briefly, the cultures were grown overnight in 3 ml nutrient broth with shaking at 30ºC. A 1.5 ml of the culture was centrifuged at 12 000 g for 10 min and the resultant pellet was resuspended in 567 µl 1× TE buffer (10 mM Tris pH 8.0, 1 mM EDTA). Proteinase K and SDS were added to final concentrations of 100 µg/ml and 0.5% respectively, and incubated at 37ºC for 1 h. After incubation, NaCl (5 M) and CTAB/NaCl (10% w/v cetyl trimethyl ammonium bromide in 0.7 M NaCl) were added and incubated at 65ºC for 10 min. The mixture was extracted once each with an equal volume of chloroform-isoamyl alcohol (24:1) and phenol-chloroform-isoamyl alcohol (25:24:1). DNA was precipitated from the aqueous phase using 0.6 volumes of isopropanol and washed once with 70% ethanol. The DNA pellet obtained after final centrifugation was vacuum dried and dissolved in 50 µl 1× TE buffer. DNA quantification was done using a UV-1601 spectrophotometer (Shimadzu Corporation, Japan).
16S rDNA sequencing and AP-PCR
The 16S rDNA of 4 strains (NIOB 005, NIOB 133, NIOB 485 and NIOB 525) were PCR amplified using universal primers and PCR conditions described by Iwamoto et al. (16) (Table 1). The resultant 454 bp products were purified using a PCR purification kit (Qiagen, Germany) and sequenced. The sequences were subjected to homology search using BLAST programme (2) of the National Center for Biotechnology Information (NCBI).
AP-PCR was performed using primer CRA22 described by Neilan (18) (Table 1). All the reactions were carried out in 30 µl volumes consisting of a 10×buffer (100 mM Tris-HCl, 500 mM KCl and 20 mM MgCl2), 200 µM concentrations of each of the four dNTPs, 30 picomoles of primer, 3 U of Taq polymerase (MBI Fermentas). All PCR amplifications were carried out in an eppendorf mastercycler (Eppendorf, Germany). In all the reactions, 300 ng of the pure genomic DNA was used. The amplification products were separated on a 2% agarose gel, stained with ethidium bromide and photographed. Amplification profiles obtained were analyzed and a dendrogram was generated using BioNumerics version 4.6 software (Applied Maths, Belgium)
PCR detection of cereulide synthetase genes
B. pumilus were screened for the presence of cereulide synthetase genes cesA and cesB using primers previously described (Table 1). The expected amplicons with cesA and cesB gene-specific primers were 188 bp and 635 bp respectively.
Nucleotide sequence accession numbers
The partial 16S rDNA and cesB sequences derived in this study have been deposited in GenBank under the accession numbers EU283326, EU283325, EU283323, EU283322, EU283320, EU283318, EU167933, EU283321, EU167932, EU167924, EU289221.
RESULTS
Taxonomic identification of Bacillus spp. isolated from coastal environment of Cochin
Eighty-two Bacillus spp. were isolated and identified by biochemical tests and fatty acid methyl ester analysis (FAME). These included B. pumilus (16), B. cereus (15), B. sphaericus (11), B. subtilis (10), B. amyloliquefaciens (8), B. megaterium (6), B. lentimorbus (5), B. coagulans (4), B. licheniformis (4), B. circulans (1), B. flexus (1) and Bacillus GC group 22 (1). The Bacillus GC group 22 corresponds to the gas chromatographic profile of a Bacillus species in the Sherlock TSBA Library version 3.9 (Microbial ID, MIDI Inc.), the 16S rDNA sequence of which does not match any known species of the genus Bacillus. In our study, B. pumilus was the most predominant species followed by B. cereus and B. sphaericus. Ivanova et al. (15) in their study found that B. pumilus and B. subtilis were the most abundant Bacillus spp. associated with marine sponges, ascidians, soft corals, and seawater.
Characterization of B. pumilus
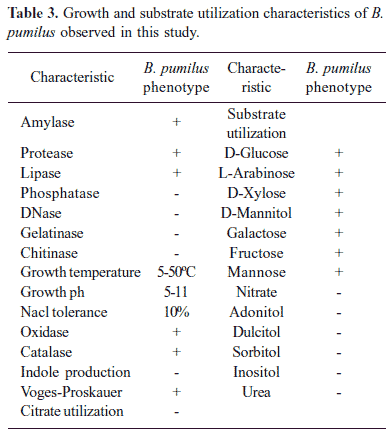
Sixteen B. pumilus strains isolated from different sources (Table 2) exhibited uniform phenotypic properties (Table 3). Physiological tests revealed the production of detectable protease and lipase but not amylase, phosphatase, DNase, gelatinase and chitinase. The partial 16S rDNA sequences (≈ 500 bp) of B. pumilus determined in this study revealed 99-100% homology with B. pumilus 16S rDNA sequences in the GenBank. The antibiotic susceptibility profiles of B. pumilus were identical. All 16 isolates were uniformly inhibited by amoxicillin, ciprofloxacin, gentamycin, cotrimaxazole, chloramphenicol, bacitracin, tetracycline, kanamycin, erythromycin, vancomycin. All isolates exhibited resistance to penicillin.
AP-PCR typing of isolates
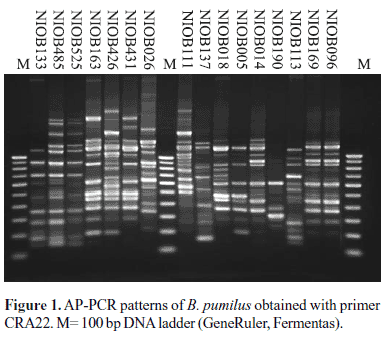
The random primer CRA 22, which consistently yielded 4-10 bands with B. pumilus, was chosen for typing of isolates by AP-PCR. The analysis of AP-PCR fingerprints revealed heterogeneity among B. pumilus isolates with 10 distinct patterns (Fig. 1). Despite this overall genetic diversity, near identical profiles were obtained between strains NIOB 485 and NIOB 525 (from crab), strains NIOB111 (fish) and NIOB 426 (from sediment), strains NIOB096 (from crab) and NIOB169 (from fish), strains NIOB163 (from fish) and NIOB431 (from starfish).
Detection of cereulide synthetase gene in B. pumilus by PCR
One isolate B. pumilus NIOB 133 isolated from an estuarine fish yielded 635 bp amplicon with cesB-targeted primers EM1-f and EM1-r (Fig. 2). The product of cesB PCR was sequenced and nucleotide sequence analysis of the PCR product revealed 96% similarity with corresponding sequence of cesB of B. cereus (GenBank accession no. DQ889676) (26), while the deduced amino acid sequence showed 92% homology with a few amino acid mismatches (Fig. 3). This strain was negative by cesA PCR using primers CER1 and EMT1.
DISCUSSION
This study on the diversity of Bacillus spp. isolated from a coastal environment by biochemical assays, FAME analysis and 16S rDNA sequencing revealed that B. pumilus was the predominant species followed by B. cereus. B. pumilus belongs to B. subtilis group of aerobic spore-forming organisms, which has lately evoked considerable research interest due its involvement in cases of food-poisoning. Recently, a pumilacidin-producing B. pumilus has been implicated in a case of food poisoning (13). A study by Brophy and Knoop (9) reported experimental induction of enterocolitis in guinea pigs, while some compounds produced by B. pumilus were reportedly toxic to mice, eukaryotic cells and humans (20,21,24,32).
Sixteen environmental B. pumilus strains isolated from different sources (Table 2) exhibited uniform phenotypic properties (Table 3). The identity of these isolates was further confirmed by fatty acid methyl ester analysis with a similarity index of >60% (data not shown). The partial 16S rDNA sequences (500 bp) of these strains revealed 99-100% similarity with B. pumilus 16S rDNA sequences in the GenBank. Thus, a combination of conventional physiological tests and genetic analysis enabled unambiguous identification of B. pumilus from coastal environments of Cochin. None of the strains exhibited resistance to the antibiotics tested in this study except to penicillin. This observation is interesting, since penicillin resistance has not been reported in B. pumilus. Studies on antibiotic resistance of B. pumilus are limited, since the organism is not considered infectious to humans and animals. However, some recent studies have revealed that several Bacillus species including B. pumilus can cause infections, ranging from skin infection to life threatening bacteremia in immunocompromised individuals (23,33). Thus, more studies need to be performed to understand the human health significance of B. pumilus, genetic basis of infections and resistance to antimicrobials.
The whole genome comparison of B. pumilus strains by AP-PCR demonstrated that B. pumilus were genetically diverse. Though the isolates shared several common amplification bands, overall heterogeneity among B. pumilus studied was apparent (Fig. 1). Despite this, some strains isolated from similar sources during different points time exhibited identical or near identical profiles. However, it was not possible to attribute isolates to a particular source solely based on the AP-PCR profiles, since some strains isolated from different sample types such as fish and sediment also exhibited identical AP-PCR profiles. The random primer CRA 22 used in this study has sufficient discriminating power and will be useful for studying genetic diversity among the Bacillus group of bacteria.
In order to understand the toxigenic potential of environmental strains of B. pumilus, the isolates obtained in this study were screened for the presence of cereulide synthetase genes cesA and cesB by PCR. Surprisingly, one strain (NIOB 133) was PCR positive for cesB-targeted gene (Fig. 2). This strain however, did not harbour cesA gene as revealed by cesA-specific PCR using primers CER1 and EMT1. Cereulide is a small heat stable cyclic dodecadepsipeptide produced by some strains of B. cereus which has high toxicity to humans (1,36). Cereulide is synthesized by a non-ribosomal peptide synthetase, encoded by the ces genes located on a 270-kb pXOI-like virulence plasmid named pCER270 (10,26), more often found associated with clinical isolates of B. cereus. A recent study has reported the presence of cesA, but not cesB, in B. pumilus and B. licheniformis isolated from bovine mastitis (20). However, the cesB-positive strain in our study lacked cesA. Therefore, it appears that cereulide-encoding genes may be distributed over a wide species range within the Bacillus group of bacteria in the environment. The cesB-positive B. pumilus in the present study was indistinguishable from the rest of the strains by biochemical assays or by its fatty acid profile. Further, the AP-PCR profile of this strain was similar to other non-toxigenic strains (Fig. 1). The production of toxin and the consequent ability of this strain to initiate emetic symptoms need to be established by suitable animal feeding experiments. Our study constitutes the first report on the presence of cesB in B. pumilus.
In conclusion, Bacillus spp. constitute key components of coastal-marine heterotrophic bacterial communities owing to their diverse and flexible physiological properties. Isolation and characterization of Bacillus spp. from these environments will help in identifying novel mechanisms of environmental survival, diverse metabolic activities, production of biotechnologically valuable compounds such as enzymes and antimicrobial substances and the presence of putative toxigenic factors. In our study, a combination of methods involving FAME, 16S rDNA sequencing and biochemical assays enabled complete identification of B. pumilus. Though, the biochemical and enzymatic properties of B. pumilus isolated in this study were uniform, intraspecific genetic diversity was evident from AP-PCR analysis. The detection of cereulide synthetase gene cesB in B. pumilus is significant, since ces genes were previously thought to be restricted to emetic strains of B. cereus. The organization of ces operon in B. pumilus is a subject for further study.
ACKNOWLEDGEMENTS
The authors are grateful to the Director, NIO, Goa and Scientist-In-Charge, NIO, RC, Cochin, for their encouragement and support. Partial financial support from the projects GAP1035 and SIP 1302 is acknowledged. This is NIO contribution number 4513.
Submitted: April 4, 2008; Returned to authors for corrections: July 10, 2008; Approved: February 26, 2009
- 1. Agata, N.; Ohta, M.; Mori, M.; Isobe, M. (1995). A novel dodecadepsipeptide, cereulide, is an emetic toxin of Bacillus cereus FEMS. Microbiol. Lett ., 129 (1), 17-20.
- 2. Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. (1990). Basic local alignment search tool. J. Mol. Biol., 215 (3), 403-410.
- 3. Ash, C.; Farrow, J.A.; Dorsch, M.; Stackebrandt, E.; Collins, M.D. (1991). Comparative analysis of Bacillus anthracis, Bacillus cereus, and related species on the basis of reverse transcriptase sequencing of 16S rRNA. Int. J. Syst. Bacteriol., 41 (3), 343-346.
- 4. Aunpad, R.; Na-Bangchang, K. (2007). Pumilicin 4, A novel bacteriocin with anti-MRSA and anti-VRE activity produced by newly isolated bacteria Bacillus pumilus strain WAPB4. Curr. Microbiol, 55 (4), 308-313.
- 5. Ausubel, F.M.; Brent, R.; Kingsten, R.E.; Moore, D.D.; Seidman, J.G.; Smith, J.A.; Struhl, K. (1995). Short protocols in molecular biology John Wiley and Sons, New York.
- 6. Banerjee, S.; Devaraja, T.N.; Shariff, M.; Yusoff, F.M. (2007). Comparison of four antibiotics with indigenous marine Bacillus spp. in controlling pathogenic bacteria from shrimp and artemia. J. Fish. Dis., 30 (7), 383-389.
- 7. Bauer, A.W.; Kirby, W.M.M.; Skerris, J.C.; Turck, M. (1966). Antibiotic susceptibility testing by a standardized single diffusion method. Am. J. Clin. Pathol., 45 (4), 493-496.
- 8. Boeye, H.; Aerts, M. (1976). Numerical taxonomy of Bacillus isolates from North Sea sediments. Int. J. Syst. Bacteriol., 26, 427-441.
- 9. Brophy, P.F.; Knoop, F.C. (1982). Bacilluspumilus in the induction of clindamycin-associated enterocolitis in guinea pigs. Infect. Immun., 35 (1), 289-295.
- 10. Ehling-Schulz, M.; Fricker, M.; Scherer, S. (2004). Identification of emetic toxin-producing Bacillus cereus strains by a novel molecular assay. FEMS Microbiol. Lett., 232 (2), 189-195
- 11. Farrow, J.A.; Wallbanks, S.; Collins, M.D. (1994). Phylogenetic interrelationship of round-spore-forming bacilli containing cell walls based on lysine and the non-spore-forming genera Caryophanon, Exiguobacterium, Kurthia, and Planococcus Int. J. Syst. Bacteriol., 44 (1), 74-82.
- 12. Fritze, D. (2002). Bacillus identification- traditional approaches. In: Berkeley, R.; Heyndrickx, M.; Logan, N.; De Vos, P. (eds.). Applications and systematics of Bacillus and relatives. Blackwell Science Ltd.
- 13. From, C.; Hormazabal, V.; Granum, P.E. (2007). Food poisoning associated with pumilacidin-producing Bacillus pumilus in rice. Int. J. Food Microbiol., 115 (3), 319-324.
- 14. Horwood, P.F.; Burgess, G.W.; Oakey, H.J. (2004). Evidence for nonribosomal peptide synthetase production of cereulide (the emetic toxin) in Bacillus cereus FEMSMicrobiol. Lett., 236 (2), 319-324.
- 15. Ivanova, E.P.; Vysotskii, M.V.; Svetashev, V.I.; Nedashkovskaya, O.I.; Gorshkova, N.M.; Mikhailov, V.V.; Yumoto, N.; Shigeri, Y.; Taguchi, T.; Yoshikawa, S. (1999). Characterisation of Bacillus strains of marine origin. Int. Microbiol., 2 (4), 267-271.
- 16. Iwamoto, T.; Tani, K.; Nakamura, K.; Suzuki, Y.; Kitagawa, M.; Eguchi, M.; Nasu, M. (2000). Monitoring impact of in situ biostimulation treatment on groundwater bacterial community by DGGE. FEMS Microbiol. Ecol., 32 (2), 129-141.
- 17. La Duc, M.T.; Kern, R.; Venkateswaran, K. (2004). Microbial monitoring of spacecraft and associated environments. Microb. Ecol., 47 (2), 150-158.
- 18. Neilan, B.A. (1995). Identification and phylogenetic analysis of toxigenic cyanobacteria by multiplex randomly amplified polymorphic DNA PCR. Appl. Environ. Microbiol., 61 (6), 2286-2291.
- 19. Nicholson, W.L.; Munakata, N.; Horneck, G.; Melosh, H.J.; Setlow, P. (2000). Resistance of Bacillus endospores to extreme terrestrial and extraterrestrial environments. Microbiol. Mol. Biol. Rev., 64 (3), 548-572.
- 20. Nieminen, T.; Rintaluoma, N.; Andersson, M.; Taimisto, A.M.; Ali-Vehmas, T.; Seppälä, A.; Priha, O.; Salkinoja-Salonen, M. (2007). Toxinogenic Bacilluspumilus and Bacillus licheniformis from mastitic milk. Vet. Microbiol., 124 (3-4), 329-339.
- 21. Obi, S.K. (1980). Lecithinase and toxin production in Bacillus species. Zentralbl.Bakteriol. A., 246(3), 415-422.
- 22. Ortigosa, M.; Garay, E.; Pujalte, M-J. (1997). Gram-positive bacteria of marine origin: a numerical taxonomic study on Mediterranean isolates. Microbiologia, 13 (4), 453-462.
- 23. Ozkocaman, V.; Ozcelik, T.; Ali, R.; Ozkalemkas, F.; Ozkan, A.; Ozakin, C.; Akalin, H.; Ursavas, A.; Coskun, F.; Ener, B.; Tunali, A. (2006). Bacillus spp. among hospitalized patients with haematological malignancies: clinical features, epidemics and outcomes. J. Hosp. Infect., 64 (2), 169-176.
- 24. Peltola, J.; Andersson, M.A.; Haahtela, T.; Mussalo-Rauhamaa, H.; Rainey, F.A.; Kroppenstedt, R.M.; Samson, R.A.; Salkinoja-Salonen, M.S. (2001). Toxic-metabolite-producing bacteria and fungus in an indoor environment. Appl. Environ. Microbiol., 67 (7), 3269-3274.
- 25. Priest, F.G. (1993). Systematics and ecology of Bacillus In: Hoch, J.A.; Losick, R. (eds.). Bacillus subtilis and other Gram-positive bacteria: Biochemistry, physiology and molecular genetics ASM Press, Washington DC.
- 26. Rasko, D.A.; Rosovitz, M.J.; Okstad, O.A.; Fouts, D.E.; Jiang, L.; Cer, R.Z.; Kolsto, A.B.; Gill, S.R.; Ravel, J. (2007). Complete Sequence analysis of novel plasmids from emetic and periodontal Bacillus cereus isolates reveals a common evolutionary history among the B. cereus-group plasmids, including Bacillus anthracis pXO1. J. Bacteriol., 189 (1), 52-64.
- 27. Ruger, H.J. (1989). Benthic studies of the northwest African upwelling region psychrophilic bacterial communities from areas with different upwelling intensities. Mar Ecol. Prog. Ser., 57, 45-52.
- 28. Sasser, M. (1990). Identification of bacteria through fatty acid analysis. In: Klement, Z.; Rudolph, K.; Sands, D.C. (eds.). Methods in Phytobacteriology Akademiai Kiado, Budapest, Hungary.
- 29. Smibert, R.M.; Kreig, N.R. (1994). Phenotypci characterization. In: Gerhardt, P.; Murray, R.G.E.; Wood, W.A.; Kreig, N.R. (eds.). Methods for general and molecular bacteriology ASM Press, Washington DC.
- 30. Sneath P.H.A.; Mair N.S.; Sharpe, M.E.; Holt, J. (1986). Bergey's manual of systematic bacteriology, Vol. 2 Williams and Wilkins, Baltimore.
- 31. Stead, D.E.; Sellwood, J.E.; Wilson, J.; Viney, I. (1992). Evaluation of a commercial microbial identification system based on fatty acid profiles for rapid, accurate identification of plant pathogenic bacteria. J. Appl. Bacteriol., 72 (4), 315-321.
- 32. Suominen, I.; Andersson, M.A.; Andersson, M.C.; Hallaksela, A.M.; Kampfer, P.; Rainey, F.A.; Salkinoja-Salonen, M. (2001). Toxic Bacilluspumilus from indoor air, recycled paper pulp, Norway spruce, food poisoning outbreaks and clinical samples. Syst.Appl. Microbiol., 24 (2), 267-276.
- 33. Tena, D.; Martinez-Torres, J.A.; Perez-Pomata, M.T.; Sáez-Nieto, J.A.; Rubio, V.; Bisquert, J. (2007). Cutaneous infection due to Bacillus pumilus: report of 3 cases. Clin. Infect. Dis. 44 (4), 40-42.
- 34. Woese, C.R. (1987). Bacterial evolution. Microbiol. Rev., 51(2), 221-271.
- 35. Yamamoto, S.; Harayama, S. (1995). PCR amplification and direct sequencing of gyrB genes with universal primers and their application to the detection and taxonomic analysis of Pseudomonas putida strains. Appl. Environ. Microbiol., 61 (3), 1104-1109.
- 36. Yokoyama, K.; Ito, M.; Agata, N.; Isobe, M.; Shibayama, K.; Horii, T.; Ohta, M. (1999). Pathological effect of synthetic cereulide, an emetic toxin of Bacillus cereus, is reversible in mice. FEMS Immunol. Med. Microbiol., 24 (1), 115-120.
Publication Dates
-
Publication in this collection
24 July 2009 -
Date of issue
June 2009
History
-
Accepted
26 Feb 2009 -
Reviewed
10 July 2008 -
Received
04 Apr 2008