Abstract
Studies on lipase production were carried out with a bacterial strain (Bacillus sp LBN 2) isolated from soil sample of hotspring of Arunachal Pradesh, India. The cells were cultivated in a mineral medium with maximum production at 1% groundnut oil. The optimum temperature and initial medium pH for lipase production by the organism were 50ºC and 9.0 respectively. The molecular mass was found to be 33KDa by SDS PAGE. The optimal pH and temperature for activity were 10 and 60ºC respectively. The enzyme was found to be stable in the pH range of 8-11 with 90% retention of activity at pH 11. The enzyme retained 90% activity at 60ºC and 70% of activity at 70ºC for 1h. The lipase was found to be stable in acetone followed by ethanol. The present findings suggested the enzyme to be thermophilic alkaline lipase.
Thermophilic bacteria; Thermostable; Alkaline lipase and Bacillus sp
INDUSTRIAL MICROBIOLOGY
Optimization of extracellular thermophilic highly alkaline lipase from thermophilic Bacillus sp isolated from Hotspring of Arunachal Pradesh, India
Limpon BoraI,* * Corresponding Author. Mailing address: Dept of Molecular Biology & Biotechnology, Tezpur University, Tezpur-784028.; E-mail: limponbioera@gmail.com ; Minakshi BoraII
IDept of Molecular Biology & Biotechnology, Tezpur University, Tezpur-784028
IIDept of Biotechnology, Dibrugarh University, Dibrugarh - 786004
ABSTRACT
Studies on lipase production were carried out with a bacterial strain (Bacillus sp LBN 2) isolated from soil sample of hotspring of Arunachal Pradesh, India. The cells were cultivated in a mineral medium with maximum production at 1% groundnut oil. The optimum temperature and initial medium pH for lipase production by the organism were 50ºC and 9.0 respectively. The molecular mass was found to be 33KDa by SDS PAGE. The optimal pH and temperature for activity were 10 and 60ºC respectively. The enzyme was found to be stable in the pH range of 8-11 with 90% retention of activity at pH 11. The enzyme retained 90% activity at 60ºC and 70% of activity at 70ºC for 1h. The lipase was found to be stable in acetone followed by ethanol. The present findings suggested the enzyme to be thermophilic alkaline lipase.
Key words: Thermophilic bacteria, Thermostable, Alkaline lipase and Bacillus sp
INTRODUCTION
Lipases (Triacylglycerol acylhydrolase, EC 3.1.1.3) constitute a group of enzymes defined as carboxyesterases that hydrolyse long chain acylglycerols at the lipid water interface (7,5). The natural function of lipases is to hydrolyze triglycerides into diglycerides, monoglycerides, fatty acids and glycerol. In addition to triacyl glycerol a number of other low and high molecular eight carboxylic-, thiol-, poly-acid esters and amides are also accepted as substrates by these hydrolase enzymes.
Most lipases isolated and reported are found to be mesophilic in nature, which cannot hydrolyze a substrate that exists in solid form at room temperature (4). To overcome this problem a search for thermophilic nature of enzyme is on as thermophilic lipases show higher thermostability, higher activity at elevated temperatures, and often shows more resistance to chemical denaturation. This makes them ideal tools in industrial and chemical processes where relatively high reaction temperatures and organic solvents are used. Thermostable enzymes are usually derived from thermophilic microbial strains, which may be expected to produce intrinsically more heat stable enzymes. The industrial demand for the thermostable enzymes continues to stimulate the search of novel thermophilic microorganisms from various unexploited regions of the earth, as small numbers of bacterial strains producing thermophilic lipases have been reported in the last decade (13).
Optimization of fermentation conditions for microbial lipase is of great interest, since culture conditions influence the properties of the enzyme produced as well as ratio of extra cellular and intracellular lipases (37). Therefore, optimization of the lipase production has been focused on improving fermentation conditions such as carbon or nitrogen source, temperature, pH, aeration, using inducers and source of inoculum etc (31, 33, 11, 4, 17). The induction in lipase production in microorganisms is a process that results in change in the phenotype, which allows the production of energy, required for its metabolism and microbial growth. Edible oils such as vegetable oils, canola oil, and olive oil are used mainly as inducers.
Because of huge variation in application and emerging need of lipases with specific characterstics has prompted to search for new lipases from newer areas with different characterstics. In the present study, we report the findings of thermophilic alkaline lipases produced by thermophilic Bacillus sp isolated from hotsprings of Arunachal Pradesh, India. These findings will provide a meaningful addition to the database on bacterial lipase research in the screening and identification of lipolytic bacteria. The Bacillus strain was sent to IMTECH Chandigarh for identification and will be deposited in MTCC after identification
MATERIALS AND METHODS
Microorganism, media, and growth conditions
Bacillus sp was isolated from hot spring of Dirang district of Arunachal Pradesh, India. All analytical chemicals and media components used were of highest purity grade available commercially. A shaking water bath with temperature controller was used. All optical measurements were performed on UV visible spectrophotometer using matched cells of path length 1cm.
The Bacillus sp isolated were maintained on Nutrient agar slants. The mineral medium was composed as (w/v) MgSO4.7H2O 0.04%, KH2PO4 0.3%, Na2CO3 0.1% and NaCl 0.01%. The pH of the medium was adjusted to 10.0.
Experimental procedures
Lipase production: The mineral medium (50ml in 250ml Erlenmeyer flasks) was prepared and the olive oil was added after emulsification for 15 minutes as sole carbon source. The sodium carbonate was sterilized separately and added to the rest of the medium to adjust the pH upto 10.0. The medium was inoculated with 104 cellsmL-1 of an overnight culture and incubated at 50ºC on shaking incubator at 200rpm for 48h. The cultures so obtained were centrifuged at 10,000g for 15 min. The cell free supernatant thus obtained was used as source of enzyme. The samples were taken and assayed for lipase activity at regular intervals. The measurement of enzyme activity was done in triplicates.
Lipolytic activity
Enzyme activity was assayed spectrophotometrically (Beckman model DU 530, UV-visible spectrophotometer, Beckman instruments Inc., Fullerton, CA, USA) using p-nitrophenyllaurate (p-NPL) (Sigma, St Louis, MO, USA) as substrate. The reaction mixture consisted of 0.8mL of 0.05 mol L−1 phosphate buffer (pH 8.0), 0.1mL of enzyme and 0.01 mol L−1 p-NPL. The mixture was incubated at 50ºC for 15 min, after which 0.25mL of 0.1mol L−1 Na2CO3 was added and the release of p-nitrophenol was measured at 410 nm. One unit of enzyme corresponds to the release of 1mmol of p-nitrophenol liberated per minute under standard assay conditions. The lipase activity was done in initial rate conditions and the activity was measured over time (35).
Determination of protein
Protein content was measured following the method of Lowry et al. (1951) using Bovine Serum Albumin as standard (24).
Time course of lipase production
Time course of the lipase production was studied with the medium in shake flasks, for 70 h. 5mgmL-1 inoculum was added to 50 ml medium, in 500 ml Erlenmeyer flask and incubated at 220 rpm on a rotary shaker, at 50ºC, for 70 h. Samples were removed periodically and growth as well as lipase activity in the culture supernatant was determined. The growth was measured in terms of OD at 620nm. Productivity was defined as lipolytic activity units per gram of dry biomass.
Optimization of culture conditions for production of lipase
The effect of carbon source on the production of lipase, different carbon sources including glucose, galactose, maltose, fructose, trehalose, mannose and sucrose at 1% (w/v) concentration were added to the medium. Various nitrogen sources including malt extract, peptone, ammonium sulphate, yeast extract, beef extract, meat extract, and casein at 1% (w/v) was supplemented to the production medium. These data were obtained using 100mL of the basal medium at an initial pH of 10.0 at 50ºC for 48 h. The effect of initial pH on lipase production was studied using shake flask culture at different initial value of pH. The pH was adjusted to the desired value using 6N NaoH or 6N HCl. The effect of temperature on lipase production was studied by incubating the organism at different temperature (30- 80ºC ). The effect of NaCl concentration on the enzyme activity was determined at different salt concentration of 0, 5, 10 and 15% NaCl (w/v) at pH and temperature optimum.
Purification of lipase
The bacterial isolate was grown for 48 h at 50ºC under submerged fermentation. The enzyme of cell free supernatant was precipitated by ammonium sulphate at 60% saturation at 4ºC. The precipitate was collected by centrifugation at 10,000g for 15 min at 4ºC and dissolved in 1M tris buffer of pH 7.0. The ammonium sulphate which was associated with precipitate, was removed by dialysis against distill water with nitrocellulose dialyzing bag. The dialysed protein was purified using Sephadex G-100 column (2 cm dia ×150 cm length) gel filtration chromatography. The elution was allowed to proceed at 1ml/min of flow rate. The column was run and the fractions were collected in 3ml volume. UV monitor at 280nm measures the absorbance of the fractions. The quantitative activity of lipase was determined as previously described.
Effect of pH on activity and stability
The pH effect on activity and stability profiles were studied in a pH range of 3.0- 12.0 using different buffers at 50mM concentration. For stability 1 ml of enzyme (5mg/ml) was mixed with respective buffer and incubated from 0.5 to 24 hours at 50ºC and then estimating the residual activity.
Effect of temperature on activity and stability
Temperature effect on lipase activity was determined by carrying out the enzyme reaction at different temperatures in the range of 10-80ºC at pH 10.0. The thermo stability of the lipase was determined by preincubating the enzyme (5mg/ml) at different temperatures ranging form 30-80ºC for 1 h. After treatment the residual enzyme activities were determined.
Effect of surfactants and commercial detergents on enzyme stability
Effect of surfactants (Tween 20, Tween 40, Tween 80, Triton X-100, sodium cholate and Sodium dodecyl sulphate (SDS)) and commercial detergents (Ariel, Surf, Rin, Wheel, Tide, and Fena) on enzyme stability was determined under standard assay conditions. 50µl of the enzyme sample were incubated in 50µl of surfactants and detergents (Solution 0.2, 1.0 and 10%w/v at pH 8.0) for 1 hour in 50ºC at pH 10.0 and the residual activity was determined at pH 10.0 for 10 Minutes.
Substrate specificity of the enzyme
The substrate specificity of the enzyme was studied by titrimetry. The reaction was done in 20 ml Erlenmeyer flask. The substrates used were, castor oil, coconut oil, cottonseed oil, groundnut oil, olive oil, sunflower oil, Span 20, triacetin and tributyrin. The reaction mixture was composed of 1 g of the above substrate, 23.9 ml glycine-NaOH buffer (0.1 M, pH 9.0) and 0.1 ml enzyme. The mixture was incubated at 40ºC on rotary shaker at 200 rpm for 1 h. The reaction was terminated by addition of 50 ml of acetone-ethanol mixture (1:1). The fatty acids liberated were estimated by titration with 0.05 M NaOH. One unit of enzyme was defined as the amount of enzyme, which liberated one μ mol of fatty acid, per minute. The end point of the enzyme reaction was 11.0 and the enzyme activity was analyzed at initial reactions.
Effect of organic solvents on lipase activity
To study the effect of organic solvents on lipase activity was analysed by incubation of enzyme for 1h at 40°C in 50mM Tris-Cl buffer (pH 7.0) containing 30%v/v organic solvents. The control was estimated without organic solvents. The activity was measured at the beginning and the end of the incubation.
RESULTS AND DISCUSSION
Time course of lipase production
The production of lipase by Bacillus sp was correlated with growth curve/ biomass. The biomass increased up to 56 hours, after that there was a decline in its amount. Extracellular lipase production exhibited a similar increase with biomass and remains in higher amount in medium upto 72h with the production at stationary phase (Fig. 1). The lipase activity appeared to couple with growth .The enzyme production was found to be maximum when the cell population entered into the stationary phase suggesting that enzyme secretion is growth associated. The lipase activity was observed to start soon after beginning of incubation, and the crude enzyme extract obtained after 48 h of growth in the culture medium exhibited highest activity at the beginning of the stationary phase. A decrease in lipase activity was apparent during the late stationary phase presumably due to the presence of proteases in the culture medium. These results are in agreement with earlier results (32, 14) where maximum lipase activities were observed at the onset of the stationary phase of bacterial growth.
Effect of carbon source on lipase production
The lipase production was studied with different categories of carbon substrates such lipids, complex nutrient media and readily assimilated substrates. Among these carbon substrates the maximum lipase production was observed in groundnut oil, but it has been found that Bacillus sp could able to utilize a number of other vegetable oils as well as other carbon sources. The maximum lipase production of 18 U/ml was observed in groundnut oil (Fig. 2). Among the sugars, sucrose and maltose were found to be good substrates for lipase production. The major factor for the expression of lipase activity has always been the carbon source, since lipases are by and large inducible enzymes (23). The enzyme production level markedly increased in the presence of lipid carbon substrates and represents the key mechanism for making a fatty acid carbon source available to the cell during growth on such compounds. The enzyme is probably induced by these substrates (1, 15) as fatty acid esters are efficient effectors leading to a significant increase in the enzyme level relative to that obtained with nonlipid substrates such as glucose.
Effect of nitrogen source on lipase production
The Bacillus sp grew well in all the nitrogen sources used in this study. Among the different nitrogen sources tested, peptone was found to be the best among all the nitrogen sources tested with a production of 20U/ml (Fig. 3). The reason to this effect is the release of NH4+ ions from peptone which stimulates the growth and at the same time increase the enzyme yield because of its protease inhibiting nature at low concentration. The results obtained are in agreement with that reported by Gulati et al. (2005) where peptone was found to be the best nitrogenous source for lipase production. Similar findings were also reported in other Bacillus sp (8, 21, 34, 38 12). The lipase production was found to be low with soybean meal and corn steep liquor, whereas in other organic and inorganic nitrogen sources the lipase production was in the range of 10-17 U/ml.
Effect of temperature and pH on lipase production
The influence of temperature on lipase production was studied in the range of 30-80ºC while keeping all the other conditions constant. The optimum temperature for lipase production lipase was found to be 50ºC (Fig. 5). The temperature regulates the enzyme synthesis at mRNA transcription level and probably translation levels of proteins, thus increases the stability of the proteins and also the production. However the other possible reason for production at elevated temperature may be temperature influences their secretion; possibly by changing the physical properties of the cell membrane (26). The production trend of Bacillus sp LBN2 was found to be at alkaline range pH of 8.0 to 10.0. The maximum production obtained when the initial pH of the medium was 9.0 (Fig. 4). Medium pH is very important in nutrients absorption and growth of bacteria, stimulation of enzyme production via signaling pathways and release of extra cellular enzymes based on proteolytic mechanism of signal peptidases (27).
The optimum temperature for lipase production corresponds with the growth temperature for lipase production (11). Our results are considerably higher with most of the findings reported earlier (30).
Effect of Salt concentration on Lipase production
The lipolytic enzyme presents a wide range of tolerance to NaCl with activation at concentration between 5 and 10% suggesting the enzyme might be an salt dependent enzyme and salt plays an essential role in either in its activity or stability (Fig. 6). The enzyme production at high salt concentration suggest that the Bacillus Sp from which the enzyme was isolated is a halotolerant bacteria.
Purification of the enzyme
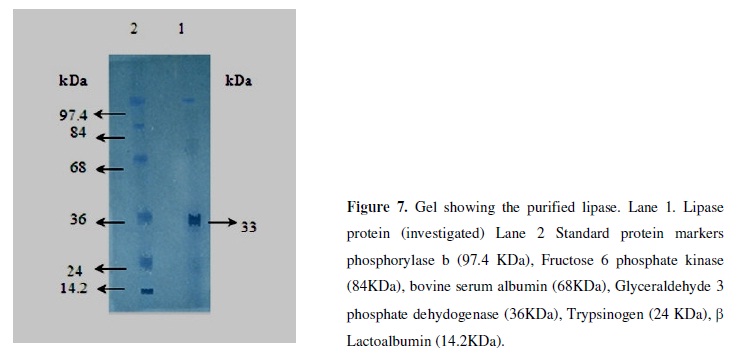
The purification steps used in this study are lesser than the reports reported by various authors (27, 14). The Enzyme was purified and characterized. The reason for the application of shorter steps is of the fact that in each step of enzyme purification an average of more than 10% of the enzyme is lost and this led to the low recovery of the enzyme (Table 1). Hence to reduce the enzyme costs it is necessary to lower the number of the processing steps in order to increase the recovery. Further importance as in our study the enzyme could be recovered at lesser steps which simplified the tedium downstream processing for isolation of pure lipase and thus lower the cost of enzyme and hence influence the overall process of economy and thus of great industrial importance.
The molecular weight of the enzyme was estimated to be 33 kDa by SDS-PAGE (Fig. 7). The pure protein smeared badly on native PAGE but showed single band with molecular weight of 33 kDa on SDS-PAGE. This indicated that the lipase might be forming high molecular weight aggregates with other lipophilic materials. Lipases are reported to be monomeric proteins having molecular weights in the range of 16 kDa-90 kDa (9, 25, 30, 17). In the present study it was found to have molecular weight in the aforesaid range therefore it might be a monomeric protein. Sodium dodecyl sulphate poly acryl amide gel electrophoresis analysis revealed the lipase protein to be monomer with a single distinct band. The molecular weight of the protein was determined to be 33 KDa.
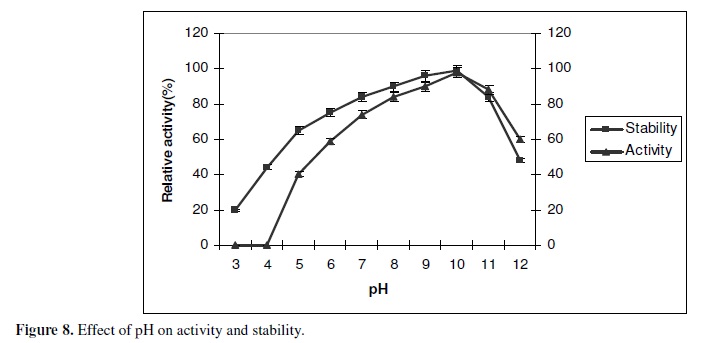
Effect of pH on the enzyme activity and stability
The effect of pH on enzyme activity was studied in the pH range of 3.0-10 at 50ºC for 1 Hour. In the present work the optimum pH of the alkaline lipase was found to be 10.0. The enzyme was found to be stable in the pH range of 8-11 (Fig. 8). The lipase activity showed marked activation at alkaline pH of 8-10 with the retention of 90% of the residual activity at pH 11. Based on the present findings the enzyme was stated to be alkaline lipase. The stability of the lipase at higher pH, could be explained that the enzyme in the present study may contains more acidic amino acid that are exposed to or excreted to the external medium and a Na+ cycle that facilitates the solute uptake. The pH optima in the present finding are relatively higher in comparison to the results reported earlier. In Bacillus thermoleovorans ID 1 (22), pH 6.5 in Staphylococcus aureus (25), pH 8.0 in Bacillus sp RSJ1 (33), pH 7.0 in Fusarium oxyporum (28), with an optimum pH in most of the Bacillus sp in the range of 5-8 (11).
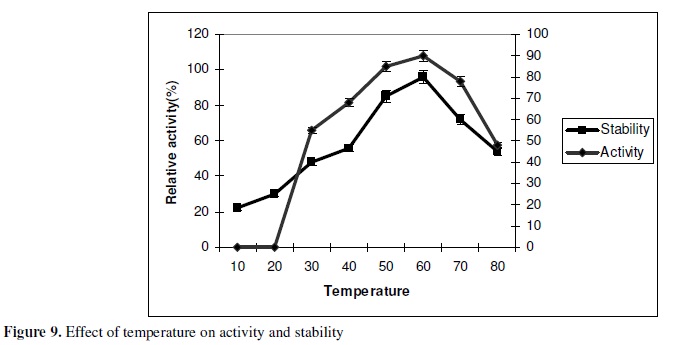
Effect of temperature on lipase activity and stability
Lipase activity was assayed at different temperatures ranging from 10-80ºC at pH 10. The optimum reaction temperature was 60ºC . Thermal stability was investigated by incubating the enzyme at various temperatures (30-80ºC ) for 1h. Thermo stability profile indicated that the enzyme showed great stability up to 60ºC (Fig. 9). The lipase seems to keep 90% of activity at 60ºC and almost 70% 0f the activity retained at 70ºC . The enzyme showed better stability than those reported by earlier authors (16,19). The thermal stability of the lipase and retention of activity over 60ºC could be explained by the presence of polyamines in the protein structure. However increased number of hydrogen bonds, salt bridges and higher proportion of thermophilic amino acids may also contribute towards the thermal stability of the enzyme at higher temperature. The results obtained are in agreement with lipase from Pseudomonas sp KWI-56 which was found to be stable at 60ºC but at a lower pH of 7.0 (6). Presently it is important to develop enzymes active and stable in broad range of temperatures, essentially due to the energy costs involved in the heating of the water, and the use of new and fragile materials. Higher temperatures are often used to remove difficult strains, such as food stains in cotton or synthetic materials.
Effect of Surfactants and detergents on lipase activity
The effects of various surfactants and commercial detergents on the alkaline lipase activity were tested at 10% and 0.1% (w/v) (Table 2) shows that the enzyme is stable in both ionic and nonionic surfactants. Tween 40 and Tween 80 were found to inhibit the lipase activity after 1h of incubation at 50ºC . On the other hand, Triton X-100 and Triton X-114. It is interesting to note that Triton x-100 and Triton x-114 stimulated the enzyme activity up to 60% in our study. The reason may be that surface active agents might have increased the turnover number of lipase by increasing the contact frequency between the active site of the enzyme and the substrate by lowering the surface tension of the aqueous medium and with the maintenance of open form of lipase which may enhance the activity. SDS was found to be strong inhibitor as in the case of Bacillus thermoleovorans (7) lipases causing almost total inhibition of enzyme activity. The cause of strong inhibition by SDS could be due to the local conformation changes in the active site of the enzyme molecule that results in inhibition, partial reversible unfolding, and subsequent inactivation.
Substrate specificity of enzyme
Microbial lipases have been divided into three categories namely non specific, regiospecific and fatty acid specific based on the substrate specificity. The present lipase caused rapid hydrolysis of the vegetable oils containing C-16 and C18 fatty acid ester suggesting that the present lipase is a "true" lipase. It acted preferentially on oils with lower chain fatty acid. It showed marked activation with tributyrin as compared to triolein (20) (Table 3). The activity of lipase in palm oil was considred to be 100% activity. Triacetin was found to be poor substrate and was hardly hydrolyzed. On the basis of present results the lipolytic activity is found to be dependent upon the unsaturated fatty acids as lipolytic activity increased with increase in the degree of unsaturation. Lipases from Bacillus thermocatenulatus, Staphylococcus hyicus and Staphylococcus epidermidis found to have hydrolysed tributyrin (31).
Effect of organic solvents on lipase activity
Acetone was found to be best solvent for the extraction of lipase which showed an increase in lipase activity over the control. However the activity of the purified lipase was decreased by the addition of 40% of organic solvents including methanol, and isopropyl alcohol (Table 4). In contrast to the present study acetone was found to be the best solvent for the lipase. The present lipase is found to be stable in organic solvents with enhanced activity in the presence of acetone. The present findings were in agreement to that reported for Bacillus thermocatenulatus (31) and Pseudomonas sp. (17) where acetone has found to enhance enzyme activity. The observed stability of lipase is suggestive of the notion that the solvent maintains the enzyme in open conformation; the lid of the enzyme does not cover the active site crevice, and thus maintains the flexible conformation. The organic solvents were used in the concentration of 30% and not 1%.
CONCLUSION
It is evident from the above study that the lipase has the requisite properties of significant industrial importance. A lipase that is stable at high alkaline conditions and high temperature is rare. In the present study Bacillus sp. LBN2, which produces inducible, extracellular, alkalophilic lipase stable in commercial detergents and surfactants, makes it ideally suited for detergent formulations. However, further investigation is required to ascertain the molecular characteristics, kinetic properties and substrate specificity to access the true potential of this microbial lipase.
Submitted: November 05, 2009; Returned to authors for corrections: July 19, 2011; Approved: January 16, 2012.
- 1. Abdel-Fattah, Y.R. (2002). Optimization of thermostable lipase production from a thermophilic Geobacillus sp. using Box-Behnken experimental design. Biotechnol Lett 24:1217-1222.
- 2. Ateslier, Z.B.B.; Metin, K. (2006). Production and partial characterization of novel thermostable esterase from a thermophilc Bacillus sp. Enzyme Microb Technol 38:628-635.
- 3. Baral, A.; Fox, P.F. (1997). Isolation and characterization of an extracellular lipase from Pseudomonas tolaassi Fd.. Chem 58: 33-38.
- 4. Bayoumi, R.A.; El-louboudey, S.S.; Sidkey, N.M.; Abd-El-Rahman, M.A. (2007). Production, Purification and Characterization of Thermoalkalophilic Lipase for Application in Bio-detergent Industry. J. Apll. Sci. Res 3: 1752-1765.
- 5. Boekema, K.H.A.; Beselin, M.; Breuer, B.; Hauer, M.; Koster, F.; Rosenau, K.; Tommassen, J. (2007). Hexadecane and Tween 80 stimulate lipase production in Burkholderis glume by different mechanisms. Appl. Environ. Microbiol, 73: 3838-3844.
- 6. Brune, A.K.; Gotz, F. (1992). Degradation of lipids by bacterial lipases. In Winkelmen G(ed) Microbial degradation of natural products, VCH, Weinhein, pp 243-266.
- 7. Castro-Ochoa, L.D.K.; Rodriguez-Gomez, G.; Valerio-Alfaro, R.; Oliart-Ros (2005). Screening, purification, and characterization of the thermoalkalophilic lipase produced by Bacillus thermoleovorans CCR11. Enzyme and Microbial Technology, 37: 648-654.
- 8. Ghanem, E.H.; Al-Sayed, H.A.; Saleh, K.M. (2000). An alkalophilic thermostable lipase produced by new isolate of Bacillus alcalophilus World. J. Microbiol. Biotechnol 16: 459-464.
- 9. Gilbert, E.J.; Cornish, A.; Jones, C.W. (1991b). Purification and properties of extrcellular lipase from Pseudomonas aeruginosa EF2. J. Gen. Microbiol. 137: 2223-2229.
- 10. Gulati, R.; Isar, J.; Kumar, V.; Prasad, A.K.; Parmar, V.S.; Saxena, R.K. (2005). Production of novel alkaline lipase from Fusarium globosum using neem oil. and its applications. Pure Appl Chem 77: 251-262.
- 11. Gupta, R.; Gupta, N.; Rathi, P. (2004). Bacterial lipases: an overview of production, Purification and Biochemical properties. Appl Microbiol Biotechnol 64: 763-781.
- 12. Kanwar, L.; Gogoi, B.K.; Goswami, P. (2002). Production of Pseudomonas lipase in n- Alkane substrate and its isolation using ammonium sulphate precipitation technique. Bioresource Technol 84: 207-211.
- 13. Kader, R.; Yousuf, A.; Hoq, M.M. (2007). Optimization of lipase production by Rhizopus MR12 in shake culture. J Appl Sci 7: 855-860.
- 14. Kambourova, M.; Kirilova, N.; Mandeva, R.; Derekova, A. (2003). Properties of thermostable lipase from a thermophilic Bacillus steathermophilus MC7. J Mol Cat B. Enzymatic 22: 307-313.
- 15. Kaushik, R.; Saran, S.; Isar, J.; Saxena, R.K. (2006). Statistical optimization of medium components and growth conditions by response surface methodology to enhance lipase production byAspergillus carneus J Mol Catal B: Enzym 40:121-126.
- 16. Kim, M.H.; Kim, H.K.; Lee, J.K.; Park, S.Y.; Oh, T.K. ( 2000). Thermostable lipase from Bacillus steathermophilus: high level production, purification, and calcium dependent thermostability. Biosci Biotechnol Biochem 64: 280-286.
- 17. Kiran, S.G.; Shanmughapriya, S.; Jayalakshmi, J.; Selvin, J.; Gandhimathi, R.; Sivaramakrishnan, S.; Arunkumar, M.; Thangavelu, T.; Natrajaseenivasan, K. (2008). Optimization of extracellular psychrophilic lipase produced by marine Pseudomonas sp. (MSIO57). Bioprocess Biosyst Eng 31: 483-492.
- 18. Konstantinou, P.; Roussis I..G. (1998). Some properties of extracellular lipase from Pseudomonas 92. Mlichwissenschaft 53:622-624.
- 19. Kulkarni, N.; Gadre, R.V. (2002). Production and properties of an alkaline thermophilic lipase from Pseudomonas flourescens NS2W. J Ind Food Microbiol 28: 344-348.
- 20. Kumura, H.; Mikawa, K.; Saito, Z. (1993) Purification and characterization of lipase from Pseudomonas fluorescens No. 33. Milchwissenschaft 48: 431-434.
- 21. Lanser, A.C.; Manthey, L.K.; Hou, C.T. (2002). Regioselectivity of new bacterial lipases detemined by hydrolysis of triolein. Curr Microbiol 44: 336-340.
- 22. Lee, D.W.; Koh, Y.S.; Kim, K.J.; Kim, B.C.; Choi, H.J.; Kim, D.S.; Suhartono, M.T.; Pyun, Y.R. (1999). Isolation and characterization of thermophilic lipase from Bacuillus thermoleovorans ID-1. FEMS Microbiol Lett 179: 393-400.
- 23. Lotti, M.; Monticelli, S.; Montesinos, J.L.; Brocca, S.; Valero, F.; Lafuente, J. (1998). Physiological control on the expression and secretion of Candida rugosa lipase. Chem Phys Lipids 93: 143-148.
- 24. Lowry, O.H.N.G.; Rosebrough, A.L.; Farr Randall, R.J. (1951). Protein measurement with the Folin- phenol reagent. J. Bio. Chem 193: 265-275.
- 25. Paiva, A.L.; Balaco, V.M.; Malacta, F.X. (2000). Review: Kinetics and mechanisms of reaction catalaysed by immobilized lipases. Enz Microbiol Technol 27: 187- 204.
- 26. Paetzel, M.; Karla, A.; Strynadka, N.C.J.; Dalbey, R.E. (2002). Signal Peptidases. Chem Rev 2002, 102:4549-4579.
- 27. Palekar, A.A.; Vasudevan, P.T.; Yan, S. (2000). Purification of lipase: A review. Biocatal Biotransform 18: 177-200.
- 28. Prazeres, J.N.; Cruz, J.A.B.; Pastore, G.M. (2006). Characterization of an alkaline lipase from Fusarium oxysporum and the effect of different surfactants and detergents on the enzyme activity. Braz J Microbiol 37: 505-509.
- 29. Rahman, R.N.Z.A.; Lee, P.G.; Basri, M.; Salleh, A.B. (2005). Physical factors affecting the production of organic solvent-tolerant protease by Pseudomonas aeruginosa strain K. Bioresource Technol 96:429-436.
- 30. Rathi, P.; Saxena, R.K.; Gupta, R. (2001). A novel alkaline lipase from Burkholderia cepacia for detergent formulation. Process Biochem 37: 187-192.
- 31. Schmidt-Dannert, C. (1999). Recombinant microbial lipases for biotechnological application. Bioorg Med Chem 7: 2123-2130.
- 32. Schuepp, C.; Kermasha, S.; Michalski, M.C.; Morin, A. (1997). Production, partial purification and characterization of lipases from Pseudomonas fragi CRDA 037. Process Biochem 32: 225-232.
- 33. Sharma, R.; Chisti, Y.; Banerjee, U.C. (2001). Production, Purification, Characterization and applications of lipases. Biotechnol. Advances 19: 627-662.
- 34. Sharma, R.; Soni, S.K.; Vohra, R.M.; Gupta, L.K.; Gupta, J.K. (2002). Purification and characterization of a thermostable alkaline lipase from a new thermophilic Bacillus sp RSJ -1. Process Biochem 37: 1075-1084.
- 35. Sigurstdair, S.; Konraosdottir, M.; Jonsson, A.; Kristjansson, J.K.; Mathiasson, E.; (1993).Lipase activity of thermophilic bacteria from Icelendic hotsprings. Biotechnol Lett 15: 361-366.
- 36. Sugihara, A.; Tadaki, T.; Yosio, T. (1991). Purification and characterization of a novel thermostable lipase from Bacillus sp. J. Biochem. 109: 211-216.
- 37. Taipa, M.A.; Aires-Barros, M.R.; Cabral, J.M.S. (1992). Production of lipases. J. Biotechnol 26: 111-142.
- 38. Thomas, A.; Manoj, M.K.; Valsa, A.; Mohan, S.; Manjula, R. (2003). Optimization of growth condition for the production of extra cellular lipase by Bacillus mycoides, Indian J. Microbiol 43: 67-69.
Publication Dates
-
Publication in this collection
02 May 2012 -
Date of issue
Mar 2012
History
-
Received
05 Nov 2009 -
Accepted
16 Jan 2012 -
Reviewed
19 July 2011