Abstract
This study evaluated 26 pigeonpea rhizobial isolates according to their cultural characteristics, intrinsic antibiotic resistance, salt and temperature tolerance, carbon source utilization and amylolytic activity. The cultural characterization showed that the majority of them presented the ability to acidify the YMA. Among the 27 isolates evaluated, 25 were able to grow when incubated at 42° C and 11 showed tolerance to 3% (w/v) of NaCl in YMA medium. The patterns of carbon sources utilization was very diverse among the isolates. It was observed the capacity of three strains to metabolize all the carbon sources evaluated and a total of 42% of the bacterial isolates was able to grow in the culture medium supplemented with at least, six carbon sources. The carbon sources mannitol (control) and sucrose were metabilized by all isolates evaluated. The profile of intrinsic resistance to antibiotics showed that the isolates were mostly resistant to streptomycin and ampicillin, but susceptible to kanamycin and chloranphenicol. High amylolytic activity of, at least, four isolates was also demonstrated, especially for isolated 47.3b, which showed the highest enzymatic index. These results indicate the metabolic versatility of the pigeonpea rhizobia, and indicates the isolate 47.3b to further studies regarding the amylase production and characterization.
strain selection; biological nitrogen fixation; biotechnology
MICROBIAL PHYSIOLOGY
Phenotypic diversity and amylolytic activity of fast growing rhizobia from pigeonpea [Cajanus cajan (L.) Millsp.]
Paulo Ivan Fernandes JúniorI, * * Corresponding Author. Mailing address: Embrapa Semiárido, BR 428, Km 152, Zona Rural, CEP.: 56302-970, Petrolina-PE, Brasil.; E-mail: paulo.ivan@embrapa.br ; Andréa Aparecida de LimaII; Samuel Ribeiro PassosII; Carlos Alberto Tuão GavaI; Paulo Jansen de OliveiraIII; Norma Gouvêa RumjanekIV; Gustavo Ribeiro XavierIV
IEmbrapa Semiárido, Petrolina, PE, Brasil
IIUniversidade Federal Rural do Rio de Janeiro, Instituto de Agronomia, Departamento de Solos, Seropédica, RJ, Brasil
IIIUniversidade Federal Rural do Rio de Janeiro, Instituto de Tecnologia, Departamento de Engenharia Química, Seropédica, RJ, Brasil
IVEmbrapa Agrobiologia, Seropédica, RJ, Brasil
ABSTRACT
This study evaluated 26 pigeonpea rhizobial isolates according to their cultural characteristics, intrinsic antibiotic resistance, salt and temperature tolerance, carbon source utilization and amylolytic activity. The cultural characterization showed that the majority of them presented the ability to acidify the YMA. Among the 27 isolates evaluated, 25 were able to grow when incubated at 42° C and 11 showed tolerance to 3% (w/v) of NaCl in YMA medium. The patterns of carbon sources utilization was very diverse among the isolates. It was observed the capacity of three strains to metabolize all the carbon sources evaluated and a total of 42% of the bacterial isolates was able to grow in the culture medium supplemented with at least, six carbon sources. The carbon sources mannitol (control) and sucrose were metabilized by all isolates evaluated. The profile of intrinsic resistance to antibiotics showed that the isolates were mostly resistant to streptomycin and ampicillin, but susceptible to kanamycin and chloranphenicol. High amylolytic activity of, at least, four isolates was also demonstrated, especially for isolated 47.3b, which showed the highest enzymatic index. These results indicate the metabolic versatility of the pigeonpea rhizobia, and indicates the isolate 47.3b to further studies regarding the amylase production and characterization.
Key words: strain selection, biological nitrogen fixation, biotechnology
INTRODUCTION
Rhizobia is a group of soil bacteria which are able to form root and/or stem nodules with legume plants. These bacteria exhibit a great taxonomic and phenotypical diversity and some groups are distributed worldwide. Studies evaluating the rhizobial diversity have shown that some of these bacteria belong to groups which were not formerly described as nodule forming bacteria, such as β-proteobacteria (16) and more recently γ-proteobacteria (23). In the same extent as taxonomic diversity, the rhizobia strains show a wide range of metabolic profiles with multiple antibiotic resistance, carbon sources metabolism, high temperature and salt tolerance (11,12, 17, 29).
The main technological application of rhizobia in Brazil is regarding the rhizobial inoculant production. The metabolic characterization of Brazilian rhizobial strains has shown a great versatility of their potential to other biotechnological applications than inoculant production, such as production of microbial biopolymers (2, 8, 9), and enzymes with industrial application (19, 20, 21).
The production of enzymes with biotechnological interests is an important issue on industrial microbiology and the amylase production reaches almost 25% of the total world enzyme market (25). The constant bioprospection of microorganisms able to produce amylase may result in microbial strains, which show higher amylase production than those, industrially available nowadays. Besides contributing to the selection of microorganisms to industrial production of amylase, the characterization of rhizobia regarding the amylolytic activity also must contribute to the understanding of the very diverse metabolic profiles of the root nodule bacteria (20, 21, 25). The aim of this study was to characterize rhizobial isolates from pigeonpea root nodules, by means of its phenotypic characteristics and evaluate the amylolytic capacity of the bacteria.
MATERIALS AND METHODS
Cultural characterization of rhizobial isolates
A total of 27 fast growing rhizobial isolates were evaluated in this study. These bacteria were isolated from root nodules of pigeonpea [Cajanus cajan (L.) Millsp.] genotypes cultivated in soils of Rio de Janeiro State, under different management practices (7). The isolates were inoculated at Petri dishes containing YMA (Yeast Extract-Manitol-Agar) (27) medium and incubated at 28°C for 2-3 days. The isolates were characterized according to the following features: pH alteration of culture medium, colony color, colony size, gum production and uniformity (7, 26, 28). As reference strains, it was also culturally characterized two strains belonging to the Rhizobium tropici (BR 322) end Ensifer saheli (BR 526) species. These bacteria were obtained from the culture collection of Embrapa Agrobiologia.
Intrinsic antibiotic resistance
The evaluation of intrinsic antibiotic resistance (IAR) of rhizobial isolates were carried out through the gradient plates method described by Bromfield et al. (4) and adapted by Xavier et al. (28). Petri dishes containing double YMA medium layers were prepared using low melting point agar. The bottom agar layer, supplemented with 500 mg.L-1 of kanamycin (kan), chloramphenicol (clo), ampicillin (amp) or streptomycin (str), was distributed and dishes solidified in a inclined surface around 30°. After basal layer solidification, the upper layer, without antibiotic supplement, was added. The antibiotic diffusion from bottom to upper layer gave rise to three sections of antibiotic concentration: zero to 167 mg.L-1 (first section), 168 to 333 mg.L-1 (second section) and 334 to 500 mg.L-1 (third section).
To inoculation, the bacterial isolates were cultured at the YM (Yeast extract-Mannitol) liquid medium (27) at orbital stirring (120 rpm) by 2-3 days. One mL of each culture broth was centrifuged (8000 g for five minutes), the supernatant was discarded and the pellet was resuspended in 1 mL of sterile saline solution (NaCl 0,85% w/v). To the inoculation, 10 µL of each cell suspension were dropped at the first section of Petri dishes that were inclined to drain the cell suspension towards the end of the third section. The dishes were incubated at 28°C until the proper growth time of each isolate. It was attributed score 1 to isolates which were able to grow only at the first section (low resistance); 2 to the isolates which were able to grow in first and second sections of the dish, but were not at the third section (intermediate resistance); 3 to the isolates that presented positive growth through every three zones (high resistance) (10, 28).
Salt and high temperature tolerance and carbon source
metabolism
The bacterial isolates were cultured in liquid YM medium and the cell suspension was prepared as described previously. A 10 µL aliquot of the cell suspension was inoculated at Petri dishes containing YMA medium and scratched with a platinum loop.
To the evaluation of salt tolerance, the bacterial isolates were inoculated at YMA medium supplemented with zero (control); 0,5; 1; 2 and 3% (w/v) of NaCl and the Petri dishes were incubated at 28°C. To evaluate the growth ability of the bacterial isolates subjected to different incubation temperatures, the bacteria were inoculated in Petri dishes containing original YMA medium and the incubated at 28 (control), 32, 37 or 42°C, in different growth chambers. The bacterial isolates were evaluated for their growth ability in basal YMA media prepared using 1% (w/v) of the following carbon sources: mannitol (control), glucose, xylose, maltose, fructose, sucrose, lactose and malic acid.
Amylolytic capacity
The bacterial isolates were cultivated in YM liquid medium and a cell suspension with saline solution was prepared as described above. Three aliquots with 10 µL of cell suspension were inoculated at equidistant points of each Petri dish containing YMA media with starch (1% w/v) as sole carbon source and without bromotimol blue (20, 21). The bacterial isolates were incubated at 28°C by 5 days. After the incubation, an iodine solution was abundantly distributed at medium surface and incubated at room temperature by 20 minutes. A clear zone surrounding the colonies indicated the amylolytic activity. The amylolytic activity was estimated by means of the "enzymatic index", estimated by the ratio between the diameter of the clear zone surrounding the colonies and the colony diameter (14). The enzymatic index for each replicate was calculated using the mean of the three colonies of each Petri dish.
Data analysis
The cultural characterization data were transformed in a binary matrix and the clustering carried out with the unweighted pair-group method with arithmetic mean (UPGMA) algorithm and Dice coefficient through the software PaSt (13). All the culture experiments were conducted with three replicates. In those cases which the evaluation profiles do not showed the same results at the three replications, the experiments were repeated to assure the results.
The amylolytic activity experiment was conducted at a completely random design. The enzymatic index data was subjected to a analysis of variance (ANOVA) and to Skott-Knott mean comparison range test (p<0,05) using the software Sisvar (Ferreira, D.F. Sisvar version 4.5. DEX/UFLA, 2003).
RESULTS AND DISCUSSION
The clustering based at the cultural characteristics showed two main clusters (Figure 1). The cluster A differs from B mainly by colony color. The cluster A, which encompasses both reference strains evaluated, presents colony with white color, while the cluster B encompasses isolates with yellow colonies. In the cluster A two main sub-clusters are observed. The cluster A1 segregates 16 isolates which acidified the culture medium, while the sub-cluster A2 has only 2 isolates that are able to alkalinize the culture media. At the sub-cluster A1 is also possible to observe other dichotomization, the upper group (9 isolates) had low mucus production and the bottom group (7 isolates) gathered bacteria that were able to produce high amount of mucus at solid culture media. The cluster B encompasses six isolates with yellow colonies. The sub division in that cluster was mainly due to the pH reaction on the culture media. The sub-cluster B1 has three isolates that did not change the pH of YMA culture medium while the sub-cluster B2 harbored isolates that acidify the medium.
The prevalence of bacterial isolates that decrease the pH of the culture media is very commonly found in association with tropical legumes (6, 7, 15). This characteristic, associated with the fast growing ability, probably is an adaptation to the tropical edapho-climatic conditions (5, 26). Recent studies have shown that some originally slow growing and alkalinizing rhizobia strains inoculated at tropical soils may reduce the generation time and acidify the medium as adaptive responses to their conditions (3).
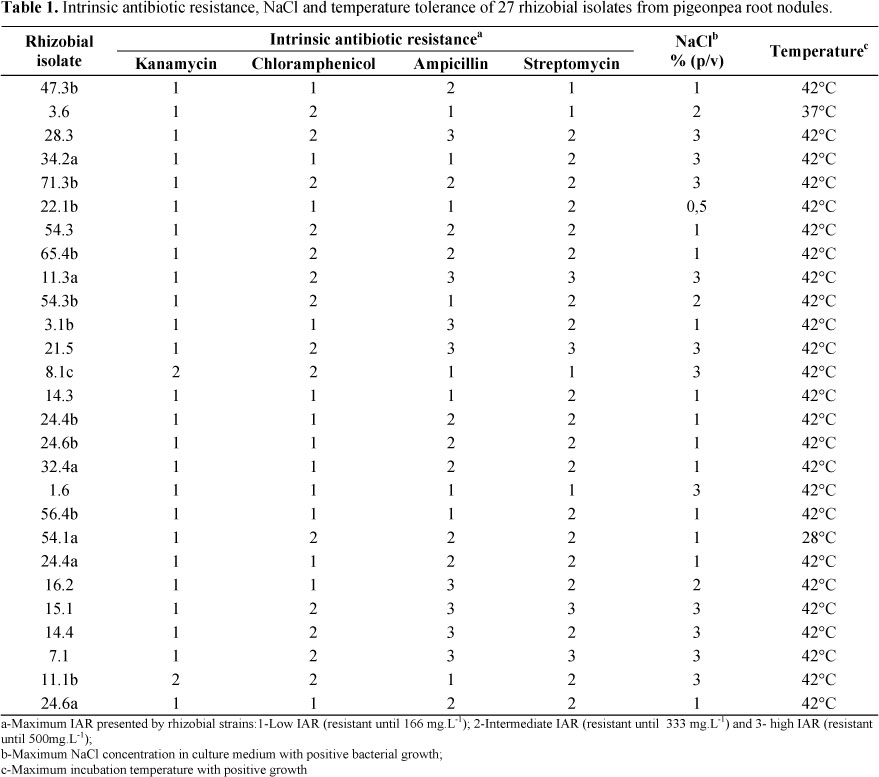
The evaluation of phenotypical characteristics demonstrated that the isolates evaluated in the present study, showed a great diversity of metabolic profiles. Regarding the IAR, eight (31%) presented the highest resistance level to ampicillin, 10 isolates (38%) presented intermediate IAR and other eight isolates presented low ampicillin resistance (Table 1). Four (16%) isolates showed high streptomycin resistance while 19 (73%) and four isolates showed intermediate and low streptomycin resistance, respectively. Among the isolates evaluated we did not found isolates with the high IAR level to chloramphenicol and kanamycin once only two (8%) isolates presented intermediate resistance and the majority of bacteria (92%) showed low resistance to kanamycin. The half part of rhizobial isolates presented intermediate resistance, while the other half only showed low chloramphenicol IAR.
Although some studies have shown that fast growing strains are not resistant to streptomycin (10), the IAR results found in the present study, corroborate to some recent results which pigeonpea rhizobial IAR also presented high resistance to streptomycin and low kanamycin resistance (6). It is very common to find fast growing rhizobial isolates that exhibiting high tolerance to streptomycin (6, 28). Regarding ampicillin IAR, the results obtained in the present study indicated that several rhizobia isolates presented high or intermediate resistance, which is different from previous results to pigeonpea rhizobia (1), pointing out the great diversity metabolic profiles of the bacteria isolated from this green manure. Other common characteristic found in fast growing rhizobia is the low resistance to kanamycin and chloramphenicol. High resistance to both antibiotics among the isolates evaluated was not observed. Intermediate resistance presented by 13 rhizobial isolates for chloramphenicol and in only two isolates for kanamycin. Similar results were already shown to rhizobia isolated from common bean (24), pigeonpea (6) and cowpea (28).
The evaluation of rhizobial tolerance to different salt concentrations at culture media and different incubation temperatures showed that a large amount of isolates evaluated (42%) was able to grow in YMA media supplemented with 3 % (w/v) of NaCl and the majority part of the isolates (92%) was able to grow when incubated at 42°C (Table 1). Temperatures higher than 40°C were easily reached at the tropical soils, the evaluation of in vitro tolerance to high temperatures may help in the selection of more stable strains to heat conditions (29). Regarding the salt tolerance evaluation, in vitro assays are cheap and feasible to large numbers of isolates, allowing the selection of potential strains to further field experiments, especially in soils with high salt content (17, 29).
Evaluating the rhizobial isolates ability to metabolize different carbon sources, we observed that the isolates 34.2a, 71.3b and 11.1b were able to grow in culture medium supplemented with all carbon sources evaluated (Table 2). All isolates were able to grow at the control treatment (original YMA medium), and also using sucrose as sole carbon source. On the other hand, few (19%) of the rhizobial isolates were able to metabolize malic acid, the majority of them (54%) was able to grow in the culture media supplemented with at least six carbon sources evaluated. A total of 92, 89, 89, 85 and 80% of rhizobial isolates presented positive growth using fructose, xylose, glucose, lactose and maltose respectively. Five isolates (19%) metabolized five carbon sources and four (15%) isolates presented positive growth in YMA medium supplemented with four carbon sources. Only one isolate grew in culture media with only three sugars as sole carbon source.
The ability of rhizobia in metabolize a large range of carbon sources, associated with other physiological characteristics such as salt and drought tolerances, indicates that there is a great saprophytic capacity of these bacteria, a relevant information on rhizobial ecology (22). The poor growth of the rhizobial isolates using the malic acid as sole carbon source was expected. As we observed in the present study, Anand and Dogra (1) showed that few fast growing rhizobia from pigeonpea were able to metabolize malic acid, comparing with other carbon sources as mannitol, xylose, sucrose among others.
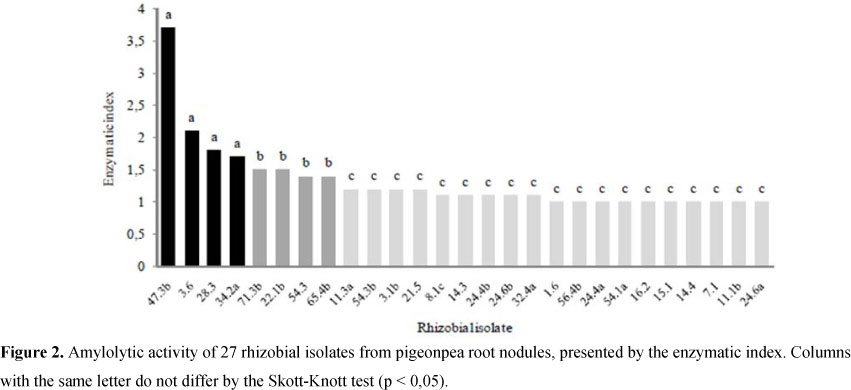
At the Figure 2 is possible to observe that four isolates stand out regarding the amylolytic capacity, four other isolates present an intermediate and 18 of then showed low amylolytic activity (in spite of being able to grow at the culture medium supplemented with starch). Qualitative evaluations of amylolytic capacity by assays using iodine solution staining is a cheap and feasible approach to select potential strains to amylase isolation and characterization (25, 26). This exploratory assay of rhizobial bacteria revealed a very promising isolate; the isolate 47.3b, which reached the highest rate of starch hydrolysis surrounding the colony (Figure 3).
The main bacteria used industrially to the production of amylases belongs to Bacillusgenus (25), but nodule forming bacteria has already demonstrated their potential for this purpose, being able to hydrolyze starch in culture medium supplemented with several starchy compounds (27). By the way, some strains evaluated, in the present study, present the capacity to produce some biopolymers industrially applied (EPS and PHB), using starch as sole carbon source (8, 9). This result shows the potential to multiple biotechnological applications of rhizobia.
The results found in this study confirm the great variability on phenotypical characteristics among the rhizobial isolates and indicates the potential of some strains to amylolytic activity. The isolate 47.3b, for example, showed the potential to further inclusion in programs of optimization of the culture conditions to produce amylases. The ability of rhizobial strains to grow using inexpensive carbon sources as substrates is desirable. In the present study, it was possible to evaluate that several bacterial strains were able to grow using inexpensive substrates as sole carbon source, such as starch and sucrose. Their ability to metabolize these substrates increases their biotechnological potential, reducing the costs of cultivation and opening new possibilities to rhizobial applications.
ACKNOWLEDGEMENTS
To CPGA-CS (UFRRJ), CAPES, Embrapa and CNPq for financial support.
Submitted: March 15, 2011
Approved: June 07, 2012
- 1. Anand, R.C; Dogra, R.C. (1991) Physiological and biochemical characteristics of fast- and slow-growing Rhizobium sp. pigeon pea (Cajanus cajan). J Appl Bacteriol 70:197-202
- 2. Aranda-Selverio, G.; Penna, A.L.B.; Campos-Sás, L.F.; Santos Júnior, O.; Vasconcelos, A.F.D.; Silva, M.L.C.; Lemos, E.G.M.; Capanharo, J.C.; Silveira, J.L.M.S. (2010) Propriedades reológicas e efeito da adição de sal na viscosidade de exopolissacarídeos produzidos por bactérias do gênero Rhizobium Quím. Nova. 33: 895-899.
- 3. Batista, J.S.S.; Hungria, M.; Barcellos, F.G.; Ferreira, M.C.; Mendes, I.C. (2007). Variability in Bradyrhizobium japonicum and B. elkanii seven years after introduction of both the exotic microsymbiont and the soybean host in a Cerrados soil. Microb. Ecol. 53: 270-284.
- 4. Bromfield, E.S.P.; Stein, M.; White, R.P. (1982) Identification of Rhizobium on antibiotic concentration gradient. Ann. Appl. Biol. 101: 269-277.
- 5. Bushby, H.V.A.; Marshall, K.C. (1977) Some factors affecting the survival of root nodule bacteria on desiccation. Soil Biol. Biochem. 9: 143-147.
- 6. Costa, F.M.; Fernandes Júnior, P.I. ; Leite, J. ; Xavier, G.R.; Schiavo, J.A. (2010). Resistência intrínseca a antibióticos e tolerância a altas temperaturas por rizóbios isolados de guandu [Cajanus cajan (L.) Millsp.]. In: FERTBIO, 2010. Guarapari, ES. CD-Rom.
- 7. Fernandes Júnior, P.I. (2009). Caracterização fenotípica e produção de biopolímeros por bactérias isoladas de nódulos de guandu [Cajanus cajan (L.) Millsp.]. Seropédica, Brasil, 186 p. (D.Sc. Thesis. Departamento de Solos. Instituto de Agronomia UFRRJ).
- 8. Fernandes Júnior, P.I.; Oliveira, P.J.; Rumjanek, N.G.; Xavier, G.R. (2011). Poly-B-hydroxybutyrate and exopolysaccharide biosynthesis by bacterial isolates from pigeonpea [Cajanus cajan (L.) Millsp] root nodules. Appl. Biochem. Biotech. 163. 473-484.
- 9. Fernandes Júnior; P.I.; Almeida, J.P.S.; Passos, S.R.; Oliveira, P.J.; Rumjanek, N.G.; Xavier, G.R. (2010) Produção e comportamento reológico de exopolissacarídeos sintetizados por rizóbios de isolados de guandu. Pesq. Agropec. Bras. 45. 1465-1471.
- 10. Fernandes, M.F.; Fernandes, R.P.M.; Hungria, M. (2003). Seleção de rizóbios nativos para guandu, caupi e feijão-de-porco nos tabuleiros costeiros de Sergipe. Pesq. Agropec. Bras. 38: 835-842.
- 11. Florentino, L.A.; Sousa, P.M.; Silva, J.S.; Silva, K.B.; Moreira, F.M.S. (2010). Diversity and efficiency of Bradyrhizobium strains isolated from soil samples collected from around Sesbania virgata roots using cowpea as trap species. R. Bras. Ci. Solo. 34: 1113-1123.
- 12. Garcia-Fraile, P.; Rivas, R.; Willems, A.; Peix, A.; Martens, M.; Martinez-Molina, E.; Mateos, P.F.; Velazquez, E. (2007). Rhizobium cellulosilyticum sp nov, isolated from sawdust of Populus alba Int. J. Syst. Evol. Microbiol. 57: 844-848.
- 13. Hammer, Ø., Harper, D.A.T., Ryan, P.D. (2001). PAST: Paleontological Statistics Software Package for Education and Data Analysis. Palaeontol. Electron. 4. http://palaeo-electronica.org/2001_1/past/issue1_01.htm
- 14. Hankin, L.; Anagnostakis, S.L. (1975) The use of solid media for detection of enzyme production by fungi, Mycol. 16: 597-607.
- 15. Leite, J.; Seido, S.L.; Passos, S.R.; Xavier, G.R.; Rumjanek, N.G.; Martins, L.M.V. (2009). Biodiversity of rhizobia associated with cowpea cultivars in soil of the lower half of the São Francisco River Valley. Rev. Bras. Ci. Solo. 33: 1215-1226.
- 16. Moulin, L.; Munive, A.; Dreyfus, B.; Boivin-Masson, C. (2001) Nodulation of legumes by members of the B-subclass of Proteobacteria. Nature. 411: 948-950.
- 17. Nóbrega, R.S.A.; Motta, J.S.; Lacerda, A.M.; Moreira, F.M.S. (2004). Tolerância de bactérias diazotróficas simbióticas à salinidade in vitro Ci. Agrotec. 28: 899-905.
- 18. Odee, D.W.; Sutherland, J.M.; Makatiani, E.T.; McInroy, S.G.; Sprent, J.I. (1997) Phenotypic characteristics and composition of rhizobia associated with woody legumes growing in diverse Kenyan conditions. Plant Soil. 188, 65-75.
- 19. Oliveira, A.N.; Oliveira, L.A. Souza, J. (2010) Partial characterization of amylases of two indigenous Central Amazonian rhizobia strains. Braz. Arch. Biol. Technol. 53: 35-45.
- 20. Oliveira, A.N.; Oliveira, L.A.; Andrade, J. S.; Chagas Júnior, A.F. (2007). Produção de amilase por rizóbios, usando farinha de pupunha como substrato. Ci. Tecnol Alim. v. 27: 61-66.
- 21. Oliveira, A.N.; Oliveira, L.A.; Andrade, J.S.; Chagas Júnior, A.F. (2007). Rhizobia amylase production using various starchy substances as carbon substrates. Braz. J. Microbiol. 38: 208-216.
- 22. Rynne, F.G.; Glenn, A.R.; Dilworth, M.J. (1994). Effect of mutations in aromatic catabolism on the persistence and competitiveness of Rhizobium legvminosarum bv. trifolii. Soil Biol. Biochem. 26: 703-710.
- 23. Shiraishi, A.; Matsushita, N.; Hougetsu, T. (2010). Nodulation in black locust by the gammaproteobacteria Pseudomonas sp. and the betaproteobacteria Burkholderia sp. Syst. Appl. Microbiol. 33: 269-274.
- 24. Souza, M.F.M.; Vale, H.M.M.; Straliotto, R. (2003) Competitividade de estirpes pertencentes a diferentes espécies de rizóbio para ocupação nodular em feijoeiro (Phaseolus vulgaris L.). Rev. Agron. 37:59-63.
- 25. Souza, P.M.; Magalhães, P.O. (2010) Application of microbial a-amylase in industry - A review. Braz. J. Microbiol. 41:850-861.
- 26. Teixeira, F.C.P.; Borges, W. L.; Xavier, G.R.; Rumjanek, N.G. (2010) Characterization of indigenous rhizobia from Caatinga. Braz. J. Microbiol. 41: 201-208.
- 27. Vincent, J.M. (1970). A manual for the practical study of root - nodules bacteria Blackwell Science Publication, Oxford, UK.
- 28. Xavier, G.R.; Martins, L.M.V.; Neves, M.C.P.; Rumjanek, N.G. (1998). Edaphic factors as determinants for the distribution of intrinsic antibiotic resistance in a cowpea, rhizobia population. Biol. Fertil. Soil. 27: 386-392.
- 29. Xavier, G.R.; Martins, L.M.V.; Rumjanek, N.G.; Neves, M.C.P. (2007) Tolerância de rizóbio de feijão-caupi à salinidade e à temperatura em condição in vitro. Caatinga. 20: 1-9. 2007.
Publication Dates
-
Publication in this collection
19 Feb 2013 -
Date of issue
Dec 2012
History
-
Received
15 Mar 2011 -
Accepted
07 June 2012