Abstract
Manganese peroxidase (MnP) was produced from white rot edible mushroom Pleurotus ostreatus on the culture filtrate. The enzyme was purified to homogeneity using (NH4)2SO4 precipitation, DEAE-Sepharose and Sephadex G-100 column chromatography. The final enzyme activity achieved 81UmL-1, specific activity 78 U mg-1 with purification fold of 130 and recovery 1.2% of the crude enzyme. SDS-PAGE indicated that the pure enzyme have a molecular mass of approximately 42 kDa. The optimum pH was between 4-5 and the optimum temperature was 25 ºC. The pure MnP activity was enhanced by Mn2+,Cu2+,Ca2+ and K+ and inhibited by Hg+2 and Cd+2.H2O2 at 5 mM enhanced MnP activity while at 10 mM inhibited it significantly. The MnP-cDNA encoding gene was sequenced and determined (GenBank accession no. AB698450.1). The MnP-cDNA was found to consist of 497 bp in an Open Reading Frame (ORF) encoding 165 amino acids. MnP from P. ostreatus could detoxify aflatoxin B1 (AFB1) depending on enzyme concentration and incubation period. The highest detoxification power (90%) was observed after 48 h incubation at 1.5 U mL-1 enzyme activities.
manganese peroxidase; aflatoxin B1; Pleurotus ostreatus; purification
Aflatoxin detoxification by manganese peroxidase purified from Pleurotus ostreatus
Ramy Sayed Yehia
Department of Botany, Faculty of Science, Cairo University, Giza, Egypt
Send correspondence to Send correspondence to R.S. Yehia Department of Botany, Faculty of Science, Cairo University 12613 Giza, Egypt E-mail: drramy4@hotmail.com
ABSTRACT
Manganese peroxidase (MnP) was produced from white rot edible mushroom Pleurotus ostreatus on the culture filtrate. The enzyme was purified to homogeneity using (NH4)2SO4 precipitation, DEAE-Sepharose and Sephadex G-100 column chromatography. The final enzyme activity achieved 81UmL-1, specific activity 78 U mg-1 with purification fold of 130 and recovery 1.2% of the crude enzyme. SDS-PAGE indicated that the pure enzyme have a molecular mass of approximately 42 kDa. The optimum pH was between 4-5 and the optimum temperature was 25 ºC. The pure MnP activity was enhanced by Mn2+,Cu2+,Ca2+ and K+ and inhibited by Hg+2 and Cd+2.H2O2 at 5 mM enhanced MnP activity while at 10 mM inhibited it significantly. The MnP-cDNA encoding gene was sequenced and determined (GenBank accession no. AB698450.1). The MnP-cDNA was found to consist of 497 bp in an Open Reading Frame (ORF) encoding 165 amino acids. MnP from P. ostreatus could detoxify aflatoxin B1 (AFB1) depending on enzyme concentration and incubation period. The highest detoxification power (90%) was observed after 48 h incubation at 1.5 U mL-1 enzyme activities.
Key words: manganese peroxidase, aflatoxin B1, Pleurotus ostreatus, purification.
Introduction
Manganese peroxidase (MnP) (Ec 1.11.1.13) catalyses the H2O2 dependent oxidation of Mn II to Mn III which is a non-specific oxidant and seems to play an important role in initiating lignin attack (Kuwahara et al., 1984; Paszcynski et al., 1986; Perez and Jeffries, 1992). The enzyme is an extracellular heme protein produced by wood rotting and little degrading basidomycetes. It is small enough to diffuse into the compact lignocelluloses complex, and oxidizes the aromatic structures via formation of radicals, leading finally to the breakdown of covalent bonds (Evans et al., 1994). The purified MnP differentiated by different column chromatography into two isoenzymes, MnP1 and MnP2 with very close molecular masses of 42 kDa and 44 kDa, respectively (Schneegaβ et al., 1997). The main and more active isoenzyme was MnP2 from Nematoloma frowardii (Schneegaβ et al., 1997), Lentinula edodes (Boer et al., 2006), Pleurotus eryngii (Martinez et al., 1996) and Phanerochate flavido-alba (de la Rubia et al., 2002).
The mushroom genus Pleurotus has the capacity to degrade lignin with degradation of cellulose (Kamra and Zadrazil, 1986); this phenomenon is of relevant biotechnological importance in delignification process for feed production and paper-pulp industry (Kamra and Zadrazil, 1986; Martinez et al., 1994). Appreciable levels of peroxidases involved in lignin biodegradation by Pleurotus species such as lignin peroxidase (LP) and MnP (Kirk and Farrell, 1987). It has been reported that MnP can oxidize Mn+2 to Mn+3 which act as a better substrate than veratryle alcohol at physiological pH 4.5. It was suggested also that the generation of chelated Mn+3 could be the main role of MnP in lignin degradation (Khindaria et al., 1995).
The increasing interest in this enzyme is due to its production by most white rot fungi including lignin degrading species that lack LP (Lobos et al., 1994; Périé and Gold, 1991), its capacity to depolymerise synthetic lignin in vitro (Wariishi et al., 1991), its oxidation to phenolic and nonphenolic compounds (Bao et al., 1994), its biodegradation of polycyclic aromatic hydrocarbon (PAH) (Pickard et al., 1999; Wariishi et al., 1992), humic acid (Ziegenhagen and Hofrichter, 1998), synthetic dyes (Heinfling et al., 1998) and more recently detoxification of mycotoxins (Motomura et al., 2003; Wang et al., 2011).
Aflatoxin B1 (AFB1) is a potent mycotoxin with mutagenic, carcinogenic, hepatogenic and immunosuppressive properties. AFB1 treated with MnP from the white rot fungus Phanerochaete sordida Yk-624 was detoxified by 86% of the original value after 48 h (Wang et al., 2011). Nuclear Magnetic Resonance (1H-NMR) and High Resolution Electrospray Mass Spectrometry (HR-ESI-MS) analysis suggested that AFB1 is first oxidized to AFB1-8, 9-epoxide by MnP and then hydrolyzed to AFB1-8, 9-dihydrodiol (Wang et al., 2011).
In the present study the detoxification of AFB1 by purified MnP extracted from the edible mushroom Pleurotus ostreatus was demonstrated. The active enzyme was characterized biochemically and in molecular basis.
Materials and Methods
Organism and culture condition
Pleurotus ostreatus was kindly supplied by Academy of Science and Technology, Alexandria, Egypt. It maintained on Potato Dextrose Agar media (PDA) at 4 ºC. For the production of MnP, the fungal strain was grown in a medium (Grgic et al., 2001) containing (g L-1) glucose (10.0), KH2PO4 (0.2), CaCl2 (0.11), (NH4)2HPO4 (0.264), MgSO4.7H2O (0.05), ZnSO4.7H2O (0.0425), MnSO4.H2O (0.175), CoCl2.6H2O (0.007), CuCl2.2H2O (0.007), FeCl3.6H2O (0.0009), NaCl (0.0009), yeast extract (0.2), veratryl alcohol (0.07), tartaric acid (3.0) and 1 g of Tween 80 (pH was adjusted at 4.5). Sterilized flasks containing 100 mL each of liquid medium were inoculated with 5.0 inoculums of 0.5 mm diameter. The flasks were incubated under stationary condition at 28 ºC. The cultures were filtered and the filtrates considered as crude MnP.
MnP activity assay
The activity of MnP was determined spectrophotometrically (Gold and Glenn, 1988) by monitoring the absorbance change at λ = 240 nm due to the formation of Mn+3 lactate and using the molar extinction coefficient value of 65.00 M-1cm-1 . The reaction mixture 1 mL consisted of 50 µM MnSO4,50 µM H2O2 and 0.5 mL of the enzyme solution in 50 mM sodium lactate/lactic acid buffer pH 4.5 at 30 ºC. One enzyme activity unit was defined as enzyme transformed 1 µmole of the Mn+2 into the Mn+3 under the specified assay condition.
Enzyme purification
All procedures were performed at 4 ºC. The culture filtrate containing crude MnP was centrifuged at 15,000 g for 30 min to remove particulate material then resuspended in bi-distilled water. One liter of culture supernatant was precipitated with solid ammonium sulfate to 80% saturation under constant stirring. The solution was centrifuged at 20000 g for 30 min and the precipitates were dissolved in 50 mM sodium acetate buffer (pH 5.0) followed by overnight dialysis against buffer A (50 mM sodium acetate, pH 5.0, supplemented with 0.1 mM phenylmethanesulfonyl fluoride (PMSF). The dialyzed sample was applied to a DEAE-Sepharose column (Pharmacia Biotech, Uppsala, Sweden) pre-equilibrated with buffer A. The proteins were eluted with buffer A containing linear gradient of 0.0-0.6 M NaCl. The soluble proteins were applied to Sephadex G-100 column (80 cm x 2.0 cm) previously equilibrated with the same buffer A. The protein fractions (2.0 mL) were eluted at a flow rate of 1 mL min-1. Each fraction was assayed for protein (A280 nm) and MnP activity. The pooled active fractions were dialyzed against water, lyophilized and stored at -20 ºC.
Protein determination
Total protein was determined by using bovine serum albumin as protein standard (Bradford, 1976).
Sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE)
SDS-PAGE was performed in 12.5% polyacrylamide gels according to the method of Laemmli (1970). The separated proteins were stained with Coomassie Brilliant Blue R-250 (Fluka, Switzerland), and their molecular weights were determined by comparison with low range molecular weight markers (Bio-Rad) was used.
Enzyme characterization
The effects of temperature, pH and metals on MnP activity were tested successively. The optimum temperature for enzyme activity was determined in the range of 20 ºC to 45 ºC at 5 ºC intervals. The optimum pH was tested in the range of 4.0 to 10.0 using 50 mM sodium acetate, sodium phosphate or glycine sodium hydroxide buffers. The effects of Ca+2, Ni+2, Cu+2, Hg+2, Mn+2, Cd+2, K+ and H2O2 on MnP activity were also assayed.
Assay of AFB1 detoxification by MnP
Reaction mixture containing 500 µL of the finally purified MnP, 10 mL of 1 mM AFB1 in 10% dimethylsulfoxide, 1 mM MnSO4, 0.1% Tween 80, 200 µL glucose oxidase, and 2.5 mM glucose in 50 mM sodium lactate /lactic acid buffer, pH 4.5. Reactions were performed for 48 h at 30 ºC and mixing at 150 r.p.m. The residual amount of AFB1 was determined by HPLC (Shimadzu) under the following conditions: column, RP18 (ODS) (4.6 mm x 150 mm, Wako Pure Chemical Industries); mobile phase, 40% aqueous methanol; flow rate, 0.5 mL min-1; and detection wavelength, 365 nm.
Molecular characterization of MnP
Isolation of RNA was carried out according to the method outlined by Huang et al. (2007) by using Biozol reagent (BioFlux, Tokyo, Japan), and then purified using tri-reagent RNA kit (Sigma). cDNA synthesis was carried out by using 1st strand cDNA kit (BIO BASIC INC.) according to the manufacturer instructions, in a reaction mixture (20 µL final volume). Specific products were amplified by PCR, (using T-GRADIENT thermal cycler from Biometra) in 25 µL volume containing 2.5 µL of cDNA; 25 pmol of each primer forward primer AAT ACCAGGGTGAAGTTGAATCG and reverse AAT GCGTCTTGGGAACAGGAATA; 10 mM of each dNTP; 1U of Taq DNA polymerase; 10 mM Tris-Hcl, pH 9.0; 50 mM KCl; 1 mM MgCl2. Conditions for PCR were, denaturation at 94 ºC for 3 min was followed by 35 cycles of 1 min at 94 ºC; 2 min at 60 ºC and 1 min at 72 ºC with a final extension step at 72 ºC for 7 min. Two oligonucleotide primers were synthesized according to the published sequence. The amplified fragment was separated, purified and sequenced.
Results and Discussion
It was found that when the dialyzed protein solution, obtained by ammonium sulphate precipitation of the culture supernatant was loaded on DEAE-Sepharose column and three major peaks with 6.8 purification fold and 20% recovery were detected (Table 1 and Figure 1A). As the eluted peaks of high MnP activity were subsequently submitted to further purification in Sephadex G-100, single activity peak was observed with final enzyme activity of 81 U mL-1, specific activity of 78 U mg-1 protein, purification fold 130 and recovery 1.2% of the crude preparation (Figure 1B).
The apparent molecular mass of the purified MnP enzyme was estimated to be 42 kDa by SDS-PAGE (Figure 2). Similar molecular mass to that of P. ostreatus MnP was recorded in Phlebia radiate (47-48 kDa), Ridigo lignosus (42 kDa) (Hatakka, 1994) and Phanerochaete Flavido-alba (45 kDa) (de la Rubia et al., 2002).
The effects of pHs on purified MnP enzyme activity showed a pH optima in the range of 4.0 to 5.0 (Figure 3A). The enzyme activity then attenuated as the pH increased. This optimum pH is lower than the pH optima reported for MnP of P. chrysosporium (Kuan et al., 1993) and comparable to MnP obtained from Lentinula edodes range (4.0-6.0) (Boer et al., 2006). In this study the optimum temperature was found to be 25 ºC (Figure 3B) which is near to value of 28 ºC for MnP of P. chrysosporium and similar to that of Fomes durissimus (Singh et al., 2011) whereas it is much lower than that obtained from Lentinula edodes (45 ºC) (Boer et al., 2006).
The purified enzyme was inhibited by Cd2+ and Hg2+ whereas it enhanced by Mn2+ > K+ > Cu2+ > Ca2+ > Ni2+ (Table 2). MnP was also resistant to presence of up to 5 mM H2O2, but it was inhibited in the presence of 10 mM H2O2. In terms of industrial applications, the stability of MnP in the presence of high concentration of H2O2 is very important property. MnP requires H2O2 for its activity. However, H2O2 could also inactivate MnP quickly when it was applied in high concentrations (Bermek et al., 2002; Wariishi et al., 1988).
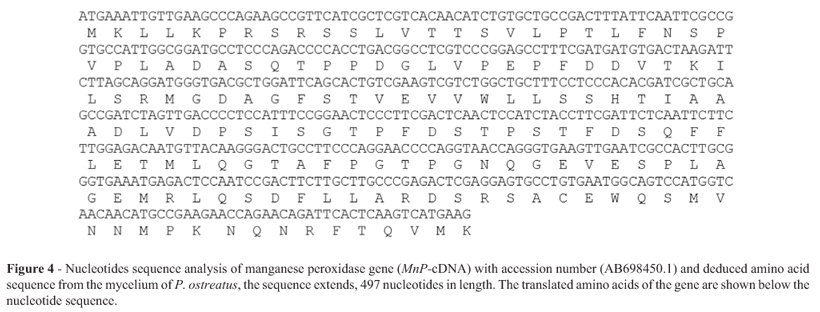
The isolated MnP enzyme gene from P. ostreatus mycelia has been sequenced. The nucleotide sequence of the gene revealed a single open reading frame (ORF) of 497 bp encoding 165 amino acids (Figure 4). These sequence date has been submitted to the GenBank under accession number AB698450.1. The MnP-cDNA gene (AB698450.1) exhibited high sequence homology (100%) with manganese peroxidase isolated from P. ostreatus isolate 5145 (GenBank Accession No. JN020144.1), (97%) of (GenBank Accession No. AJ243977.1), (95%) of Mnperoxidase isolated from P. pulmonarius (GenBank Accession No. AY916529.1). The deduced amino acid sequence showed high homology (97%) with P. ostreatus isolate 5145 (GenBank Accession No. AER35423.1). This enzyme gene belong to a group of peroxidases containing a heme prosthetic group (ferriprotoporphyrin IX), (plant heme-dependent peroxidase superfamily) which catalyzes a multistep oxidative reaction involving hydrogen peroxide as the electron acceptor. The plant peroxidase-like superfamily is found in all three kingdoms of life and carries out a variety of biosynthetic and degradative fractions (Marchler-Bauer et al., 2011).
The detoxification potentiality of MnP to aflatoxin B1 showed that after 24 h the level of AFB1 was reduced to 67% in the presence of 1.5 U mL-1 enzyme while the maximum detoxification (90%) was attained after 48 h at enzyme concentration of 1.5 U mL-1 (Table 3). MnP from Phanerochaete sordida Yk-624 removed approximately 70% of AFB1 after 24 h and reach to complete detoxification by multitreatment with MnP (Wang et al., 2011). Culture supernatant of Myxococcus fulvus was able to degrade AFB1, AFG1 and AFM1 effectively in solution. Significant reduction of AFB1 (71.89%), AFG1 (68.13%) and AFM1 (63.82%) were observed after 48 h treatment with the culture supernatant from strain ANSM068 (Zhao et al., 2010). These findings are in agreement with those obtained by Alberts et al. (2006) who reported that AFB1 was effectively degraded by extracellular extracts from Rhodococcus erythropolis liquid cultures, with only 33.2% residual AFB1 after 72 h. These results also went parallel with those obtained by Lillehoj et al. (1971) who reported that AFM1 (8 µgmL-1) is completely removed from the liquid medium of Flavobacterium aurantiacum by incubation with 5 x 1010 resting cells per milliliter for 4 h.
Acknowledgement
The author is grateful to Dr. Tahany M.A. Abd El-Rahman, professor of microbiology, Department of Botany, Faculty of Science, Cairo University, for her continuous encouragement and for revising the manuscript.
Submitted: March 30, 2012; Approved: April 1, 2013.
All the content of the journal, except where otherwise noted, is licensed under a Creative Commons License CC BY-NC.
- Alberts JF, Engelbrecht Y, Steyn PS, Holzapfel WH, Van Zyl WH (2006) Biological degradation of aflatoxin B1 by Rhodococcus erythropolis cultures. Int J Food Microbiol 109:121-126.
- Bao WL, Fukushima Y, Jensen KA, Moen MA, Hammel KE (1994) Oxidative degradation of non-phenolic lignin during lipid peroxidation by fungal manganese peroxidase. FEBS Lett 354:297-300.
- Bermek H, Li K, Eriksson K-EL (2002) Studies on inactivation and stabilization of manganese peroxidase from Trametes versicolor In: Biotechnology in the pulp and paper industry 8th ICBPPI Meeting. Progress Biotechnol, Elsevier Science 21:141-149.
- Boer CG, Obici L, de Souza CGM, Peralta RM (2006) Purification and some properties of Mn peroxidase from Lentinula edodes Proc Biochem 41:1203-1207.
- Bradford MA (1976) Rapid and sensitive method for quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248-254.
- de la Rubia T, Linares A, Pérez J, Muñoz-Dorado J, Romera J, Martínez J (2002) Characterization of manganesedependent peroxidase isoenzymes from the ligninolytic fungus Phanerochaete flavido-alba Res Microbiol 153:547-554.
- Evans CS, Dutton MV, Guillen F, Veness RG (1994) Enzymes and small molecular mass agents involved with lignocellulose degradation. FEMS Microbiol Rev 13:235-240.
- Gold MH, Glenn JK (1988) Manganese peroxidase from Phanerochaete chrysosporium Meth Enzymol 161:258-264.
- Grgic I, Podgornik H, Berovic M, Perdih A (2001) Improvements in the determination of Manganese peroxidase activity. Biotechnol Lett 23:1039-1042.
- Hatakka A (1994) Lignin-modifying enzymes from selected white-rot fungi production and role in lignin degradation. FEMS Microbiol Rev 13:125-135.
- Heinfling A, Martinez MJ, Martinez AT, Bergbauer M, Szewzyk U (1998) Purification and characterization of peroxidases from the dye-decolorizing fungus Bjerkandera adusta FEMS Microbiol Lett 165:43-50.
- Huang Y, Peng K, Su J, Huang Y, Xu Y, Wang S (2007) Different affects of homocysteine and oxidized low density lipoprotein on methylation status in the promoter region of the estrogen receptor a gene. Acta Biochem Biophys Sin 39:19-26.
- Kamra DN, Zadrazil F (1986) Influence of gaseous phase, light and substrate pretreatment on fruit body formation, lignin degradation and in vitro digestibility of wheat straw fermented with Pleurotus spp. Agric Wastes 18:1-17.
- Khindaria A, Barr DP, Aust SD (1995) Lignin peroxidases can also oxidize manganese. Biochemistry 34:7773-7779.
- Kirk TK, Farrell RL (1987) Enzymatic combustion: the microbial degradation of lignin. Annu Rev Microbiol 41:465-505.
- Kuan IC, Johnson KA, Tien M (1993) Kinetic analysis of Manganese Peroxidase: The reaction with Manganese complexes. J Biol Chem 268:20064-20070.
- Kuwahara M, Glenn JK, Morgan MA, Gold MH (1984) Separation and characterization of two extracellular hydrogen peroxide-dependent oxidases from ligninolytic cultures of Phanerochaete chrysosporium FEBS Lett 169:247-250.
- Laemmli UK (1970). Cleavage of structural proteins during the assembly of head of bacteriophage T4. Nature 227:680-685.
- Lillehoj EB, Stubblefield RD, Shannon GM, Shotwell OL (1971) Aflatoxin M1 removal from aqueous solutions by Flavobacterium aurantiacum Mycopathol Mycol Appl 45:259-266.
- Lobos S, Larrain J, Salas L, Cullen D, Vicufia R (1994) Isoenzymes of manganese dependent peroxidase and laccase produced by the lignin-degrading basidiomycete Ceriporiopsis subvermispora Microbiology-UK 140:2691-2698.
- Marchler-Bauer A, Lu S, Anderson JB, Chitsaz F, Derbyshire MK, DeWeese-Scott C, Fong JH, Geer LY, Geer RC, Gonzales NR, Gwadz M, Hurwitz DI, Jackson JD, Ke Z, Lanczycki CJ, Lu F, Marchler GH, Mullokandov M, Omelchenko MV, Robertson CL, Song JS, Thanki N, Yamashita RA, Zhang D, Zhang N, Zheng C, Bryant SH (2011) CDD: a Conserved Domain Database for the functional annotation of proteins. Nucleic Acids Res 37:225-229.
- Martinez AT, Camarero S, GuillCn F, Gutikrrez A, MuRoz C, Varela E, Martinez MJ, Barrasa JM, Ruel K, Pelayo M (1994) Progress in biopulping of non-woody materials: chemical, enzymatic and ultra-structural aspects of wheat-straw delignification with ligninolytic fungi from the genus Pleurotus FEMS Microbiol Rev 13:265-274.
- Martinez MJ, Ruiz-duenas FJ, Cuillen F, Martinez AT (1996) Purification and catalytic properties of two manganese peroxidase isoenzymes from Pleurotus eryngii Eur J Biochem 237:424-432.
- Motomura M, Toyomasu T, Mizuno K, Shinozawa T (2003) Purification and characterization of an aflatoxin degradation enzyme from Pleurotus ostreatus Microbiol Res 158:237-242.
- Paszcynski A, Huynh VB, Crawford RL (1986) Comparison of ligninase-I and peroxidase-M2 from the white-rot fungus Phanerochaete chrysosporium Arch Biochem Biophys 244:750-756.
- Perez J, Jeffries TW (1992) Roles of manganese and organic acid chelators in regulating lignin degradation and biosynthesis of peroxidases by Phanerochaete chrysosporium Appl Environ Microbiol 58:2402-2409.
- Périé FH, Gold MH (1991) Manganese regulation of manganese peroxidase expression and lignin degradation by the white rot fungus Dichomitus squalens Appl Environ Microbiol 57:2240-2245.
- Pickard MA, Roman R, Tinoco R, Vazquez-Duhalt R (1999) Polycyclic aromatic hydrocarbon metabolism by white rot fungi and oxidation by Coriolopsis gallica UAMH 8260 laccase. Appl Environ Microbiol 65:3805-3809.
- Schneegaβ I, Hofrichter M, Scheibner K, Fritsche W (1997) Purification of the main manganese peroxidase isoenzyme MnP2 from the white-rot fungus Nematoloma frowardii b19. Appl Microbiol Biotechnol 48:602-605.
- Singh SK, Yadav M, Yadava S, Yadav KDS (2011) Coal depolymerising activity and haloperoxidase activity of Mnperoxidase from Fomes durissimus MTCC-1173. Bioinorg Chem Appl 2011:1-8.
- Wang J, Ogata M, Hirai H, Kawagishi H (2011) Detoxification of aflatoxin B1 by manganese peroxidase from the white-rot fungus Phanerochaete sordidaYK-624. FEMS Microbiol Lett 314:164-169.
- Wariishi H, Akileswaran L, Gold MH (1988) Manganese peroxidase from Phanerochaete chrysosporium: spectral characterization of the oxidized states and the catalytic cycle. Biochemistry 27:5365-5370.
- Wariishi H, Valli K, Gold MH (1991) In vitro depolymerization of lignin by manganese peroxidase of Phanerochaete chrysosporium Biochem Biophys Res Commun 176:269-275.
- Wariishi H, Valli K, Gold MH (1992) Manganese (II) oxidation by manganese peroxidase from the basidiomycete Phanerochaete chrysosporium J Biol Chem 267:23688-23695.
- Zhao LH, Guan S, Gao X, Ma QG, Lei YP, Bai XM, Ji C (2010) Preparation, purification and characteristics of an aflatoxin degradation enzyme from Myxococcus fulvus ANSM068. J Appl Microbiol 110:47-155.
- Ziegenhagen D, Hofrichter M (1998) Degradation of humic acids by manganese peroxidase from the white-rot fungus Clitocybula dusenii J Basic Microbiol 38:289-299.
Send correspondence to
Publication Dates
-
Publication in this collection
11 Apr 2014 -
Date of issue
2014
History
-
Received
30 Mar 2012 -
Accepted
01 Apr 2013