Abstract
Many studies have evaluated the effects of biochar application on soil structure and plant growth. However, there are very few studies describing the effect of biochar on native soil microbial communities. Microbial analysis of environmental samples requires accurate and reproducible methods for the extraction of DNA from samples. Because of the variety among microbial species and the strong adsorption of the phosphate backbone of the DNA molecule to biochar, extracting and purifying high quality microbial DNA from biochar-amended soil is not a trivial process and can be considerably more difficult than the extraction of DNA from other environmental samples. The aim of this study was to compare the relative efficacies of three commercial DNA extraction kits, the FastDNA® SPIN Kit for Soil (FD kit), the PowerSoil® DNA Isolation Kit (PS kit) and the ZR Soil Microbe DNA Kit MiniprepTM (ZR kit), for extracting microbial genomic DNA from sand treated with different types of biochar. The methods were evaluated by comparing the DNA yields and purity and by analysing the bacterial and fungal community profiles generated by PCR-DGGE. Our results showed that the PCR-DGGE profiles for bacterial and fungal communities were highly affected by the purity and yield of the different DNA extracts. Among the tested kits, the PS kit was the most efficient with respect to the amount and purity of recovered DNA and considering the complexity of the generated DGGE microbial fingerprint from the sand-biochar samples.
biochar; DNA extraction; PCR-DGGE; microbial communities; DNA purity indices
SHORT COMMUNICATION
Comparison of DNA extraction protocols for microbial communities from soil treated with biochar
D.C.A. LeiteI; F.C. BalieiroII; C.A. PiresII; B.E. MadariIII; A.S. RosadoI; H.L.C. CoutinhoII; R.S. PeixotoI
ILaboratório de Ecologia Microbiana Molecular, Instituto de Microbiologia Prof. Paulo de Góes, Universidade Federal do Rio de Janeiro, Rio de Janeiro, RJ, Brazil
IIEmpresa Brasileira de Pesquisa Agropecuária Solos, Rio de Janeiro, RJ, Brazil
IIIEmpresa Brasileira de Pesquisa Agropecuária Arroz e Feijão, Goiás, GO, Brazil
Send correspondence to Send correspondence to: R.S. Peixoto Laboratório de Ecologia Microbiana Molecular, Instituto de Microbiologia Prof. Paulo de Góes Av. Carlos Chagas Filho 373, Edifício do Centro de Ciências da Saúde, Bloco E sub-solo, Cidade Universitária 21941-902, Ilha do Fundão, Rio de Janeiro, RJ, Brazil E-mail: raquelpeixoto@micro.ufrj.br
ABSTRACT
Many studies have evaluated the effects of biochar application on soil structure and plant growth. However, there are very few studies describing the effect of biochar on native soil microbial communities. Microbial analysis of environmental samples requires accurate and reproducible methods for the extraction of DNA from samples. Because of the variety among microbial species and the strong adsorption of the phosphate backbone of the DNA molecule to biochar, extracting and purifying high quality microbial DNA from biochar-amended soil is not a trivial process and can be considerably more difficult than the extraction of DNA from other environmental samples. The aim of this study was to compare the relative efficacies of three commercial DNA extraction kits, the FastDNA® SPIN Kit for Soil (FD kit), the PowerSoil® DNA Isolation Kit (PS kit) and the ZR Soil Microbe DNA Kit MiniprepTM (ZR kit), for extracting microbial genomic DNA from sand treated with different types of biochar. The methods were evaluated by comparing the DNA yields and purity and by analysing the bacterial and fungal community profiles generated by PCR-DGGE. Our results showed that the PCR-DGGE profiles for bacterial and fungal communities were highly affected by the purity and yield of the different DNA extracts. Among the tested kits, the PS kit was the most efficient with respect to the amount and purity of recovered DNA and considering the complexity of the generated DGGE microbial fingerprint from the sand-biochar samples.
Key words: biochar, DNA extraction, PCR-DGGE, microbial communities, DNA purity indices.
Introduction
The addition of biochar to soil has been promoted as a tool to minimise atmospheric CO2 emissions in agricultural systems (Laird, 2008; Novak et al., 2009) and to improve plant productivity (Kwapinski et al., 2010). Biochar is the product of the thermal degradation of organic materials in the absence of oxygen (pyrolysis), and it differs from charcoal in its use in soil amendment (Lehmann et al., 2009).
Currently, well known benefits of addition of biochar to soil include increasing the pH and humidity, preserving nutrients, improving the soil structure and reducing N2O and CH4 emissions from the soil (Amonette et al., 2009). Some studies have also demonstrated that biochar can modify the composition and abundance of soil biological communities (Pietikäinen et al., 2000; Yin et al., 2000; Kim et al., 2007; O'Neill et al., 2009; Liang et al., 2010; Grossman et al., 2010; Jin, 2010). The changes produced by biochar treatment may have effects on the recycling of nutrients (Steiner et al., 2008) or even on the soil structure (Rillig et al., 2006) and may indirectly affect plant growth (Warnock et al., 2007). Bacteria and fungi from the rhizosphere may also directly promote plant growth (Schwartz et al., 2006; Compant et al., 2010). The possible connections between biochar properties and the soil microbiota, and the implications of these connections, have not been systematically described, and Lehmann et al. (2011), have indicated that this is a priority research area.
In the last few decades, molecular techniques based on the 16S rRNA gene, the fungal ITS region and other genetic markers have been developed to analyse microbial communities from environmental samples (Ovreas et al., 1997; Nakatsu et al., 2007). These methods have advantages over classical protocols because only 1-10% of the microorganisms in an environmental sample can be cultured (Hugenholtz et al., 1998; Zeyaullah et al., 2009). Fingerprinting techniques, which combine target gene amplification with amplicon separation by DGGE can provide important information about changes in the microbial community in response to environmental changes (Ovreas et al., 1997; Nakatsu et al., 2007). The success of molecular tools, including PCR-DGGE, is strongly affected by the DNA or RNA extraction method used (Santos et al., 2012). DNA isolation methods involving insufficient cell lysis or too much DNA degradation may result in underestimates (Holland et al., 2000; McOrist et al., 2002). Inhibitors within environmental samples, such as humic acids and excess protein, may create similar problems (Ariefdjohan et al., 2010).
Because of the strong adsorption of the phosphate backbone of DNA to biochar, microbial DNA extraction and purification from biochar-amended soil can be more difficult than the extraction of DNA from other environmental samples (Wilson et al., 1997; Thies et al., 2003). Jin (2010) added 10 µg of DNA to pure biochar particles and observed that the DNA recovery decreased in the presence of biochar. Approximately 30.6% of the added DNA was recovered in the absence of biochar, in contrast to 2.7% in the presence of biochar. This result indicates that biochar effectively adsorbed the DNA, demonstrating the necessity of efficient methods to improve to recovery of DNA from biochar-associated microbial communities.
Many commercial DNA extraction kits have been developed to simplify and accelerate DNA purification. The goal of this study was to compare the relative efficacies of three commercial DNA extraction kits (the FastDNA® SPIN Kit for Soil, the PowerSoil DNA Isolation Kit and the ZR Soil Microbe DNA Kit) for extracting microbial genomic DNA from soil treated with different types of biochar. These kits were compared based on the amount and the purity of the recovered DNA and on the complexity of the bacterial and fungal DGGE profiles obtained from the extracted DNA.
Materials and Methods
Sampling
Sampling was performed using microcosms maintained at Embrapa Arroz e Feijão (Santo Antônio de Goiás, GO, Brazil) on December 2009. The microcosms were constructed in flowerpots with sand (7 kg each). Biochar (12:1, vol/vol) was milled, passed through a 2-mm sieve and then carefully mixed with the sand to produce a homogenous mixture. Two different types of biochar were mixed with the sand: sugar cane straw biochar and Eucalyptus sp. wood biochar. Three field repetitions were made for each treatment, for a total of 9 microcosms. The treatments were soil + sugar cane straw biochar (SSB); soil + Eucalyptus sp. wood biochar (EWB) and soil (control, SC).
Soil DNA extraction
In this study, we tested three different methods for DNA extraction: The FastDNA® SPIN Kit for Soil (MP Biomedicals, Solon, CA, USA) (FD kit), the PowerSoil® DNA Isolation Kit (MoBio Laboratories, Carlsbad, CA, USA) (PS kit) and the ZR Soil Microbe DNA Kit MiniprepTM (Zymo Research, Irvine, CA, USA) (ZR kit). All methods are based on direct cell lysis with subsequent recovery and purification of nucleic acids. Up to 500 mg of soil was added to the lysis tubes, and the samples were homogenised in a FastPrep® for 40 s at a speed setting of 6.0 m/s. The DNA extraction was performed according to manufacturer's recommendations for each kit.
DNA quantification and purity
The DNA purity was quantified using a NanoDrop Spectrophotometer (ND-1000, NanoDrop Technology, Wilmington, DE, USA) by measuring the A260/280 and A260/230 ratios. The DNA yield was quantified using a Qubit® 2.0 Fluorometer (Invitrogen, Carlsbad, CA, USA). The Qubit® assay uses an ultrasensitive fluorescent nucleic acid stain to quantify double-stranded DNA (Invitrogen, 2010).
Co-extracted humic acids and proteins are two major contaminants of DNA extracted from environmental samples. The levels of humic acids and proteins were determined by measuring the absorbances at 230 nm and 280 nm, respectively, whereas the amount of DNA was determined by measuring the absorbance at 260 nm. In this study, the purity of the DNA was assessed spectrophotometrically by calculating the A260/A230 and A260/A280 ratios to evaluate the levels of humic acid contamination and protein impurities, respectively. A260/A230 ratios greater than 2 and A260/A280 ratios greater than 1.7 indicate high-purity DNA, and lower ratios indicate humic acid and protein contamination, respectively (Ning et al., 2009).
PCR-DGGE of the 16S gene and ITS region
The amplification of specific regions of the gene encoding the 16S rRNA was performed using the primers U968f GC (5' CGC CCG CCG CGC GCG GCG GGC GGG GCG GGG GCA CGG GGG GAA CG CGA AGA ACC TTA C 3') and L1401r (5' GCG TGT GTA CAA GAC CC 3') (Heuer et al., 1997). The amplification was performed in a solution containing 1X PCR buffer, 0.2 mM dNTPs, 2.5 mM MgCl2, 2.5 U of recombinant Taq DNA polymerase (Promega), 4 ng of total DNA, 200 µmol of each primer and sterile Milli-Q water in a final volume of 50 µL. The reaction was performed in a thermocycler (Mastercycler Gradient, Eppendorf, Hamburg, Germany) with the following conditions: initial denaturation at 94 ºC for 3 min; 35 cycles at 94 ºC for 1 min, 55 ºC for 1 min and 72 ºC for 1 min and a final extension at 72 ºC for 10 min. The DGGE gels (45-65% urea and formamide) were prepared with a solution of polyacrylamide (6%) in Trisacetate (pH 8.3). Electrophoresis was performed in Trisacetate-EDTA buffer at 60 ºC at a constant voltage of 75 V for 16 hours. The DGGE gels were stained with SYBR Green (Molecular Probes) and visualised using a Storm 860 Imaging System (GE Healthcare).
Fungal ITS regions were PCR-amplified using the primers EF4 (5' GGA AGG GRT GTA TTT ATT AG 3') (Smit et al., 1999) and ITS4 (5' TCC TCC GCT TAT TGA TAT GC 3') (White et al., 1990). The first amplification was performed in a solution containing 1X PCR buffer,
0.2 mM dNTPs, 3.75 mM MgCl2, 2.5 U of recombinant Taq DNA polymerase (Promega), 4 ng of total DNA, 200 µmol of each primer, 0.25 µL of bovine serum albumin (BSA) solution (20 mg mL-1) and sterile Milli-Q water in a final volume of 25 µL. The reaction was performed in a thermo cycler (Mastercycler Gradient, Eppendorf, Hamburg, Germany). The cycling parameters were 94 ºC for 5 min; 35 cycles of 94 ºC for 30 s, 55 ºC for 30 s and 72 ºC for 30 s; and a final extension at 72 ºC for 5 min.
The second amplification was performed with the primers ITS1-F GC (5' CTT GGT CAT TTA GAG GAA GTA A 3') (Gardes et al., 1993) and ITS2 (5' GCT GCG TTC TTC ATC GAT GC 3') (White et al., 1990). A GC clamp (5'-CGC CCG CCG CGC GCG GCG GGC GGG GCG GGG GCA CGG GGG G-3') (Muyzer et al., 1993) was added to the 5' end of the ITS1-F primer. The PCR and cycling conditions were as described above except that BSA was omitted from the reactions. The DGGE gels (30 to 60% urea and formamide) were prepared with a solution of polyacrylamide (8%) in Tris-acetate (pH 8.3). Electrophoresis was performed in Tris-acetate-EDTA buffer at 60 ºC at a constant voltage of 75 V for 16 hours. The DGGE gels were stained with SYBR Green (Molecular Probes) and visualised using a Storm 860 Imaging System (GE Healthcare).
Analysis of the bacterial and fungal DGGE profiles
The similarities between the banding patterns in the DGGE profiles were calculated based on the presence and absence of bands and were expressed as similarity coefficients. In this study, the Jaccard similarity coefficient was used for pairwise comparisons of the DGGE fingerprint profiles obtained from the three DNA extraction kits. Dendrograms that show clustering according to the similarity of the DNA fragment patterns between samples were constructed using the unweighted pair group method of arithmetic averages (UPGMA) using BioNumerics software (BioNumerics, Applied Maths, Inc., Austin, TX).
Statistical analysis
All extractions were performed in triplicate to account for analytical variability. The number of DGGE bands was analysed using Statistica (version 9.1; SAS Institute, Cary, NC) by repeated measures ANOVA. Differences between treatments were grouped using the Tukey test. Data were expressed as the mean ± SD. Differences were considered significant when the p value was < 0.05.
Results
Purity and quantity of extracted DNA
Although the three evaluated commercial kits are all based on direct lysis, significant differences were observed in the quantity and purity of the recovered DNA. In general, the FD and PS kits were able to recover more DNA than the ZR kit (Figure 1). No significant differences were observed between the FD and PS DNA extraction kits when using the control treatment (SC) samples. However, differences were detected between these DNA extraction methods for samples containing biochar (SSB and EWB), in agreement with the previously described difficulties with DNA extraction from biochar-treated soil samples (Jin, 2010).
Based on the purity index of the extracted DNA (Table 1), we observed that the FD kit resulted in a higher amount of humic acids co-extracted together with the DNA (about three times more than with the ZR kit). The PS kit resulted in the lowest level of humic acids (it extracted 36 times less humic acids than the FD kit). The three methods resulted in similar amounts of protein co-extraction.
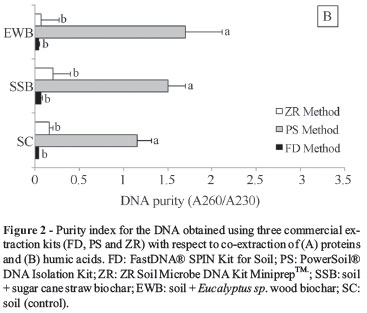
According to Ning et al. (2009), an A260/A230 ratio greater than 2 and an A260/A280 ratio greater than 1.7 indicate high-purity DNA, and lower ratios indicate humic acid and protein contamination, respectively. In this study, the ZR kit yielded the worst A260/280 ratio (~ 1.0), and the FD and PS kits both showed better results, ranging from 1.8 to 2.5 (Figure 2A). The PS kit had A260/A230 ratio values near 2, demonstrating better performance with respect to removing humic substances than the ZR and FD kits (Figure 2B).
Bacterial and fungal profiles accessed by DGGE
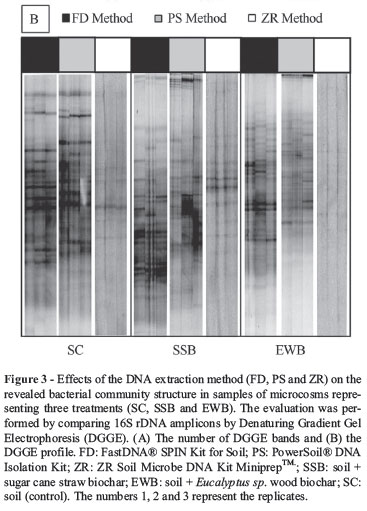
PCR products representing the total bacterial and fungal communities were evaluated by PCR-DGGE. The DGGE profiles for each community (bacterial and fungal) were evaluated for each DNA extraction method, and the number and diversity of detected bands were compared (Figures 3 and 4).
The ZR extraction method presented serious limitations for the PCR-DGGE analyses due to the low quantity of recovered DNA. The bacterial community was represented by a reduced number of bands when compared with the number of bands obtained with the other extraction methods (Figure 3A); for fungal communities, amplification was only achieved for the SSB samples (Figure 4A-B).
For the bacterial community, the number of bands in the DNA extracted using the FD and PS kits differed for the SSB samples, whereas there were no significant differences in the number of bands for the EWB and SC samples between kits (Figure 3A). In contrast, there was no significant difference in the number of bands for the fungal community between the FD and PS kits (Figure 4A).
The differences in the band resolution were best illustrated by comparisons of the profiles for the same sample obtained using different DNA extraction kits, according to Figures 3B and 4B.
Using the DGGE profiles of the microbial communities, we performed a cluster analysis to verify the influence of the extraction method on the final dendrogram. Each DNA extraction method resulted in different clusters for the bacterial community (Figure 5). The DNA extracted with the ZR and PS kits (Figure 5B-C) showed that bacterial communities of the EWB and SC treatments were similar. However, the PS kit had a higher similarity coefficient (: 72%) than the ZR kit (: 38%). This result was not observed when the FD kit (Figure 5A) was used for the DNA extraction. There was no clear clustering of the SC, SSB and EWB samples, which exhibited only approximately 58% similarity to each other. The FD and PS kits yielded very similar results for the fungal community profiles (Figure 6A-B) and indicated that the addition of biochar (EWB and SSB) modified the native fungal community (SC). When using the ZR kit, only the fungal community in the SSB samples was detected, and therefore, it was not possible to do the cluster analysis.
Discussion
The use of molecular biology methods to analyse microbial communities from environmental samples requires reproducible and efficient strategies for DNA extraction. The first DNA extractions were performed using protocols developed in-house that then became the basis for different commercially available kits (Park et al., 2005; Miller et al., 2009). Different extraction methods can vary with respect to efficiency due to the physical and chemical characteristics of samples. Therefore, diversity analysis and/or specific gene quantification is influenced by the DNA extraction method (Park et al., 2005). In our study, we investigated the feasibility of using different methods to extract microbial DNA from biochar-treated sand.
Jin (2010) suggests that biochar has a high affinity for the phosphate backbone of DNA. This hypothesis is consistent with studies performed by Chattoraj and Mitra (2009) in which genomic DNA was found to have a high binding affinity for charcoal. Jin (2010) also verified that the amount of DNA that could be extracted decreased with an increasing biochar application rate over a range from 0tha-1 (control treatment, no biochar added) to 30 t biochar ha-1. The quantity of recovered DNA decreased by half in samples containing high levels of biochar relative to control samples (no biochar added).
Our results indicate that, among the three kits tested, the FD and PS kits were the most efficient with regard to the quantity of DNA that was obtained from the biochartreated sand samples (Figure 1). However, the FD kit coextracted more humic acids than the other kits; this result is a key factor that may compromise the utility of the extracted DNA and thus interfere with DNA longevity and PCR amplification.
As reported in previous studies, biochar is a compound with a high adsorption capacity for organic matter (Baldrock et al., 2002) and soluble nutrients such as ammonium (Lehmann et al., 2002), nitrate (Mizuta et al., 2004), phosphate (Beaton et al., 1960) and other ionic solutes (Radovic et al., 2002). It is necessary to use methods that remove substances that behave similarly to DNA and may inhibit PCR amplification. The PS DNA extraction method uses three steps to separate DNA from organic and inorganic material including humic substances, cellular debris and proteins to increase the DNA purity and facilitate downstream DNA applications.
After cell lysis, the ZR kit uses column filtration during DNA purification, which had low efficiency with respect to DNA purity and yield for our samples (Figures 1 and 2). Just because biochar easily adsorbs ions, and because DNA is a negatively charged molecule, we suggest that the filtration column method may not be effectively to separate DNA from other contaminants derived from biochar due to characteristics that are shared between DNA and these contaminants.
The results obtained by PCR-DGGE analysis of the biochar-associated microbial communities revealed a clear influence of the DNA extraction method on the detected microbial diversity and community composition.
Some studies have shown that the incorporation of biochar into soil may influence soil microbial communities in a different way (Lehmann et al., 2001). After biochar addition, the soil pH may increase or decrease depending on the pH of the biochar. Biochar can have pH as low as 4 or as high as 12, depending on the feedstock type, the pyrolysis temperature (Lehmann, 2007; Chan et al., 2009) and the degree of oxidation (Cheng et al., 2006). Therefore, very different environmental conditions can be created in biochar pores when using different types of biochar. For example, fungal and bacterial populations react differently to changes in pH. Bacteria are likely to increase in abundance with increasing pH up to values of approximately 7, whereas fungi may exhibit no change in total biomass (Rousk et al., 2010) or may show dramatic reductions in growth at higher pHs (Rousk et al., 2009).
Bacterial communities
The PS and FD kits yielded DNA with an enriched band profile when compared with the DNA obtained with the ZR kit, which yielded a lower number of weaker bands with lower intensities. The results obtained for the DNA isolated with the ZR kit may be related to the lower amount of DNA recovered after extraction, which may have interfered with the PCR-DGGE profiles (Figure 3B). The DNA extracted with the ZR and PS kits demonstrated that bacterial communities in the SC (control) and EWB (eucalyptus wood biochar) samples are more similar to each other than to the communities in the SSB (sugarcane straw biochar) samples (Figure 5B-C). The same patterns were not observed when the FD (Figure 5A) kit was used to extract the DNA; in this case, the samples containing biochar (SSB and EWB) and the control (SC) were 58% similar.
These results demonstrate that the choice, evaluation and standardisation of DNA extraction methods are critical, further highlighting the importance of the DNA extraction step in microbial ecology studies. These data are in accord with the result of the studies of De Lipthay et al. (2004) and Carrigg et al. (2007), who observed that bacterial community profiles, when analysed by denaturing gradient gel electrophoresis (DGGE), were affected by different methods of DNA extraction.
Fungal communities
We did not observe differences between the fungal community profiles generated by PCR-DGGE analysis of DNA samples extracted using the PS and FD kits (Figure 6A-B). In both cases, the fungal communities profiles from biochar-treated samples were grouped together (SSB and EWB). The same pattern was not observed with the ZR kit, which only yielded detectable fungal DNA from the SSB samples (Figure 6A). Furthermore, the DNA extracts recovered from replicate samples were not reproducible. Reproducibility is a very important consideration for microbial ecology studies because variability can come not only from environmental heterogeneity but also from variability introduced by sampling and analysis (Prosser, 2010).
In conclusion, the PCR-DGGE results for bacterial and fungal communities were affected by the purity and yield of the DNA obtained with different extraction methods. The appropriate DNA extraction method depends not only on the sample type and microbial population targets but also on the analytical method used. The PS and FD kits both extracted large quantities of DNA, but only the PS kit yielded high DNA purity and a tendency towards a large numbers of bands in the DGGE profiles. The ZR kit did not cover the bacterial and fungal diversity of biochar-treated sand with the same efficiency as the PS and FD kits.
Acknowledgments
The authors would like to thank to Embrapa, the National Council for Scientific and Technological Development (CNPq), the National Council for the Improvement of Higher Education (CAPES) and the Carlos Chagas Filho Foundation for Research Support of Rio de Janeiro State (FAPERJ) for their support of this work.
Submitted: March 1, 2013
Approved: April 1, 2013.
All the content of the journal, except where otherwise noted, is licensed under a Creative Commons License CC BY-NC.
- Amonette JE, Joseph S (2009) Characteristics of biochar: microchemical properties. In: Lehmann, J., Joseph, S. (eds). Biochar for Environmental Management: Science and Technology, London, pp 33-52.
- Ariefdjohan MW, Savaiano DA, Nakatsu CH (2010) Comparison of DNA extraction kits for PCR-DGGE analysis of human intestinal microbial communities from fecal specimens. Nutr J 9-23:1-8.
- Beaton JD, Peterson HB, Bauer N (1960) Some aspects of phosphate adsorption by charcoal. Proc Soil Sci Soc Am 24:340-346.
- Baldrock JA, Smernick RJ (2002) Chemical composition and bioavailability of thermally altered Pinus resinosa (Red pine) wood. Org Geochem 33:1093-1109.
- Carrigg C, Rice O, Kavanagh S, Collins G, O'Flaherty V (2007) DNA extraction method affects microbial community profiles from soils and sediment. Appl Microbiol Biotechnol 77:955-964.
- Chan KY, Xu Z (2009) Biochar: nutrient properties and their enhancement. In: Lehmann, J., Joseph, S. (eds). Biochar for Environmental Management: Science and Technology, London, pp 67-84.
- Chattoraj DK, Mitra A (2009) Adsorption of DNA at solid-water interfaces and DNA -Surfactant binding interaction in aqueous media. Curr Sci 97-10:1430-1438.
- Cheng, CH.; Lehmann, J.; Thies, J.E.; Burton, S.D.; Engelhard, M.H. (2006). Oxidation of black carbon by biotic and abiotic processes. Org Geochem 37:1477-1488.
- Compant S, Clément S, Sessitsch A (2010) Plant growthpromoting bacteria in the rhizo-and endosphere of plants: their role, colonization, mechanisms involved and prospects for utilization. Soil Biol Biochem 42:669-678.
- De Lipthay JR, Enzinger C, Johnsen K, Aamand J, Sørensen SJ (2004) Impact of DNA extraction method on bacterial community composition measured by denaturing gradient gel electrophoresis. Soil Biol Biochem 36:1607-1614.
- Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycetes: application to the identification of mycorrhiza and rusts. Mol Ecol 2:113-118.
- Grossman JM, O'Neill BE, Tsai SM, Liang B, Neves E, Lehmann J, Thies JE (2010) Amazonian anthrosols support similar microbial communities that differ distinctly from those extant in adjacent, unmodified soils of the same mineralogy. Microb Ecol 60:192-205.
- Heuer H, Smalla K (1997) Application of denaturing gradient gel electrophoresis and temperature gradient gel electrophoresis for studying soil microbial communities. In: Elsas J.D., Trevors J.T., Wellington E.M.H. (eds) Modern Soil Microbial. Marcel Dekker, New York, pp 353-373.
- Holland JL, Louie L, Simor AE, Louie M (2000) PCR detection of Escherichia Coli O157:H7 directly from stools: evaluation of commercial extraction methods for purifying fecal DNA. J Clin Microbiol 38(11):4108-4113.
- Hugenholtz P, Goebel BM, Pace NR (1998) Impact of Culture-Independent Studies on the Emerging Phylogenetic View of Bacterial Diversity. J Bacteriol 180(18):4765-4774.
- Invitrogen. 2010. Accurate and sensitive quantitation of nucleic acids, even at low concentrations: Comparison of the Qubit IM quantitation platform with spectrophotometry. Available at: http://www.invitrogen.com/site/us/en/home/support/Newslet-ters-and-Journals/BioProbes-Journal-of-Cell-Biology-Applications/BioProbes-Issues-2010/BioProbes-62/Comparison-of-Qubit-Quantitation-Platform-With-Spectrophotometry.html Acessed 03 november 2011.
- Jin H (2010) Characterization of microbial life colonizing biochar and biochar amended soils. NY, USA (PhD Thesis, Cornell University).
- Kim JS, Sparovek S, Longo RM, De Melo, WJ, Crowley D (2007) Bacterial diversity of terra preta and pristine forest soil from the Western Amazon. Soil Biol Biochem 39:648-690.
- Kwapinski W, Byrne CMP, Kryachko E, Wolfram P, Adley C, Leahy J J, Novotny EH, Hayes M. HB (2010) Biochar from Biomass and Waste. Waste and Biomass Valorization 1(2):177-189.
- Laird DA (2008) The charcoal vision: a winewin scenario for simultaneously producing bioenergy, permanently sequestering carbon, while improving soil and water quality. Agron J 100:178-181.
- Lehmann J (2007) Bio-energy in the black. Front. Ecol Environ 5:381-387.
- Lehmann J, da Silva J, Rondon M, Cravo MS, Greenwood J, Nehls T, Steiner C, Glaser B (2002) Proceedings of the 17th World Congress of Soil Science: Slash-and-char -A feasible alternative for soil fertility management in the central Amazon? Bangkok, Thailand, v. 1, p. 12.
- Lehmann J, Joseph S (2009) Biochar for Environmental Management: An Introduction. In: Lehmann J, Joseph S (eds) Biochar for Environmental Management: Science and Technology, London, pp 251-265.
- Lehmann J, Rillig MC, Thies J, Masiello CA, Hockaday WC, Crowley D (2011) Biochar effects on soil biota -A review. Soil Biol Biochem 43:1812-1836.
- Liang B, Lehmann J, Sohi SP, Thies JE, O'Neill B, Trujillo L, Gaunt J, Solomon D, Grossman J, Neves EG, Luizão FJ (2010) Black carbon affects the cycling of non-black carbon in soil. Org Geochem 41:206-213.
- McOrist AL, Jackson M, Bird AR (2002) A comparison of five methods for extraction of bacterial DNA from human faecal samples. J Microbiol Meth 50(2):131-139.
- Miller DN, Bryant JE, Madsen EL, Ghiorse WC (1999) Evaluation and optimization of DNA extraction and purification procedures from soil and sediment samples. Appl Environ Microbiol 65:4715-4724.
- Mizuta K, Matsumoto T, Hatate Y, Nishihara K, Nakanishi T (2004) Removal of nitrate-nitrogen from drinking water using bamboo powder charcoal. Bioresource Technol 95:255-257.
- Muyzer G, Wall EC, Uitterlinden AG (1993) Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Am Soc Microbiol 59(3):695-700.
- Nakatsu CH (2007) Soil microbial community analysis using Denaturing Gradient Gel Electrophoresis. Soil Sci Soc Am J 71(2):562-571.
- Ning J, Liebich J, Kästner M, Zhou J, Schäffer A, Burauel P (2009) Different influences of DNA purity indices and quantity on PCR-based DGGE and functional gene microarray in soil microbial community study. Appl Microbiol Biotechnol 82:983-993.
- Novak JM, Busscher WJ, Laird DA, Ahmedna M, Watts DW, Niandou M. (2009) Impact of biochar amendment on fertility of a southeastern coastal plain soil. Soil Sci 174:105-112.
- O'Neill B, Grossman J, Tsai MT, Gomes JE, Lehmann J, Peterson J, Neves E, Thies JE (2009) Bacterial Community Composition in Brazilian Anthrosols and Adjacent Soils Characterized Using Culturing and Molecular Identification. Microb Ecol 58(1):23-35.
- Ovreas L, Forney L, Daae F, Torsvik V (1997) Distribution of bacterioplankton in meromictic Lake Sąlenvannet, as determined by denaturing gradient gel electrophoresis of PCRamplified gene fragments coding for 16S rRNA. Appl Environ Microbiol 63(9):3367-3373.
- Park JW, Crowley DE (2005) Normalization of soil DNA extraction for accurate quantification of target genes by real-time PCR and DGGE. BioTech 38:579-586.
- Pietikäinen J, Kiikkilä O, Fritze H (2000) Charcoal as a habitat for microbes and its effects on the microbial community of the underlying humus. Oikos, 89:231-242.
- Prosser JI (2010) Replicate or lie. Environ Microbiol 12(7):1806-1810.
- Radovic LR, Moreno-Castilla C, Rivera-Utrilla J (2001) Carbon materials as adsorbents in aqueous solutions. In: Radovic, L.R. (eds). Chemistry and Physics of Carbon. New York, pp 227-405.
- Rillig MC, Mummey DL (2006) Mycorrihizas and Soil Structure. New Phytol 171:41-53.
- Rousk J, Bååth E, Brookes PC, Lauber CL, Lozupone C, Caporaso JG, Knight R, Fierer N (2010) Soil bacterial and fungal communities across a pH gradient in an arable soil. ISME J 4:134-151.
- Rousk J, Brookes PC, Bååth E (2009) Contrasting soil pH effects on fungal and bacterial growth suggest functional redundancy in carbon mineralization. Appl Environ Microbiol 75:1589-1596.
- Santos HF, Carmo FL, Leite DCA, Jesus HE, Maalouf PC, Almeida C, Soriano A, Altomani D, Suhett L, Volaro V, Valoni E, Francisco M, Vieira J, Rocha R, Sardinha BL, Bacelllar LB, João LB, Lacava B, Sebastian GV, Pessoa A, Elsas JDV, Pires D, Duarte G, Castro C, Rosado AS, Peixoto RS (2012) Comparison of different protocols for the extraction of microbial DNA from reef corals. Braz J Microbiol 43:517-527.
- Schwartz MW, Hoeksema JD, Gehring CA, Johnson NC, Klironomos JN, Abbott K, Pringle A (2006) The promise and the potential consequences of the global transport of mycorrhizal fungal inoculum. Ecol Lett 9:501-515.
- Smit E, Leeflang P, Glandorf B, Van Elsas JD, Wernars K (1999) Analysis of fungal diversity in the wheat rhizosphere by sequencing of cloned PCR-amplified genes encoding 18S rRNA and temperature gradient gel electrophoresis. Appl Environ Microbiol 65:2614-2621.
- Steiner C, Glaser B, Teixeira WG, Lehmann J, Blum WEH, Zech W (2008) Nitrogen retention and plant uptake on a highly weathered central Amazonian Ferralsol amended with compost and charcoal. J Plant Nutrit Soil Sci 171:893-899.
- Thies J, Suzuki K (2003) Amazonian dark earths: Biological measurements. In: Lehman, J (ed.). Amazonian Dark Earths: Origin, Properties, Management. Dordrecht, Netherlands, pp 287-332.
- Warnock DD, Lehmann J Kuyper TW, Rillig MC (2007) Mycorrhizal responses to biochar in soil e concepts and mechanisms. Plant Soil 300:9-20.
- White TJ, Bruns TD, Lee S, Taylor J (1990) Analysis of phylogenetic relationships by amplification and direct sequencing of ribosomal RNA genes. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR Protocols: a Guide to Methods and Applications. New York, pp 315-322.
- Wilson IG (1997) Inhibition and facilitation of nucleic acid amplification. Appl Environ Microbiol 63:3741-3751.
- Yin B, Crowley D, Sparovek, G, De Melo WJ, Borneman J (2000) Bacterial functional redundancy along a soil reclamation gradient. Appl Environ Microbiol 66(10):4361-4365.
- Zeyaullah M, Kamli MR, Islam B, Atif M, Benkhayal FA, Nehal M, Rizvi MA, Ali A (2009) Metagenomics -An advanced approach for noncultivable micro-organisms. Biotech Mol Biol Rev 4(3):49-54.
Send correspondence to:
Publication Dates
-
Publication in this collection
29 May 2014 -
Date of issue
2014
History
-
Accepted
01 Apr 2013 -
Received
01 Mar 2013