Abstracts
The Pampas deer (Ozotoceros bezoarticus) is one of the most endangered Neotropical cervid with populations that have been drastically reduced to small and isolated ones, mainly because of its habitat destruction. Random amplified polymorphic DNA (RAPD) markers were used to analyze population divergence and genetic variation within and between two populations corresponding to distinct subspecies. The RAPD markers displayed substantial genetic variation with all animals possessing unique RAPD phenotypes over 105 polymorphic bands produced by 15 primers. An analysis of molecular variance (AMOVA) and a neighbor-joining cluster analysis were performed to assess levels of differentiation between populations. No differentiation was recorded and about 96.0% (P < 0.00001) of the total variance was attributable to variation within populations. This result is quite distinct from data obtained by the analysis of the mtDNA control region, and is discussed on the basis of genetic differences between the different markers and the male-biased dispersal patterns generally observed in the mammal species. The data presented herein are potentially useful for future taxonomic and genetic studies in this species, for the monitoring of the genetic variation observed within these populations, and for the development of management guidelines for its conservation.
Cervidae; Ozotoceros bezoarticus; RAPD; genetic diversity; population structure
O Veado-campeiro (Ozotoceros bezoarticus) é uma das espécies de cervídeos neotropical mais ameaçadas devido à destruição de seu hábitat e conseqüente redução e isolamento de suas populações. Marcadores do tipo RAPD (Random amplified polymorphic DNA) foram utilizados na análise da divergência populacional e estimativa da variação genética dentro e entre duas populações correspondentes a diferentes subespécies. Os marcadores RAPD mostraram uma variação genética substancial, sendo que as 105 bandas polimórficas obtidas pelo uso de 15 primers produziram fenótipos únicos para todos os indivíduos analisados. Para avaliar o nível de diferenciação entre as populações, foi realizada uma análise da variância molecular (AMOVA) e uma análise de agrupamento utilizando o método de neighbor-joining. Nenhuma diferenciação foi observada, sendo aproximadamente 96,0% da variação encontrada atribuída à variação dentro das populações estudadas. Este resultado difere do obtido através da análise da região controle do mtDNA, e é discutido levando-se em consideração as diferenças genéticas entre os diferentes marcadores utilizados e o padrão de dispersão geralmente observado nas espécies de mamíferos (realizada principalmente pelos machos). Os dados aqui apresentados poderão ser úteis para futuros estudos taxonômicos e genéticos desta espécie, para o monitoramento da variação genética observada em suas populações e para o desenvolvimento de estratégias de manejo para sua conservação.
Cervidae; Ozotoceros bezoarticus; RAPD; diversidade genética; estrutura populacional
Genetic diversity of two Brazilian populations of the Pampas deer (Ozotoceros bezoarticus, Linnaeus 1758)
Diversidade genética de duas populações brasileiras de Veado-campeiro (Ozotoceros bezoarticus, Linnaeus 1758)
Rodrigues, FP.I, * * e-mail: barbanti@fcav.unesp.br, fprodrigues_consgen@yahoo.com.br ; Garcia, JF.II; Ramos, PRR.I; Bortolozzi, J.III; Duarte, JMB.IV, * * e-mail: barbanti@fcav.unesp.br, fprodrigues_consgen@yahoo.com.br
IDepartamento de Física e Biofísica, Instituto de Biociências, Universidade Estadual Paulista "Júlio de Mesquita Filho" UNESP, Distrito de Rubião Jr., s/n, CEP 18618-000, Botucatu, SP, Brazil
IIDepartamento de Apoio, Saúde e Produção Animal, Faculdade de Medicina Veterinária, Universidade Estadual Paulista "Júlio de Mesquita Filho" UNESP, Rua Clóvis Pestana 793, CEP 16050-680, Araçatuba, SP, Brazil
IIIDepartamento de Biologia, Faculdade de Ciências, Universidade Estadual Paulista "Júlio de Mesquita Filho" UNESP, Av. Eng. Luiz Edmundo C. Coube 14-01, CEP 17033-360, Bauru, SP, Brazil
IVDepartamento de Zootecnia, Faculdade de Ciências Agrárias e Veterinárias, UNESP, Via de Acesso Prof. Paulo D. Castelane, s/n, CEP 14884-900, Jaboticabal, SP, Brazil
ABSTRACT
The Pampas deer (Ozotoceros bezoarticus) is one of the most endangered Neotropical cervid with populations that have been drastically reduced to small and isolated ones, mainly because of its habitat destruction. Random amplified polymorphic DNA (RAPD) markers were used to analyze population divergence and genetic variation within and between two populations corresponding to distinct subspecies. The RAPD markers displayed substantial genetic variation with all animals possessing unique RAPD phenotypes over 105 polymorphic bands produced by 15 primers. An analysis of molecular variance (AMOVA) and a neighbor-joining cluster analysis were performed to assess levels of differentiation between populations. No differentiation was recorded and about 96.0% (P < 0.00001) of the total variance was attributable to variation within populations. This result is quite distinct from data obtained by the analysis of the mtDNA control region, and is discussed on the basis of genetic differences between the different markers and the male-biased dispersal patterns generally observed in the mammal species. The data presented herein are potentially useful for future taxonomic and genetic studies in this species, for the monitoring of the genetic variation observed within these populations, and for the development of management guidelines for its conservation.
Keywords: Cervidae, Ozotoceros bezoarticus, RAPD, genetic diversity, population structure.
RESUMO
O Veado-campeiro (Ozotoceros bezoarticus) é uma das espécies de cervídeos neotropical mais ameaçadas devido à destruição de seu hábitat e conseqüente redução e isolamento de suas populações. Marcadores do tipo RAPD (Random amplified polymorphic DNA) foram utilizados na análise da divergência populacional e estimativa da variação genética dentro e entre duas populações correspondentes a diferentes subespécies. Os marcadores RAPD mostraram uma variação genética substancial, sendo que as 105 bandas polimórficas obtidas pelo uso de 15 primers produziram fenótipos únicos para todos os indivíduos analisados. Para avaliar o nível de diferenciação entre as populações, foi realizada uma análise da variância molecular (AMOVA) e uma análise de agrupamento utilizando o método de neighbor-joining. Nenhuma diferenciação foi observada, sendo aproximadamente 96,0% da variação encontrada atribuída à variação dentro das populações estudadas. Este resultado difere do obtido através da análise da região controle do mtDNA, e é discutido levando-se em consideração as diferenças genéticas entre os diferentes marcadores utilizados e o padrão de dispersão geralmente observado nas espécies de mamíferos (realizada principalmente pelos machos). Os dados aqui apresentados poderão ser úteis para futuros estudos taxonômicos e genéticos desta espécie, para o monitoramento da variação genética observada em suas populações e para o desenvolvimento de estratégias de manejo para sua conservação.
Palavras-chave: Cervidae, Ozotoceros bezoarticus, RAPD, diversidade genética, estrutura populacional.
1. Introduction
The Pampas deer (Ozotoceros bezoarticus) is a medium-size cervid that could be originally found in a vast area at the south-eastern part of South America (between 5° and 41° S) occupying open habitats as grasslands, pampas, savannas, and cerrado (Brazil). Due to human activities, these habitats have been drastically reduced so that this species is, perhaps, the most endangered tropical Latin American deer and is currently listed as in danger of extinction according to Appendix 1 of the CITES (Wemmer, 1998; Weber and González, 2003). The largest population is found in Brazil, where at least 60,000 individuals are found in the Pantanal biome (Mourão et al., 2000). In other Brazilian sites and in Argentina, Bolivia, Paraguay, and Uruguay, it exists in small and isolated populations (Merino et al., 1997).
Actually, there are five recognized Pampas deer subspecies. Based on morphological data, Cabrera (1943, 1960) suggested that O. b. bezoarticus can be found from central Brazil, south of Amazonia, between the Mato Grosso plateau and upper the São Francisco River, while O. b. leucogaster can be found from south-western Brazil, south-eastern Bolivia, Paraguay, and north Argentina. In the Argentinean pampas, O. b. celer subspecies could be found from the Atlantic coast to Andean foothills and southward to the Negro River. More recently, González et al. (2002), based on morphometric and genetic differences, proposed that populations from Uruguay should be considered as two separated subspecies, O. b. uruguayensis and O. b. arerunguaensis.
Until now, only one genetic study was done in this species. In this work, González et al. (1998) found a remarkable high level of genetic diversity within Pampas deer populations when analyzing the control region of the mitochondrial DNA (mtDNA). A high sequence divergence correlated with geographic distance was found between populations, suggesting that genetic differentiation in this species is largely explained by the limited dispersal abilities of individuals rather than the presence of long-standing ecological or geographical barriers. The pronounced mtDNA sequence divergence found between populations from central and western Brazil (classified as O. b. bezoarticus and O. b. leucogaster, respectively) led these authors to suggest that populations from these regions should be treated as distinct management units.
It is consensus in the scientific community that, whenever possible, more than one kind of genetic marker should be used to characterize the genetic diversity and population structure of a species. Comparing markers with different modes of inheritance is useful to avoid biased results originated by differences in its characteristics and to provide insights into patterns of sex-biased dispersal and gene flow. The use of female-inherited markers (such as mtDNA) and bi-parentally inherited autosomal markers (like RAPDs and microsatellites, among others) has been widely reported in the literature revealing contrasting genetic patterns that can be found accordingly to the kind of marker that was used (Ishibashi et al., 1997; Gibbs et al., 2000; Prugnolle and Meeus, 2002).
Random Amplified Polymorphic DNA (RAPD) markers (Welsh and McClelland, 1990; Williams et al., 1990) present several interesting characteristics that make them very useful for the survey of genetic diversity. Among these, are the numerous loci that can be assayed at relatively low cost and without previously knowledge of its genome, the sampling of the whole genome, and their high polymorphism (Perez-Sweeney et al., 2003). In mammals, RAPD technique has been used to solve several tasks like hybridization detection (Tsvirka et al., 2006), species diagnosis (Martinez and Danielsdottir, 2000; Basckevich et al., 2004), genetic mapping (Shiue et al., 1999), evaluation of genetic diversity and population differentiation (Petrosyan et al., 2002; Spiridonova et al., 2004), and determination of systematic relationships between taxa (Bellinvia et al., 1999; Mishra et al., 2002).
In order to improve our knowledge on the genetic structure and diversity found in the Pampas deer, we used RAPD markers to study two distinct populations: one from the cerrado biome (O. b. bezoarticus, from central Brazil) and other from the Pantanal biome (O. b. leucogaster, from western Brazil).
2. Material and Methods
2.1. Sample collection
Forty-two free-ranging individuals were sampled and used in the analysis; 23 from Emas National Park (Goiás State, central Brazil - 18° 15 S and 52° 53 W), and 19 from Pantanal (Mato Grosso do Sul State, western Brazil - 19° 59 S and 56° 39 W). Animals were captured using the "fast-setting net" method (Nunes et al., 1997) and anesthetized with a mixture of intramuscular xylazine hydrochloride (1 mg.kg-1) and ketamine (5 mg.kg-1). Blood samples were collected using vacuum tubes containing heparin salt (Vacuntainer®) and placed on ice until they could be processed in a field lab, which was done within 5 hours of collection. In the field lab, leukocytes were separated from blood samples and preserved in liquid nitrogen (196 °C) until DNA extraction was performed.
2.2. Molecular methods
DNA was extracted from leukocytes using a standard phenol-chloroform method (Sambrook and Russel, 2001) and diluted to 4 ng.µL-1 as the working solution. In order to select primers that produce clearly identifiable polymorphic bands, we screened 80 random primers (Kits B, J, K, and M, from Operon Technologies, Inc.; Alameda, CA, USA) on five individuals from each population. Fifteen out of the eighty primers yielded the desirable pattern and were used in all the samples. PCR reactions were performed in a volume of 15 µL containing 10 mM Tris-HCl (pH 8.4), 50 mM KCl, 3.5 mM MgCl2, 0.2 mM of each dNTP, 0.2 µM of primer, 1.25 U.I. of Taq DNA polymerase, and 4 ng of template DNA, covered with a thin layer of mineral oil. DNA amplifications were carried out in a M. J. Research PTC-100 thermocycler programmed for an initial denaturation step of 94 °C for 5 minutes, followed by 40 cycles of 15 seconds at 94 °C, 30 seconds at 40 °C, and 60 seconds at 72 °C. The reaction was completed with a final run at 72 °C for 5 minutes. Amplification products were separated on 1.4% agarose gels in 0.5x TBE buffer for 2 hours and 15 minutes at 100V. After electrophoresis, gels were stained with ethidium bromide (0.5 mg.mL-1), visualized under UV light, and photographed for later scoring and analysis.
2.3. Data analysis
Each obtained marker was considered as a distinct and independent phenotype, and bands with same electrophoretic mobility were considered identical independently of their intensity. Ambiguous and extremely light bands were disconsidered. Electropherograms were used for the construction of a binary matrix, where the presence of a band was designated as 1, and its absence as 0. Using the RAPDistance 1.04 software (Armstrong et al. 1994), banding patterns were converted into pairwise distance matrices using both the Euclidean metrics of Excoffier et al. (1992) and Nei and Li (1979), which is equivalent to the Dice (1945) algorithm. The genetic distance between populations was estimated by averaging distance values obtained in pairwise comparisons between individuals of these populations. We used the Analysis of Molecular Variance (AMOVA) procedure (Excoffier et al., 1992) to estimate variance components for RAPD phenotypes, partitioning the variation among individuals within population and among populations. The AMOVAs were performed using the Arlequim 3.11 program (Excoffier et al., 2005). A cluster analysis tree was produced using the NEIGHBOUR program from the PHYLIP package, version 3.66, using the neighbor-joining option (Felsenstein, 2004). The treefile produced by the PHYLIP program was visualized and manipulated using the software MEGA4 (Tamura et al., 2007). Average gene diversity across all loci (Nei, 1987) within and among populations was calculated using Arlequim program.
3. Results
The 15 selected primers generated a total of 160 scorable bands (10.66 bands/primer), 55 monomorphic (3.66 bands/primer) and 105 polymorphic (7.0 bands/primer). Unique RAPD fragments were found at both populations: eight in individuals from Emas National Park, and three in individuals from Pantanal. Eight of the 105 polymorphic markers were fixed at Emas National Park, while 11 were fixed into Pantanal.
Pairwise genetic distances were computed between all individuals in the study, according to Nei and Li (1979) and Excoffier et al. (1992) coefficients. Since results obtained with both metrics were almost identical, here we present only those obtained with Nei and Li (1979) algorithm. Distances between individuals varied from 0.12 to 0.33 for comparisons within populations, and from 0.10 to 0.35 among populations. Averaged distances were very similar at both populations, corresponding to 0.210 and 0.216 to Emas and Pantanal populations respectively, and to 0.222 between them. Average gene diversity across all loci was similar for both populations too, showing a value of 0.259 for the entire sample. Estimates of genetic distance and gene diversity are shown in Table 1.
Estimates of variance components within and between populations, calculated using AMOVA (Table 2), revealed that almost 96.0% (P < 0.0001) of the total variance could be attributed to variation within populations, while among-population variation represented only 4.0% of the total variation.
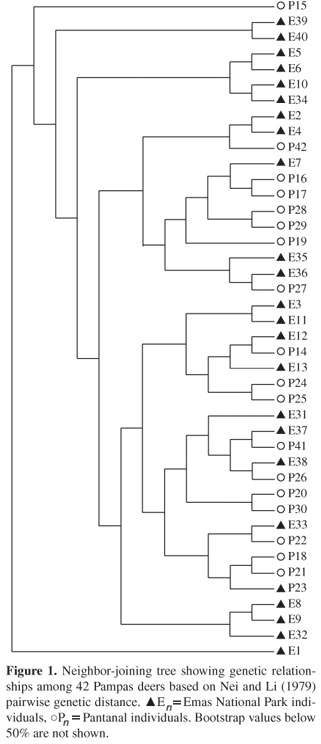
To analyze individual relationships within and among populations we used the pairwise distance comparisons to construct a neighbor-joining tree. As shown in Figure 1, cluster analysis was not able to set individuals into their population of origin.
4. Discussion
Despite the problems usually associated to its use (i.e., the dominant mode of inheritance and the considerable care required to obtain repeatable results), the RAPD technique is still been applied in a variety of situations, especially in the study of those species where other kinds of nuclear markers, such as microsatellites, are not available. In mammal species, the most frequent use of RAPD marker is related to the analysis of intraespecific genetic variation and it has been applied in a variety of animal groups like rodents (Vucetich et al., 2001; Chiappero and Gardenal, 2003; Spiridonova et al., 2004), marsupials (Cooper, 2000), carnivores (Ratnayeke et al., 2002), and primates (Neveu et al., 1998). Few studies using the RAPD technique have been reported on cervid species. Among these, there are analysis related to the establishment of systematic relationships (Comincini et al., 1996), analysis of intra and interespecific variation (Scandura et al., 1998; Tokarskaya et al., 2000), and development of specie-specific markers (Wu et al., 2006).
In the present study, we used RAPD markers to analyze genetic diversity and population structure of the Pampas deer Ozotoceros bezoarticus, one of the most endangered Neotropical cervid. Based on the average gene diversity value obtained using Nei (1987) index, we concluded that a moderate to high level of genetic diversity is present at the two studied populations. Originally derived to be used with codominant markers, the use of Neis gene diversity index for dominant markers became a measure of genetic variability of statistical value, with values ranging from 0-0.5 (Lowe et al., 2004). Here, a value of 0.26 was found when considering both populations together. The existence of considerable variation within each population is also evident by the fact that each individual was characterized by a unique RAPD phenotype. Using the Nei and Li (1979) algorithm, we found average distance values of 0.210 and 0.216 for Emas and Pantanal populations, respectively. These values seem to be in the range of those usually reported in the literature, but the lack of standard procedures for the analysis of RAPD data and the use of different kinds of similarity and distance indexes make comparisons with other studies very difficult.
These results are concordant with González et al. (1998) that found a remarkably high variability in the control region of the mitochondrial DNA. This high genetic variability at nuclear (RAPD) and mtDNA at both populations can reflect the historically large population sizes and their recent decline. The abundance of this species in the past can be deduced from archaeological sites where it is commonly found (Tonni et al., 1992). Its exploitation and consequent population decline can also be easily deduced when we check the numbers of pelts exported from Rio de La Plata in the nineteen century: about 2,130,000 only between 1860 and 1870 (Thornback and Jenkins, 1982). Actually we can find about 1,300 individuals at Emas National Park (Rodrigues, 2003) while at the Pantanal, where the largest population of this species resides, there is about 60,000 individuals (Mourão et al., 2000). Pampas deer is still suffering a strong pressure over its populations, mainly by hunting activities, contact with cattle disease, and habitat destruction promoted by agricultural expansion (Merino et al., 1997).
Based on RAPD markers, we were not able to separate individuals into their populations of origin. The data presented here show a very low degree of structuring, with a value of among population variance in AMOVA of just 4.0%. This lack of population division is clearly visible in the neighbor-joining tree generated by the cluster analysis. These results are quite distinct from those obtained by González et al. (1998) for mtDNA. This can be a consequence of the differences in the genetic characteristics of the markers used in each study, since different classes of markers can give rise to very distinct results. According to Avise (2004), a species can show distinct patterns in its geographical population structure when the analyses are done using genes with biparental transmission (which is the case for the majority of nuclear loci) or using those in which the transmission occurs mainly by one of the sexes (like in the mtDNA). One of the most important characteristics of mtDNA is it maternal inheritance in most species. It means that evolutionary relationships inferred by using this marker must be interpreted as an inference of matriarchal phylogeny (Avise, 2004). In that way, depending on the presence or not of a sex-biased dispersal pattern in the species under study, distinct genetic structures can be found when using nuclear or mitochondrial DNA markers. As a general rule, males are the dispersal sex in the mammal species (Greenwood, 1980; Handley and Perrin, 2007), and differences in geographic structure of mitochondrial and nuclear polymorphisms caused by this biased dispersal pattern have been widely reported (Ishibashi et al., 1997; Mesa et al., 2000; Escorza-Trevino and Dizon, 2000; Prugnolle and Meeus, 2002). Unfortunately, dispersal patterns in the pampas deer are poorly known, but the hypothesis that in this species the dispersion is done mainly by males can explain the differences on the data found in our study from those obtained by González et al. (1998).
Although our data analysis evidenced a lack of genetic structure and population subdivision, it is important to note that 11 exclusive RAPD markers were found at both populations. If it can not be taken as a proof of population distinctiveness, this data at least suggest a limited dispersion between these two populations.
The analyzed populations are considered two distinct subspecies: O. b. bezoarticus (samples from Emas National Park), and O. b. leucogaster (samples from Pantanal). This classification, primarily based on morphological characters (Cabrera, 1943), was reinforced by the remarkable mtDNA sequence divergence between them (González et al., 1998). Aside this, the habitats occupied by these two populations present diverse climate conditions. Emas National Park is located at the cerrado of central Brazil between 650-1,000 m of altitude and has a marked dry season lasting from April to September. Pantanal is a low altitude flooded plain (80 m) composed primarily by grassland with a very hot and humid climate during almost all year, with food and area availability depending on flooding cycles. Distinct environmental conditions can give rise to different evolutionary forces, which in turn can drive selection of adaptive characteristics that could be useful in these different environments. Until now, few studies on characteristics of adaptive value have been conduct in Pampas deer. Among these, studies on endocrine monitoring, antler cycle, and reproductive behavior seem to be insufficient for concluding that there are physiological differences between Emas and Pantanal populations (Pinder, 1992; Tomas, 1995; Rodrigues, 1996; Pereira et al., 2005).
Concerning to population structure, the results presented here for RAPD are quite different from those obtained from mtDNA analysis. More than just report these differences, we would like to stress the need for assessing patterns of genetic structure using multiple classes of genetic markers. Different kinds of markers can underscore different population histories and difficult-to-study behaviors. As demonstrated by Hoffman et al. (2006), even the same kind of marker can give rise to distinct results depending on the sample size and number of analyzed loci. By this way, conservation efforts and management recommendations need to be addressed with care and based on as much information as possible. We suggest that more studies using other nuclear markers (like microsatellites) and field works on Pampas deer dispersal abilities are needed before effective conservation and management guidelines could be done for this species and its populations.
Acknowledgments We would like to thank Renato Caparroz for comments during the work. The capture of the animals and the blood sampling were conducted with the permission of IBAMA, permits 021/96 (Process n. 02001.004425/9411). This work was supported, in part, by FAPESP (fellowship to FPR) and Environment National Fund (Fundo Nacional do Meio Ambiente FNMA).
Received May 31, 2007
Accepted October 10, 2007
Distributed December 1, 2007
- ARMSTRONG, JS., GIBBS, AJ., PEAKALL, R. and WEILLER, G., 1994. The RAP Distance Package Available from: http://www.anu.edu.au/BoZo/software/index.html [Access: May 2007].
- AVISE, JC., 2004. Molecular markers, natural history an evolution 2nd ed. New York, Sinauer Associates, 684p.
- BASKEVICH, MI., POTAPOV, SG., OKULOVA, NM. and BALAKIREV, AE., 2004. Diagnostics of syntopic mouse species of the genus Apodemus in the western Great Caucasus. Zoolog. Zhurnal, vol. 83, no. 10, p. 1261-1269.
- BELLINVIA, E., MUNCLINGER, P. and FLEGR, J., 1999. Application of the RAPD technique for a study of the phylogenetic relationships among eight species of the genus Apodemus. Folia Zoologica, vol. 48, no. 4, p. 241-248.
- CABRERA, A., 1943. Sobre la sistemática del venado y su variación individual y geográfica. Rev. Mus. La Plata, Zool, vol. 3, no. 18, p. 5-41.
- -, 1960. Catalogo de los mamiferos de America del Sur. Rev. Museo Arg. Ciências Nat. "Bernardino Rivadavia", vol. 4, no. 2, p. 309-732.
- CHIAPPERO, MB. and GARDENAL, CN., 2003. Restricted Gene Flow in Calomys musculinus (Rodentia, Muridae), the Natural Reservoir of Junin Virus. J. Heredity, vol. 94, no. 6, p. 490-495.
- COMINCINI, S., SIRONI, M., BANDI, C., GIUNTA, C., RUBINI, M. and FONTANA, F., 1996. RAPD analysis of systematic relationships among the Cervidae. Heredity, vol. 76, no. 3, p. 215-221.
- COOPER, ML., 2000. Random amplified polymorphic DNA analysis of southern brown bandicoot (Isoodon obesulus) populations in Western Australia reveals genetic differentiation related to environmental variables. Mol. Ecol, vol. 9, no. 4, p. 469-479.
- DICE, LR., 1945. Measures of the amount of ecologic association between species. Ecology, vol. 26, no. 3, p. 297-302.
- ESCORZA-TREVINO, S. and DIZON, AE., 2000. Phylogeography, intraspecific structure and sex-biased dispersal of Dalls porpoise, Phocoenoides dalli, revealed by mitochondrial and microsatellite DNA analyses. Mol. Ecol, vol. 9, no. 8, p. 1049-1060.
- EXCOFFIER, L., SMOUSE, P. and QUATTRO, J., 1992. Analysis of molecular variance inferred from metric distances among DNA haplotypes: Application to human mitochondrial DNA restriction data. Genetics, vol. 131, no. 2, p. 479-491.
- EXCOFFIER, L., LAVAL, G. and SCHNEIDER, S., 2005. Arlequin ver. 3.0: An integrated software package for population genetics data analysis. Evol. Bioinf. Online, vol. 1, p. 47-50.
- FELSENSTEIN, J., 2004. PHYLIP (Phylogeny Inference Package) version 3.6. Seattle, Department of Genome Sciences, University of Washington. Available from: http://evolution.gs.washington.edu/phylip.html [Access: May 2007].
- GIBBS, HL., DAWSON, RJG. and HOBSON, KA., 2000. Limited differentiation in microsatellite DNA variation among northern populations of the yellow warbler: evidence for male-biased gene flow? Mol. Ecol, vol. 9, no. 12, p. 2137-2147.
- GONZÁLEZ, S., MALDONADO, JE., LEONARD, JA., VILÀ, C., DUARTE, JMB., MERINO, M., BRUM-ZORRILLA, N. and WAYNE, RK., 1998. Conservation genetics of the endangered Pampas deer (Ozotoceros bezoarticus). Mol. Ecol, vol. 7, no. 1, p. 47-56.
- GONZÁLEZ, S., ÁLVAREZ-VALIN, F. and MALDONADO, JE., 2002. Morphometric differentiation of endangered pampas Deer (Ozotoceros bezoarticus), with description of new subspecies from Uruguay. J. Mammal, vol. 83, no. 4, p. 1127-1140.
- GREENWOOD, PJ., 1980. Mating systems, philopatry and dispersal in birds and mammals. Anim. Behav, vol. 28, no. 4, p. 1140-1162.
- HANDLEY, LJ. and PERRIN, N., 2007. Advances in our understanding of mammalian sex-biased dispersal. Mol. Ecol, vol. 16, no. 8, p. 1559-1578.
- HOFFMAN, JI., MATSON, CW., AMOS, W., LOUGHLIN, TR. and BICKHAM, JW., 2006. Deep genetic subdivision within a continuously distributed and highly vagile marine mammal, the Stellers sea lion (Eumetopias jubatus). Mol. Ecol, vol. 15, no. 10, p. 2821-2832.
- ISHIBASHI, Y., SAITOH, T., ABE, S. and YOSHIDA, MC., 1997. Sex-related spatial kin structure in a spring population of grey-sided voles Clethrionomys rufocanus as revealed by mitochondrial and microsatellite DNA analyses. Mol. Ecol, vol. 6, no. 1, p. 63-71.
- LOWE, A., HARRIS, S. and ASHTON, P., 2004. Ecological Genetics: Design, Analysis, and Application Oxford, Blackwell Publishing Company, 326p.
- MARTINEZ, I. and DANIELSDOTTIR, AK., 2000. Identification of marine mammal species in food products. J. Science Food Agric, vol. 80, no. 4, p. 527-533.
- MERINO, ML., GONZALES, S., LEEUWENBERG, F., RODRÍGUEZ, FHG., PINDER, L. and TOMAS, WM., 1997. Veado-campeiro (Ozotoceros bezoarticus). In DUARTE, JMB. (ed.). Biologia e conservação de cervídeos sul-americanos: Blastocerus, Ozotoceros e Mazama Jaboticabal, Funep, p. 42-58.
- MESA, NR., MONDRAGON, MC., SOTO, ID., PARRA, MV., DUQUE, C. and ORTIZ-BARRIENTOS, D., 2000. Autosomal, mtDNA, and Y-chromosome diversity in Amerinds: pre- and post-colombian patterns of gene flow in South America. Am. J. Hum. Genet, vol. 67, no. 5, p. 1277-1286.
- MISHRA M., DUBEY, N., TOTEY, SM, BHAT, KV., BABU, S., AWASTHI-KALIA, M. and ANANDA, RK., 2002. Phylogenetic relationships and genetic polymorphisms in wild Indian mice. Biomol. Engineering, vol. 18, no. 6, p. 281-288.
- MOURÃO, G., COUTINHO, M., MAURO, R., CAMPOS, Z., TOMÁS, W. and MAGNUSSON, W., 2000. Aerial surveys of caiman, marsh deer and pampas deer in the Pantanal Wetland of Brazil. Biol. Conserv, vol. 92, no. 2, p. 175-183.
- NEI, M., 1987. Molecular evolutionary genetics New York, Columbia University Press, 512p.
- NEI, M. and LI, WH., 1979. Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc. Natl. Acad. Sci. USA, vol. 76, no. 10, p. 5269-5273.
- NEVEU, H., HAFEN, T., ZIMMERMANN, E. and RUMPLER, Y., 1998. Comparison of the genetic diversity of wild and captive groups of Microcebus murinus using the random amplified polymorphic DNA method. Folia Primatol, vol. 69, suppl. 1, p. 127-135.
- NUNES ALV., GASPARINI, RL., DUARTE, JMB., PINDER, L. and BUSCHINELLI, MC., 1997. Captura, Contenção e Manuseio. In DUARTE, JMB. (ed.). Biologia e Conservação de Cervídeos Sul-Americanos: Blastocerus, Ozotoceros e Mazama, Jaboticabal, Funep, p. 142-170.
- PEREIRA, RJG., DUARTE, JMB. and NEGRÃO, JA., 2005. Seasonal changes in fecal testosterone concentrations and their relationship to the reproductive behavior, antler cycle and grouping patterns in free-ranging male Pampas deer (Ozotoceros bezoarticus bezoarticus). Theriogenology, vol. 63, no. 8, p. 2113-2125.
- PEREZ-SWEENEY, BM., RODRIGUES, FP. and MELNICK, DJ., 2003. Metodologias moleculares utilizadas em genética da conservação. In CULLEN Jr., L., RUDRAN, R., and VALLADARES-PÁDUA, C. (eds.). Manual Brasileiro em Biologia da Conservação Curitiba, Editora da Universidade Federal do Paraná e Fundação O Boticário de Proteção à Natureza, p. 343-380.
- PETROSYAN, VG., TOKARSKAYA, ON., DANILKIN, AA. and RYSKOV, AP., 2002. Quantitative analysis of genetic parameters in populations of European (Capreolus Capreolus L.) and Siberian (Capreolus pygargus Pall.) Roe Deer with RAPD markers. Russian J. Genet, vol. 38, no. 6, p. 676-683.
- PINDER, L., 1992. Comportamento social e reprodutivo dos veados campeiro e catingueiro. Anais de etologia, vol. 10, p. 167-173.
- PRUGNOLLE, F. and MEEUS, T., 2002. Inferring sex-biased dispersal from population genetic tools: a review. Heredity, vol. 88, no. 3, p. 161-165.
- RATNAYEKE, S., TUSKAN, GA. and PELTON, MR., 2002. Genetic relatedness and female spatial organization in a solitary carnivore, the raccoon, Procyon lotor. Mol. Ecol, vol. 11, no. 6, p. 1115-1124.
- RODRIGUES, FHG., 1996. História natural e biologia comportamental do veado campeiro em cerrado do Brasil Central (Dissertação de Mestrado) UNICAMP, Campinas, SP.
- -, 2003. Estimating Pampas Deer Population in Emas National Park, Brazil. Deer Specialist Group News, no. 18, p. 10-12.
- SAMBROOK, J. and RUSSEL, DW., 2001. Molecular cloning: a laboratory manual 3rd ed. New York, Cold Spring Harbor Laboratory Press.
- SCANDURA, M., TIEDEMANN, R., APOLLONIO, M. and HARTL, GB., 1998. Genetic variation in an isolated Italian population of fallow deer Dama dama as revealed by RAPD-PCR. Acta Theriol, suppl. 5, p. 163-169.
- SHIUE, YL., BICKEL, LA., CAETANO, AR., MILLON, LV., CLARK, RS., EGGLESTON, ML., MICHELMORE, R., BAILEY, E., GUÉRIN, G., GODARD, S., MICKELSON, JR., VALBERG, SJ., MURRAY, JD. and BOWLING, AT., 1999. A synteny map of the horse genome comprised of 240 microsatellite and RAPD markers. Animal Genet, vol. 30, no. 1, p. 1-9.
- SPIRIDONOVA, LN., CHELOMINA, GN., MORIWAKI, K., YONEKAWA, H. and BOGDANOV, AS., 2004. Genetic and taxonomic diversity of the house mouse Mus musculus from the Asian part of the former Soviet Union. Russian J. Genet, vol. 40, no. 10, p. 1134-1143.
- TAMURA, K., DUDLEY, J., NEI, M., and KUMAR, S., 2007. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) Software Version 4.0. Mol. Biol. Evol, vol. 24, no. 8, p. 1596-1599.
- THORNBACK, J. and JENKINS, M., 1982. The IUCN mammal red data book Switzerland, IUCN, 560p.
- TOKARSKAYA, ON., EFREMOVA, DA., KAN, NG., DANILKIN, AA., SEMPERE, A., SEMYENOVA, SK., PETROSYAN, VG. and RYSKOV, AP., 2000. Variability of multilocus DNA markers in populations of Siberian (Capreolus pygargus Pall.) and European (C. capreolus L.) roe deer. Russian J. Genet, vol. 36, no. 11, p. 1278-1287.
- TOMAS, WM., 1995. Seasonality of antler cycle of pampas deer (Ozotoceros bezoarticus leucogaster) from the Pantanal wetland, Brazil. Stud. Neotrop. Fauna Environ, vol. 30, no. 3, p. 227-233.
- TONNI, EP., ALBERDI, MT., PRADO, JL., BARGO, MS. and CIONE, AL., 1992. Changes of mammal assemblages in the Pampean region (Argentina) and their relation with the Plio-Pleistocene boundary. Palaeogeogr Palaeocl, vol. 95, nos. 3-4, p. 179-194.
- TSVIRKA, MV., CHELOMINA, GN. and KORABLEV, VP., 2006. Genetic evidence of hybridization between Paletailed Spermophilus pallidicauda Satunin, 1903 and Alashanic S. alaschanicus Büchner, 1888 Ground Squirrels in Mongolia. Russian J. Genet, vol. 42, no. 4, p. 421-428.
- VUCETICH, LM., VUCETICH, JA., JOSHI, CP., WAITE, TA. and PETERSON, RO., 2001. Genetic (RAPD) diversity in Peromyscus maniculatus populations in a naturally fragmented landscape. Mol. Ecol, vol. 10, no. 1, p. 35-40.
- WEBER, M. and GONZÁLEZ, S., 2003. Latin American deer diversity and conservation: a review of status and distribution. Ecoscience, vol. 10, no. 4, p. 443-454.
- WELSH, J. and McCLELLAND, M., 1990. Fingerprinting genomes using PCR with arbitrary primers. Nucleic Acids Res, vol. 18, no. 24, p. 7213-7218.
- WEMMER, C., 1998. Deer: Status survey and conservation action plan (IUCN/SSC Action Plans for the Conservation of Biological Diversity). Oxford, World Conservation Union, 112p.
- WILLIAMS, JGK., KUBELIK, AR., LIVAK, KJ., RAFALSKI, JA. and TINGEY, SV.,1990. DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res, vol. 18, no. 22, p. 6531-6535.
- WU, XB., LIU, H. and JIANG, ZG., 2006. Identification primers for sika deer (Cervus nippon) from a sequence-characterized amplified region (SCAR). New Zealand J. Zool, vol. 33, no. 1, p. 65-71.
Publication Dates
-
Publication in this collection
11 Feb 2008 -
Date of issue
Dec 2007
History
-
Accepted
10 Oct 2007 -
Received
31 May 2007