Abstract
ALT-C, an ECD motif (glutamic acid, cysteine, aspartic acid) disintegrin from Bothrops alternatus snake venom, induces α2β1 integrin-mediated signaling and neutrophil chemotaxis. In vitro, in human umbilical vein endothelial cells (HUVEC), ALT-C induces cell proliferation, thus showing an interesting potential for tissue regeneration studies. This work aimed to evaluate the influence of ALT-C in myoblast viability and differentiation. Myoblasts were obtained from hind limb muscles of 3 to 4-day old Wistar rats. The cells were incubated with ALT-C at different concentrations and incubation periods were followed by total RNA isolation. cDNA synthesis and real time polymerase chain reaction (PCR) were performed with primers of myoD as well as of both (slow and fast) myosin heavy chain isoforms (MHC). ECD-disintegrin increased myoblast viability in a dose-dependent way, mostly with 50 to 100 nM concentrations, and such effect was more prevalent after 48 hours. No changes in gene expression of both MHC isoforms were observed in ALT-C-treated cells. MyoD expression was not detected, which suggests that myoblasts were in mature stages. Protease activity and cytokine array tested in a medium of 50 nM ALT-C-treated cells after 48 hours were not different from controls. In conclusion, it was shown that myoblats are sensitive to ALT-C indicating an integrin-mediated intracellular signaling that increases cell viability.
viability; myosin heavy chain; myoblast; disintegrin; skeletal muscle; Bothrops alternatus
ORIGINAL PAPER
ALT-C, a disintegrin-like Cys-rich protein from Bothrops alternatus, increases skeletal myoblast viability
Mesquita-Ferrari RAI; Moraes CK de II; Micocci KCII; Selistre-de-Araújo HSII
IDepartment of Physiotherapy, Nove de Julho University, UNINOVE, São Paulo, São Paulo State, Brazil
IIDepartment of Physiological Science, Federal University of São Carlos, São Carlos, São Paulo State, Brazil
Correspondence to Correspondence to: Raquel Agnelli Agnelli Mesquita-Ferrari Departamento de Fisioterapia, Mestrado em Ciências da Reabilitação Universidade Nove de Julho, UNINOVE, Centro de Pós-graduação Av. Francisco Matarazzo, 612 São Paulo, SP, 05001-100, Brasil Phone: +55 11 3665 9325 Email: raquel.mesquita@gmail.com
ABSTRACT
ALT-C, an ECD motif (glutamic acid, cysteine, aspartic acid) disintegrin from Bothrops alternatus snake venom, induces α2β1 integrin-mediated signaling and neutrophil chemotaxis. In vitro, in human umbilical vein endothelial cells (HUVEC), ALT-C induces cell proliferation, thus showing an interesting potential for tissue regeneration studies. This work aimed to evaluate the influence of ALT-C in myoblast viability and differentiation. Myoblasts were obtained from hind limb muscles of 3 to 4-day old Wistar rats. The cells were incubated with ALT-C at different concentrations and incubation periods were followed by total RNA isolation. cDNA synthesis and real time polymerase chain reaction (PCR) were performed with primers of myoD as well as of both (slow and fast) myosin heavy chain isoforms (MHC). ECD-disintegrin increased myoblast viability in a dose-dependent way, mostly with 50 to 100 nM concentrations, and such effect was more prevalent after 48 hours. No changes in gene expression of both MHC isoforms were observed in ALT-C-treated cells. MyoD expression was not detected, which suggests that myoblasts were in mature stages. Protease activity and cytokine array tested in a medium of 50 nM ALT-C-treated cells after 48 hours were not different from controls. In conclusion, it was shown that myoblats are sensitive to ALT-C indicating an integrin-mediated intracellular signaling that increases cell viability.
Key words: viability, myosin heavy chain, myoblast, disintegrin, skeletal muscle, Bothrops alternatus.
INTRODUCTION
Function and maintenance of tissue integrity depend on specific interactions of cells with the surrounding extracellular matrix (ECM). Transmembrane receptors are involved in the polymerization and assembly of the matrix as well as in providing a mechanical connection with the cytoskeleton and a way of transducing signals from the extracellular matrix to the nucleus (1).
Integrins form a family of cell surface adhesion receptors, mediating both cell-to-cell and cell-to-matrix interactions. They are heterodimeric transmembrane glycoproteins that consist of α and β chains, non-covalently associated (2, 3). At least 18 α and eight β chains have already been identified, which differently combined form about 24 distinct dimers (1, 4).
Studies carried out with birds have indicated that β1 integrins are involved in cell migration from the somite and terminal differentiation of myoblasts into myotubes, and that particular integrins are expressed in different stages of muscular development and differentiation (5). The integrins α5β1 (fibronectin receptor), α6β1 and α7β1 (laminin receptor) are widely expressed during development and down-regulated after myotube formation, whereas α7β1 is mainly restricted to skeletal and cardiac muscles and is strongly up-regulated during myoblast fusion. The role of α5β1 and α6β1 in muscle development and the reason they coexist at the myoblast stage as ligand-opposing receptors have not bell well characterized yet (1).
A family of myogenic transcription factors, expressed at precise steps during development, coordinates the process of myogenic differentiation. Myoblasts start to express early specific markers known as muscle regulatory factors (MRF) including desmin, Myf-5 and MyoD while the expression of myogenin, muscle regulatory factor 4 (MRF4) and myosin occurs later (6-9). However, the expression of these factors is also regulated by extracellular signaling molecules from the ECM and other elements involved in cell-to-cell contact (8).
Although myosin heavy chain (MHC) is known to be closely related to muscle function, it consists of several isoforms (10, 11). Innervation or cellular signaling affects myosin expression; however, fibers can express the slow myosin in early phases independently of innervation (12, 13). Innervation-independent variation in muscle fiber phenotype based on individual myoblasts gave rise to the concept that intrinsic, cell line-dependent mechanisms yielded muscle fibers expressing contractile protein genes in fiber type specific patterns (10). Furthermore, when skeletal muscle myoblasts were mechanically stretched, cell growth was facilitated and mRNA expression in MHC 2b, 2d and 2a increased with rapid, medium and slow stretching, respectively (10, 11).
Disintegrins were first described in 1989 as a protein group with low molecular weight (5 to 9 kDa), that interact with integrin receptors on the cell surface (14-16). These peptides represent a family of cysteine-rich proteins, isolated from snake venoms, and are known to inhibit cell-to-matrix and cell-to-cell interactions mediated by integrins (2, 17). Most disintegrins contain a RGD/KGD sequence within a hairpin loop maintained by disulfide bonds and are very potent inhibitors of aIIbb3 integrin-dependent platelet aggregation as well as cell-to-matrix interactions involved in tumor cell metastasis and angiogenesis (2, 18, 19).
A different class of disintegrins, called disintegrin-like, is also found in snake venoms and does not contain the RGD motif. These proteins are larger than RGD disintegrins (about 30 kDa) and have an extra C-terminal, cysteine-rich domain. Additionally, they do not bind to αIIbβ3, α5β1 or αvβ3 integrins, but interact with the collagen receptor, α2β1 integrin, therefore inhibiting cell adhesion to collagen I. The D/ECD sequence replaces the RGD motif, and it has been suggested that this sequence is involved in integrin binding (20).
ALT-C, an ECD-disintegrin isolated from Bothrops alternatus venom, is synthesized as a precursor form with a metalloproteinase domain from which it is released after proteolytic processing, yielding a form with both disintegrin and cysteine rich domains. ALT-C induces integrin-mediated signaling and chemotaxis in neutrophils and also strongly leads to human vein endothelial cells (HUVEC) proliferation in vitro by up-regulating the expression of some growth factors, including vascular endothelial growth factor (VEGF) (2, 18, 20, 21).
There is considerable interest in skeletal muscle regeneration studies to improve efficiency of repair in sports medicine, after severe injuries or muscle transplantation, in muscular dystrophies and for recovery strength in disuse or space flight atrophy (22). The use of integrin-binding molecules that up-regulate the expression of growth factors may be useful in these studies. This is the first demonstration of the influence of a disintegrin on myoblast proliferation and myosin gene expression.
MATERIALS AND METHODS
Protein Purification
ALT-C was purified from Bothrops alternatus venom by two steps of gel filtration followed by anion exchange chromatography (16). The molecular mass of the purified protein was estimated in 29 kDa by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE). Protein concentration was estimated according to the method of Bradford (23).
Cell Culture
Myoblasts were obtained from limb muscles of 3 to 4-day old Wistar rats, according to Horn and Brodwick (24). Rats were killed and limb muscles were removed, cut into small pieces and homogenized by trituration for 10 minutes with shears. After that, tissue was submitted to an enzymatic digestion with tripsin (0.025% in PBS buffer) and collagenase V (0.1% in PBS buffer) for 40 minutes at 37° C. Then, cells were cultivated in a 10 cm plastic dish with Dubelcco's minimum medium (DMEM Cultilab, Brazil) supplemented with 10% fetal bovine serum, 1% L-glutamine, 1% antibiotic-antimycotic solution (Cultilab, Brazil) at 37° C, with 5% CO2 for 20 minutes to remove other cell types and proteins such as fibroblasts, endothelial cells and ptoteoglicans. Afterwards, cells were transferred to a culture flask containing DMEM supplemented as described before.
Cell Viability Assay
Myoblasts (105 cells/well) were incubated in a 96-well plate in 200µL of DMEM with ALT-C at different concentrations at 37° C and 5% CO2 for 24, 48 and 72 hours. After incubation time, media from cultures were saved for latter protein analysis. Cell concentration was measured by addition of a tetrazolium salt MTT [1-(4,5-dimethylthiazol-2-yl)-3,5-diphenyl formazan, Sigma, USA], 0.5 mg/mL final concentration followed by incubation at 37ºC and 5% CO2 for three hours. The bound stain was dissolved in isopropyl alcohol and measured spectrophotometrically at 595 nm. All experiments were repeated three times, and each sample was made in triplicate. Some experiments were performed in collagen I-coated plates (1 µg/well). Mean, standard deviation and one-way analysis of variance (ANOVA) were employed for data analysis. Statistical significance was assessed by the Dunnett's test and acceptable p levels were less than 0.05 (25).
Detachment Assay
Myoblasts (105 cells/well) were incubated in collagen I-coated 96-well plates in 200 µL of adhesion buffer [20 mM Hepes (Sigma, USA), 150 mM NaCl, 5 mM MgCl2, 0.25 mM MnCl2, pH 7.4] at 37° C for four hours. Subsequent to incubation, cells were washed five times with adhesion buffer, then ALT-C in adhesion buffer was added at different concentrations and cells were incubated at 37° C and 5% CO2 for 48 hours. Medium from cultures was removed, cells were washed again and cell concentration was measured by adding MTT (0.5 mg/mL, final concentration) followed by incubation at the same conditions for three hours. Data analysis was performed as previously described.
Isolation of Total RNA
Cells (106 cells/well) were incubated in 10-cm plastic dishes in DMEM medium with or without 50 nM ALT-C at 37° C and 5% CO2 for 24, 48 and 72 hours. Culture medium was removed and cells were lysed with cold TRIzol® reagent (Invitrogen Carlsbad, USA) for total RNA isolation, according to the manufacturer instructions. Total RNA was quantified by spectrophotometry and RNA samples were treated with DNAse (Invitrogen Carlsbad, USA) to avoid contamination with genomic DNA. All solutions were prepared with 0.01% diethyl pyrocarbonate-treated water (DEPC Sigma, USA), while glassware and plasticware were treated against RNase using standard procedures.
cDNA Synthesis and Real Time PCR
cDNA synthesis was performed using the AMV reverse transcriptase (Access Quick RT-PCR System® (Promega, USA) and real time PCR was accomplished using the SYBR Green PCR® Master Mix (Applied Biosystems, USA) in a Rotor-Gene RG 3000 (Corbett Research, Australia). All samples were analyzed in duplicate.
Oligonucleotide Primers
Specific primers for MHC I isoform (GenBank, accession n. X15939) and MHC IIa were used for real time PCR (26). The primer sequences were:
-
MHC I forward: 5' AGA GAA TGG CAA GAC GGT GAC T; reverse: 5' CAT GTC CTC GAT CTT GTC GAA CT (82 bp amplicon);
-
MHC IIa foward: 5 TAT CCT CAG GCT TCA AGA TTT G; reverse: 5' TAA ATA GAA TCA CAT GGG GAC A (310 bp amplicon);
-
MyoD forward: 5 GGA GAC ATC CTC AAG CGA TGC; reverse: 5' AGC ACC TGG TAA ATC GGA TTG.
A constitutive gene, β-actin, was emplyed to normalize data using the same amount of cDNA. β-actin primers were forward 5' CGTGGGCCGCCCTAGGCACCAGGG and reverse 5' CGGAGGAAGAGGATGCGGCAGTGG (604 bp amplicon).
To normalize data among control and treated groups, arbitrary units were calculated as follows, Arbitrary Unit = 2 ΔΔTC, and ΔΔTC = sample ΔTC control ΔTC (TC, threshold cycle).
Protease Activity and VEGF Expression
After incubation time, the cell medium was removed and tested for protease activity by zymography, carried out as described by Allen et al (27) with small modifications. For enzymatic assays, SDS-gels (15%) were prepared with gelatin (1 mg/mL) and 30 µg of protein was loaded per lane in sample buffer without reduction. After electrophoresis, gels were washed twice for 15 minutes each with 2.5% Triton X-100® (Sigma, USA) to eliminate SDS. Gels were then incubated overnight at 37° C in substrate buffer (50 mM Tris-HCl, pH 8.5, 5 mM CaCl2, 0.02% NaN3). After this, gels were stained for 30 minutes in 0.05% Coomassie Brilliant Blue R-250® (Sigma, USA) in acetic acid:methanol:water (1:4:5) and destained in the same solution. All gels were prepared and run at the same time. The bands were quantified by densitometry using the Image Pro-plus software (Media Cybernetics, USA). ELISA assays for VEGF detection were performed using the Quantikine® immunoassay kit for human VEGF (R&D Systems, USA) according manufacturer instructions. Briefly, 200 µL of conditioned media were added to each well, previously coated with human monoclonal anti-VEGF antibody. After two hours of incubation, wells were washed and incubated with an enzyme-linked polyclonal anti-VEGF antibody. Following another washing, substrate solution was added to wells and color developed in proportion to the amount of VEGF bound in the initial step. The plate was read on a Dynex plate reader (USA) at 450 nm.
Cytokine Array
The expression of cytokines was analyzed using the incubation medium of control cells (without Alt-C) and treated cells (with Alt-C 50 nM), after 48 hours, using the RayBio® human cytokine antibody array kit (R&D Systems, USA). The analyzed cytokines were interleukins (IL-1α, IL-2, IL-3, IL-5, IL-6, IL-7, IL-8, IL-10, IL-13, IL-15); tumor growth factor (TGF-β1); tumor necrosis factors (TNF-α, TNF-β); interferons (IFN- γ, IFN-α, IFN-β); granulocyte colony-stimulating factor (GCSF); granulocyte-monocyte colony-stimulating factor (GM-CSF); genes GRO and GRO-a; regulated upon activation, normal T-cell expressed and secreted (RANTES); monokine induced by interferon-gamma (MIG); macrophage/monocyte chemotactic proteins (MCP-1, MCP-2, MCP-3).
RESULTS
ALT-C Induces Myoblast Viability
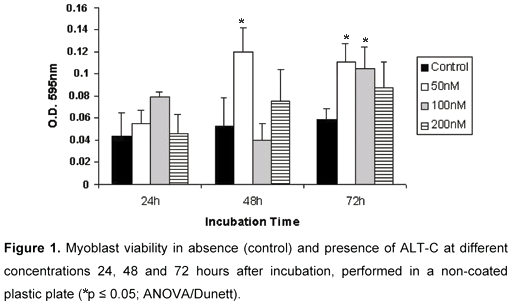
There was no significant difference of cell number in ALT-C treated myoblasts when compared with control values 24 hours after incubation in the tested concentrations (Figure 1). However, ALT-C induced a significant increase in viability of myoblasts at 50 nM concentration, 48 and 72 hours after incubation. Myoblast viability was also induced by ALT-C at 100 nM, but only after 72 hours. These effects were not observed with higher concentrations, such as 200 nM, which presented values not significantly different from controls (Figure 1).
For cell growing on collagen-coated plates, similar results were found. ALT-C also improved myoblast viability; however, higher concentrations, around 100 and 150 nM, were required (Figure 2). Furthermore, ALT-C did not provoke myoblast detachment when it was added to a culture after myoblast adhesion (results not shown).
Influence of ALT-C on Myosin Heavy Chain Expression
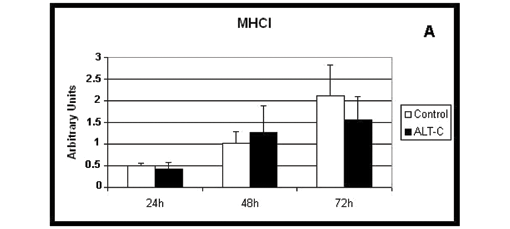
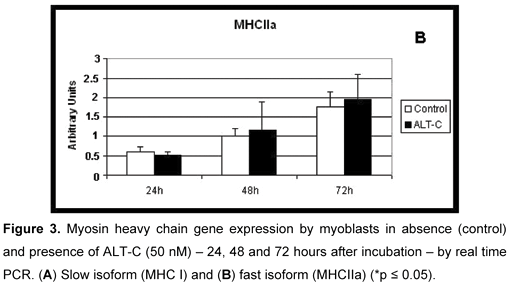
Both myosin slow (MHCI, Figure 3 A) and fast (MHCIIa, Figure 3 B) isoforms were expressed in all conditions: absence (control) and presence of ALT-C in all incubation periods and no differences were observed when ALT-C was used.
Protease, VEGF and Cytokine Expression
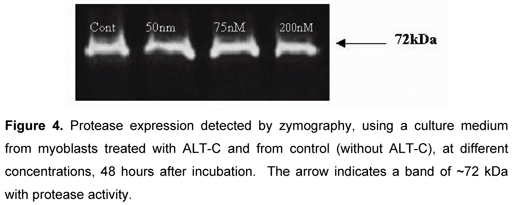
Protease activity was detected in the culture medium of myoblast as a single ~72 kDa lytic band (Figure 4). This activity was completely abolished when gels were incubated in substrate buffer with 15 mM EDTA (not shown), suggesting the presence of a gelatinase. No significant differences were observed between ALT-C-treated samples and controls. VEGF was not detected in the medium culture of both control and treated cells (results not shown). There was no difference in cytokine expression between control and treated cells (Figure 5). After membrane incubation in the culture medium, only positive controls were marked.
DISCUSSION
It was previously demonstrated that ALT-C, an α2β1-integrin ligand, induces VEGF expression in fibroblasts and endothelial cell viability in doses as low as 10 nM (19). Therefore, we found interesting to study the effect of this disintegrin on other cell types. A striking observation is that ALT-C induced myoblast viability in a dose-dependent way, mostly in concentrations ranging from 50 to 100 nM. Interestingly, in the presence of collagen I, increased protein concentrations were require to provoke similar results (150 nM). In addition, this effect was also influenced by incubation time and was more consistent after 48 hours. These results suggest that α2β1 integrin may be relevant for muscle cell viability. Moreover, ALT-C did not induce myoblast detachment as similarly observed in fibroblasts and HUVEC (19).
Cells express distinct sets of muscle-specific proteins at different stages of the myogenic pathway. Desmin, Myf-5 and MyoD are expressed at relatively early stages, whereas myogenin, myogenin regulatory factor 4 (MRF4) and myosin are expressed at later stages. The cells utilized in the current study expressed both MHC I and IIa isoforms of myosin, but not MyoD. These characteristics allow us to conclude that they belong to later differentiation stages (6-9, 28, 29).
It has been reported that myoblasts in their mature stage, as myotube, express mostly slow myosin, while the fast form can be found especially after innervation or under signaling conditions (10, 11). The present data showed that myosin expression occurred in myoblast cells and both isoforms, slow and fast, were expressed in absence of innervation. Furthermore, there was no evidence of ALT-C influence in the expression of these isoforms, since their values were similar in the presence and absence of this disintegrin in all periods of cell incubation.
ALT-C was reported to induce in vitro VEGF expression by fibroblasts, but not by HUVEC (18). However, in the present study VEGF expression was not detected either at mRNA or protein level, suggesting that this effect may be cell-specific. It also implies that the primary cultures were not contaminated with fibroblasts. We could not determine, by the assays accomplished, which factors or pathways could be involved in myoblast intracellular signaling induced by ALT-C. More studies, such as cDNA arrays, are necessary to address these relevant questions for a better understanding of the action mechanism of this desintegrin.
Disintegrin-related proteins (ADAM a disintegrin and metalloproteinase), with a similar domain organization, are found in mammals as well as in several other organisms, in which they are involved in numerous physiological processeses such as fertilization, cell differentiation and shedding of receptors (30). It has been demonstrated that ADAM disintegrin and Cys-rich domains may also trigger integrin-mediated intracellular signaling events and cell responses (31). Given the homology between disintegrins from snake venoms and ADAM domains, it is reasonable to suggest that the disintegrin-like domain of some ADAM may have a role in myoblast viability.
In conclusion, our results demonstrated that ALT-C did not modify myoblast MHC expression but induced an increase in cell viability. Therefore, ALT-C may be very valuable for myogenesis and skeletal muscle regeneration studies.
ACKNOWLEDGEMENTS
This work was supported by grants from FAPESP (98/14138-2; 04/09671-6) and CNPq (350111/1998-7), Brazil.
Received: August 26, 2008
Accepted: September 17, 2008
Abstract published online: September 26, 2008
Full paper published online: May 30, 2009
CONFLICTS OF INTEREST: There is no conflict
FINANCIAL SOURCE: FAPESP and CNPq
- 1. Mayer U. Integrins: redundant or important players in skeletal muscle? J Biol Chem. 2003;278(17):14587-90.
- 2. Mariano-Oliveira A, Coelho AL, Terruggi CH, Selistre-de-Araújo HS, Barja-Fidalgo C, De Freitas MS. Alternagin-C, a non RGD-disintegrin, induces neutrophil migration via integrin signaling. Eur J Biochem. 2003;270(24):4799-808.
- 3. Niewiarowski S, McLane MA, Kloczewiak M, Stewart GJ. Disintegrins and other naturally occurring antagonists of platelet fibrinogen receptors. Semin Hematol. 1994;31(4):289-300.
- 4. Humphries JD, Byron A, Humphries MJ. Integrin ligands at a glance. J Cell Sci. 2006;119(Pt 19):3901-3.
- 5. Velleman SG, McFarland DC. β1 integrin mediation of myogenic differentiation: implications for satellite cell differentiation. Poult Sci. 2004;83(3):245-52.
- 6. Cabane C, Englaro W, Yeow K, Ragno M, Dérijard B. Regulation of C2C12 myogenic terminal differentiation by MKK3/p38α pathway. Am J Physiol Cell Physiol. 2003;284:658-66.
- 7. Dominov JA, Dunn JJ, Miller JB. Bcl-2 expression identifies an early stage of myogenesis and promotes clonal expansion of muscle cells. J Cell Biol. 1998;142(2):537-44.
- 8. Krauss RS, Cole F, Gaio U, Takaesu G, Zang W, Kang JS. Close encounters: regulation of vertebrate skeletal myogenesis by cell-cell contact. J Cell Sci. 2005;118:2355-62.
- 9. Tannu NS, Rao VK, Chaudhary RM, Giorgianni F, Saeed AE, Gao Y, Raghow R. Comparative proteomes of the proliferating C(2)C(12) myoblasts and fully differentiated myotubes reveal the complexity of the skeletal muscle differentiation program. Mol Cell Proteomics. 2004;3(11):1065-82.
- 10. Kurokawa K, Abe S, Sakiyama K, Takeda T, Ide Y, Ishigami K. Effects of stretching stimulation with different rates on the expression of MyHC mRNA in mouse cultured myoblasts. Biomed Res. 2007;28(1):25-31.
- 11. Sakiyama K, Abe S, Tamatsu Y, Ide Y. Effects of stretching stress on the muscle contraction proteins of skeletal muscle myoblasts. Biomed Res. 2005;26(2):61-8.
- 12. Jordan T, Jiang H, Li H, DiMario JX. Inhibition of ryanodine receptor 1 in fast skeletal muscle fibers induces a fast-to-slow muscle fiber type transition. J Cell Sci. 2004;117(Pt 25):6175-83.
- 13. Miller JB, Crow MT, Stockdale FE. Slow and fast myosin heavy chain content defines three types of myotubes in early muscle cell cultures. J Cell Biol. 1985;101(5 Pt 1):1643-50.
- 14. Beviglia L, Stewart GJ, Niewiarowski S. Effect of four disintegrins on the adhesive and metastatic properties of B16F10 melanoma cells in a murine model. Oncol Res. 1995;7(1):7-20.
- 15. Gould RJ, Polokoff MA, Friedman PA, Huang TF, Holt JC, Cook JJ, Niewiarowski S. Disintegrins: a family of integrin inhibitory proteins from viper venoms. Proc Soc Exp Biol Med. 1990;195(2):168-71.
- 16. Souza DHF, Iemma MRC, Ferreira LL, Faria JP, Oliva MLV, Zingali RB, Niewiarowski S, Selistre-de-Araújo HS. The disintegrin-like domain of the snake venom metalloprotease alternagin inhibits α2β1 integrin-mediated cell adhesion. Arch Biochem Biophys. 2000;384(2):341-50.
- 17. Calvete JJ, Moreno-Murciano MP, Theakston RD, Kisiel DG, Marcinkiewicz C. Snake venom disintegrins: novel dimeric disintegrins and structural diversification by disulphide bond engineering. Biochem J. 2003;372(Pt 3):725-34.
- 18. Cominetti MR, Terruggi CH, Ramos OH, Fox JW, Mariano-Oliveira A, De Freitas MS, Figueiredo CC, Morandi V, Selistre-de-Araujo HS. Alternagin-C, a disintegrin-like protein, induces vascular endothelial cell growth factor (VEGF) expression and endothelial cell proliferation in vitro J Biol Chem. 2004;279(18):18247-55.
- 19. Kamiguti AS, Hay CRM, Zuzel M. Inhibition of collagen-induced platelet aggregation as the result of cleavage of α2β1 integrin by the snake venom metalloproteinase jararhagin. Biochem J. 1996;320(pt2):635-41.
- 20. Selistre-de-Araújo HS, Cominetti MR, Terruggi CHB, Mariano-Oliveira A, De Freitas MS, Crepin M, Figueiredo CC, Morandi V. Alternagin-C, a disintegrin-like protein from the venom of Bothrops alternatus, modulates α2β1 integrin-mediated cell adhesion, migration and proliferation. Braz J Med Biol Res. 2005;38(10):1505-11.
- 21. Ramos OH, Selistre-de-Araujo HS. Snake venom metalloproteases structure and function of catalytic and disintegrin domains. Comp Biochem Physiol C Toxicol Pharmacol. 2006;142(3-4):328-46.
- 22. Grounds MD, White JD, Rosenthal N, Bogoyevitch MA. The role of stem cells in skeletal and cardiac muscle repair. J Histochem Cytochem. 2002;50(5):589-610.
- 23. Bradford MM. A rapid and sensitive method for the quantization of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976;72:248-54.
- 24. Horn R, Brodwick MS. Acetylcholine-induced current in perfused rat myoballs. J Gen Physiol. 1980;75(3):297-321.
- 25. Löster K, Horstkorte R. Enzymatic quantification of cell-matrix and cell-cell adhesion. Micron. 2000;31(1):41-53.
- 26. Jaschinski F, Schuler M, Peuker H, Pette D. Changes in myosin heavy chain mRNA and protein isoforms of rat muscle during forced contractile activity. Am J Physiol Cell Physiol. 1998;43(2):365-70.
- 27. Allen DL, Teitelbaum DH, Kurachi K. Growth factor stimulation of matrix metalloproteinase expression and myoblast migration and invasion in vitro Am J Physiol Cell Physiol. 2003;284:805-15.
- 28. Di Carlo A, De Mori R, Martelli F, Pompilio G, Capogrossi MC, Germani A. Hypoxia inhibits myogenic differentiation through accelerated MyoD degradation. J Biol Chem. 2004;279(16):16332-8.
- 29. Dogra C, Hall SL, Wedhas N, Linkhart TA, Kumar A. Fibroblast growth factor inducible 14 (Fn14) is required for the expression of myogenic regulatory factors and differentiation of myoblasts into myotubes. Evidence for TWEAK-independent functions of Fn14 during myogenesis. J Biol Chem. 2007;282(20):15000-10.
- 30. Blobel CP. ADAMs: key components in EGFR signaling and development. Nat Rev Mol Cell Biol. 2005;6(1):32-43.
- 31. Eto K, Huet C, Tarui T, Kupriyanov S, Liu HZ, Puzon-McLaughlin W, Zhang XP, Sheppard D, Engvall E, Takada Y. Functional classification of ADAMs based on a conserved motif for binding to integrin α9β1: implications for sperm-egg binding and other cell interactions. J Biol Chem. 2002;277(20):17804-10.
Correspondence to:
Publication Dates
-
Publication in this collection
10 June 2009 -
Date of issue
2009
History
-
Accepted
17 Sept 2008 -
Received
26 Aug 2008