Abstract
Pseudomonas aeruginosa infections cause significant mortality and morbidity in health care settings. Strategies to prevent and control the emergence and spread of P. aeruginosa within hospitals involve implementation of barrier methods and antimicrobial stewardship programs. However, there is still much debate over which of these measures holds the utmost importance. Molecular strain typing may help elucidate this issue. In our study, 71 nosocomial isolates from 41 patients and 23 community-acquired isolates from 21 patients were genotyped. Enterobacterial repetitive intergenic consensus-polymerase chain reaction (ERIC-PCR) was performed. Band patterns were compared using similarity coefficients of Dice, Jaccard and simple matching. Strain similarity for nosocomial strains varied from 0.14 to 1.00 (Dice); 0.08 to 1.00 (Jaccard) and 0.58 to 1.00 (simple matching). Forty patterns were identified. In most units, several clones coexisted. However, there was evidence of clonal dissemination in the high risk nursery, neurology and two surgical units. Each and every community-acquired strain produced a unique distinct pattern. Results suggest that cross transmission of P. aeruginosa was an uncommon event in our hospital. This points out to a minor role for barrier methods in the control of P. aeruginosa spread.
Pseudomonas aeruginosa; strain typing; ERIC-PCR; nosocomial infections; health care infections
ORIGINAL PAPER
Polyclonal endemicity of Pseudomonas aeruginosa in a teaching hospital from Brazil: molecular typing of decade-old strains
Fortaleza CMCBI; Bacchi CEII; Oliveira DEII; Ramos MCIII
IDepartment of Tropical Diseases and Imaging Diagnosis, Botucatu Medical School, São Paulo State University (UNESP - Univ Estadual Paulista), Botucatu, São Paulo State, Brazil
IIDepartment of Pathology, Botucatu Medical School, São Paulo State University (UNESP - Univ Estadual Paulista), Botucatu, São Paulo State, Brazil
IIISchool of Medical Sciences, State University of Campinas, UNICAMP, Campinas, São Paulo State, Brazil
Correspondence to Correspondence to: Carlos Magno Castelo Branco Fortaleza Departamento de Doenças Tropicais e Diagnóstico por Imagem Faculdade de Medicina de Botucatu, UNESP Distrito de Rubião Junior, s/n Botucatu, SP, 18618-970, Brazil Phone: +55 14 3811 6212. Fax: +55 14 3815 9898 Email: cmfortaleza@uol.com.br
ABSTRACT
Pseudomonas aeruginosa infections cause significant mortality and morbidity in health care settings. Strategies to prevent and control the emergence and spread of P. aeruginosa within hospitals involve implementation of barrier methods and antimicrobial stewardship programs. However, there is still much debate over which of these measures holds the utmost importance. Molecular strain typing may help elucidate this issue. In our study, 71 nosocomial isolates from 41 patients and 23 community-acquired isolates from 21 patients were genotyped. Enterobacterial repetitive intergenic consensus-polymerase chain reaction (ERIC-PCR) was performed. Band patterns were compared using similarity coefficients of Dice, Jaccard and simple matching. Strain similarity for nosocomial strains varied from 0.14 to 1.00 (Dice); 0.08 to 1.00 (Jaccard) and 0.58 to 1.00 (simple matching). Forty patterns were identified. In most units, several clones coexisted. However, there was evidence of clonal dissemination in the high risk nursery, neurology and two surgical units. Each and every community-acquired strain produced a unique distinct pattern. Results suggest that cross transmission of P. aeruginosa was an uncommon event in our hospital. This points out to a minor role for barrier methods in the control of P. aeruginosa spread.
Key words:Pseudomonas aeruginosa, strain typing, ERIC-PCR, nosocomial infections, health care infections.
INTRODUCTION
Pseudomonas aeruginosa is a relevant pathogen in hospitalized patients, accounting for significant morbidity and mortality (1). In a recent report, it ranked second among agents of ventilator-associated pneumonia in North American hospitals. It was also among the seven most common agents of health care-acquired urinary, blood system and surgical site infections (2). This picture is worsened by the continuous, worldwide increase of multidrug resistance among P. aeruginosa isolates from hospitalized patients (3, 4).
Despite its high incidence, several steps in P. aeruginosa epidemiology remain unknown. Its reservoirs in the hospital and mechanisms of transmission have not been fully elucidated (5). As a consequence, a debate about which measures are utmost important to control its spread within hospitals has not come to a definite conclusion (6).
In this setting, techniques of molecular epidemiology are of great value, adding precious information and helping us fulfill gaps in our present knowledge (7). The purpose of our study was to identify distribution of P. aeruginosa clones within a teaching hospital, and thus estimate the frequency of their cross-transmission among patients.
MATERIALS AND METHODS
Setting
The study was conducted in the teaching hospital of Botucatu Medical School (HC-FMB), a 400-bed facility that provides tertiary care for an area with approximately one million inhabitants. At the time the bacterial strains from our study were isolated, the hospital had four areas for critical patients: high risk nursery (HRN), pediatric intensive care unit (P-ICU), medical-surgical intensive care unit (MS-ICU) and emergency room intensive care unit (ER-ICU). There were also several units for medical, surgical and pediatric non-critical patients.
P. aeruginosa Isolates: Collection, Storage and Susceptibility Tests
P. aeruginosa isolates recovered from clinical cultures in the microbiology laboratory of HC-FMB from August to October 1998 were collected and stored at -70°C in 10% glycerol/BHI media. An automatic procedure (Microscan Autoscan-4â, Siemens, USA) was used for antimicrobial susceptibility tests.
Epidemiological Classification of Isolates
The infection control committee data files were analyzed to identify which patients harboring P. aeruginosa isolates had a previous nosocomial infection. Isolates collected from those patients, as well as those cultured from previously sterile sites more than 48 hours after hospital admission were stated as nosocomial (Group N). All other strains we classified as Group C (community acquired). Group N comprised 71 isolates from 46 inpatients. Group C consisted of 23 isolates from 21 individuals, including inpatients and outpatients.
DNA Purification
DNA was purified using a method originally described for Mycobacterium tuberculosis (8). The final product was quantified by spectrophotometry (GeneQuant IIâ, Pharmacia, USA).
Strain Typing
Enterobacterial repetitive intergenic consensus-polymerase chain reaction (ERIC-PCR) was performed as described by Versalovic et al. (9). A single primer (ERIC 2: 5'- AAG TAA GTG ACT GGG GTG AGC G - 3', Life Technologies, USA) was used. PCR solution consisted of: 100 ng of source DNA; 1.0 mM magnesium chloride (MgCl2, Gibco, USA); 200 mM dNTP mix (dATP, dTTP, dGTP e dCTP, Gibco); 100 pmol primer (ERIC-2); 2 U Taq polimerase (Amersham Pharmacia, USA) and 5 μL of reaction buffer (Amersham Pharmacia, USA). Final volume was 50 μL. PCR cycles consisted of: an initial denaturation step (95°C for five minutes) followed by 35 cycles of denaturation at 90°C (30 seconds), annealing at 52°C (one minute) and extension at 65°C (eight minutes). Final extension was held for 16 minutes. Positive (ATCC 27853) and negative controls were used in every amplification.
After amplification, 5 μL of PCR products were submitted to electrophoresis in 2% agarose (50 V, six hours). The gel was stained in ethidum bromide (0.5 μg/mL) and photographed with Polaroid (USA).
Cluster Analysis
Images were visually analyzed and converted into binary sheets. Cluster analysis was performed in free software (Treeconâ for Windows, Bioinformatics, Belgium). Similarity coefficients of Dice (SD), Jaccard (SJ) and simple matching (SSM) were calculated (10). Unweighted pair method group with arithmetic averages (UPGMA) dendrograms were drawn for simple matching similarity.
Ethical Issues
This study was fully approved by the Research Ethics Committee of Botucatu Medical School (protocol number 0394).
RESULTS
A total of 94 isolates (71 from group N and 23 from group C) were collected. Group N isolates grew predominantly from respiratory specimens (50.1%). On the other hand, wound secretions were the source of 47.8% of community-acquired isolates (Group C).
Typing of Nosocomial Strains (Group N)
Seventy-one isolates from group N produced 40 unambiguous patterns, named N1 to N40. Similarity coefficients varied from 0.14 to 1.00 (SD), 0.08 to 1.00 (SJ), and 0.58 e 1.00 (SSM) (Figure 1).
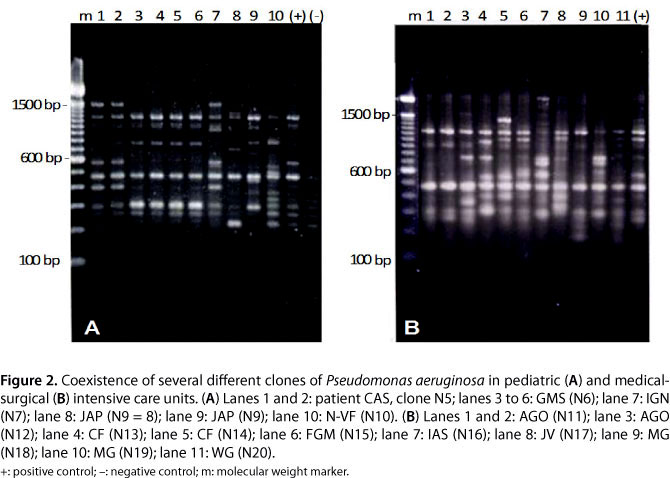
We found the coexistence of several clones in most units (Figure 2). However, some clusters with complete similarity were found:
· High risk nursery (HRN): N4 - 15 isolates from six patients (Figure 3).
· Neurology unit: N33 - five isolates from four patients.
· Neurology unit: N31- three isolates from two patients.
· Vascular surgery unit: N38 - three isolates from two patients.
· Gastric surgery unit: N23 - two isolates from two patients.
Other similarity clusters consisted of several cultures from the same patient: N5, N6, N11 and N32. We noticed some variation in antimicrobial susceptibility profiles within the same cluster (Table 1).
In six occasions, two or more different clones were cultured from the same individual. In one case there was a significant similarity between strains (N28-N29 - SD = 0.91). Mostly in these occasions, antimicrobial susceptibility patterns were quite divergent.
Typing of Community-Acquired Strains (Group C)
Each and every isolate in Group C produced a unique unambiguous pattern (Figure 3). Similarity results varied (SD: 0.20-0.86; SJ: 0.11-0.75; SSM: 0.55-0.94). In two occasions, different patterns were observed in isolates from the same patient. In both cases, the difference was only one band.
DISCUSSION
Our study did not find evidence of extensive cross transmission of nosocomial P. aeruginosa clones. Instead, multiple clones coexisted in most units. This pattern was noticed even in units dealing with critical care and advanced life support (P-ICU, CA-ICU, MS-ICU), where high occurrence of cross transmission is expected. Also, no cluster was identified among community-acquired strains.
Results from strain typing of P. aeruginosa performed during outbreaks usually report extensive dissemination of one or a few clones (11-13). On the other hand, studies that focused on endemic clones reported two different epidemiological patterns of P. aeruginosa infections. In the first, few clones infect a great number of patients (14-18). This picture points out to extensive cross transmission of P. aeruginosa strains within the hospital. However, other authors report a different pattern, in which multiple clones coexist (19-22). Those findings are coherent with ours.
The explanation for this "polyclonal endemicity" remains controversial. However, Kropec et al. (23) pointes out the gastrointestinal tract as the most important reservoir of P. aeruginosa. These authors speculate that small, undetectable community-acquired carriage becomes significant after exposure to antimicrobials and invasive procedures in the hospital setting. Otherwise, it has been suggested that strains implicated in nosocomial infections arise from water and other environmental sources (24).
In our study, clonal dissemination - probably through cross transmission - was found in four units: HRN, neurological, gastric surgery and vascular surgery. In HRN, N4 clone was isolated from six out of ten patients with positive cultures for P. aeruginosa. This clone has persisted in the unit for at least two months, showing a considerable variability in antimicrobial susceptibility patterns. Clonal dissemination of P. aeruginosa in neonatal units is expected, since newborns do not carry community-acquired strains. Other authors have described this event (25, 26). In other units, clones were also found to persist for long intervals: N23 and N33 were isolated in specimens collected in August and October.
Although it was not the main objective of this work, variation in antimicrobial susceptibility within the same clone was frequently noticed. To what extent this may be ascribed to reproducibility problems in automatic testing is a matter of concern (27). It is well known, however, that P. aeruginosa shows a strong tendency to change susceptibility patterns under environmental pressure. Several antimicrobials (some penicillins and cephalosporins) may induce the overproduction of AmpC beta-lactamase, leading to resistance to currently used anti-pseudomonal drugs (ceftazidime, cefepime). Changes in outer membrane porin proteins and expression of carbapenemases account for carbapenem resistance (28).
Overall, ERIC-PCR was useful for epidemiological studies with P. aeruginosa. As a PCR-based method, it can provide faster results than pulsed-field gel electrophoresis (PFGE) (29). In a study by Lau et al. (30), both techniques had a similar discriminatory power. Using long primers directed to known repetitive elements in the genome of gram-negative bacteria, ERIC-PCR has a greater reproducibility than methods based on arbitrary primers (31).
Contrary to PFGE, no easy method was proposed for interpretation of ERIC-PCR generated patterns (32). Cluster analysis can overcome this problem, providing numerical results that can be interpreted in the light of epidemiological findings. An interesting approach to our results is to compare clusters using three different cut-points in similarity (Table 2). A policlonal pattern results even after grouping every strain with Dice similarity above 0.75. In that case, six clusters group 24 out of 41 patients. Still, strains from different clusters coexisted in most units.
Finally, one should ask if there is anything we can learn from the typing of isolates that are more than a decade old. We believe they are valuable for understanding the epidemiology of P. aeruginosa in health care settings. In 1998, P. aeruginosa was the bacteria most frequently recovered from clinical cultures from inpatients in our hospital, accounting for 16.1% of all positive cultures. That was the peak of hyperendemicity. The fact that it was mostly polyclonal suggests that cross transmission was infrequent. These findings are consistent with the hypothesis that strain selection from inner reservoirs is the main route for P. aeruginosa colonization and infection. There is a good possibility that antimicrobial control policies, rather than isolation precautions, should be effective in reducing P. aeruginosa incidence. Further epidemiological studies are required to strengthen our findings.
ACKNOWLEDGEMENTS
We are thankful to the State of São Paulo Research Foundation (FAPESP) for the financial support.
Submission status
Received: November 16, 2010.
Accepted: March 16, 2011.
Abstract published online: March 17, 2011.
Full paper published online: May 31, 2011.
Conflicts of interest
There is no conflict.
Financial source
The State of São Paulo Research Foundation (FAPESP) provided the financial grants.
Ethics committee approval
The present study was approved by the Research Ethics Committee of Botucatu Medical School (protocol number 0394), UNESP, Brazil.
- 1. Navon-Venezia S, Ben-Ami R, Carmeli Y. Update on Pseudomonas aeruginosa and Acinetobacter baumannii infections in the healthcare setting. Curr Opin Infect Dis. 2005;18(4):306-13.
- 2. Hidron AI, Edwards JR, Patel J, Horan TC, Sievert DM, Pollock DA, et al. NHSN annual update: antimicrobial-resistant pathogens associated with healthcare-associated infections: annual summary of data reported to the National Healthcare Safety Network at the Centers for Disease Control and Prevention, 2006-2007. Infect Control Hosp Epidemiol. 2008;29(11):996-1011.
- 3. Ribeiro J, Mendes RE, Domingos R, França E, Silbert S, Jones RN, Sader HS. Microbiological and epidemiological characterization of imipenem-resistant Pseudomonas aeruginosa strains from a Brazilian tertiary hospital: report from the SENTRY Antimicrobial Surveillance Program. J Chemother. 2006;18(5):461-7.
- 4. Jones RN, Stilwell MG, Rhomberg PR, Sader HS. Antipseudomonal activity of piperacillin/tazobactam: more than a decade of experience from the SENTRY Antimicrobial Surveillance Program (1997-2007). Diagn Microbiol Infect Dis. 2009;65(3):331-4.
- 5. Jarvis WR, Sinkowitz-Cochran R. Emerging healthcare-associated problem pathogens in the United States. Postgrad Med. 2001;109(2 Suppl):3-9.
- 6. Siegel JD, Rhinehart E, Jackson M, Chiarello L; Healthcare Infection Control Practices Advisory Committee. Management of multidrug-resistant organisms in healthcare settings, 2006. Am J Infect Control. 2007;35(10 Suppl 2):S165-S93.
- 7. Jackson BR, Thomas A, Carroll KC, Adler FR, Samore MH. Use of strain typing data to estimate bacterial transmission rates in healthcare settings. Infect Control Hosp Epidemiol 2005;26(7):638-45.
- 8. Van Soolingen D. Isolation of genomic DNA from mycobacteria. In: Bunschoten A, et al. Spoligotyping a method to detect and type Mycobacterium tuberculosis complex bacteria. Bilthoven: Research Laboratory for Infectious Diseases, National Institute of Public Health and the Environment, 1996.
- 9. Versalovic J, Koeuth T, Lupski JR. Distribution of repetitive DNA sequences in eubacteria and application to fingerprinting of bacterial genomes. Nucleic Acids Res. 1991;19(24):6823-31.
- 10. Sneath PA, Sokal RR. Numerical taxonomy. San Francisco: W. H. Freeman and Company, 1973. 573 p.
- 11. Crespo MP, Woodford N, Sinclair A, Kaufmann ME, Turton J, Glover J, et al. Outbreak of carbapenem-resistant Pseudomonas aeruginosa producing VIM-8, a novel metallo-beta-lactamase, in a tertiary care center in Cali, Colombia. J Clin Microbiol. 2004;42(11):5094-101.
- 12. Pellegrino FL, Casali N, Dos Santos KR, Nouér SA, Scheidegger EM, Riley LW, et al. Pseudomonas aeruginosa epidemic strain carrying bla(SPM) metallo-beta-lactamase detected in Rio de Janeiro, Brazil. J Chemother. 2006;18(2):151-6.
- 13. Lanini S, D'Arezzo S, Puro V, Martini L, Imperi F, Piselli P, et al. Molecular epidemiology of a Pseudomonas aeruginosa hospital outbreak driven by a contaminated disinfectant-soap dispenser. PLoS One. 2011;6(2):e17064.
- 14. Mayank D, Anshuman M, Singh RK, Afzal A, Baronia AK, Prasad KN. Nosocomial cross-transmission of Pseudomonas aeruginosa between patients in a tertiary intensive care unit. Indian J Pathol Microbiol. 2009;52(4):509-13.
- 15. Agodi A, Barchitta M, Cipresso R, Giaquinta L, Antonietta Romeo M, Denaro C. Pseudomonas aeruginosa carriage, colonization, and infection in ICU patients. Intensive Care Med. 2007;33(7):1155-61.
- 16. Peña C, Guzmán A, Suarez C, Dominguez MA, Tubau F, Pujol M, et al. Effects of carbapenem exposure on the risk for digestive tract carriage of intensive care unit-endemic carbapenem-resistant Pseudomonas aeruginosa strains in critically ill patients. Antimicrob Agents Chemother. 2007;51(6):1967-71.
- 17. Cholley P, Gbaguidi-Haore H, Bertrand X, Thouverez M, Plésiat P, Hocquet D, et al. Molecular epidemiology of multidrug-resistant Pseudomonas aeruginosa in a French university hospital. J Hosp Infect. 2010;76(4):316-9.
- 18. Carvalho APD, Albano RM, Oliveira DN, Cidade DAP, Teixeira LM, Marques EA. Characterization of an epidemic carbapenem-resistant Pseudomonas aeruginosa producing SPM-1 metallo-beta-lactamase in a hospital located in Rio de Janeiro, Brazil. Microb Drug Resist. 2006;12(2):103-8.
- 19. Gruner E, Kropec A, Huebner J, Altwegg M, Daschner F. Ribotyping of Pseudomonas aeruginosa strains isolated from surgical intensive care patients. J Infect Dis. 1993;167(5):1216-20.
- 20. Arruda EA, Marinho IS, Boulos M, Sinto SI, Caiaffa HH, Mendes CM, et al. Nosocomial infections caused by multiresistant Pseudomonas aeruginosa Infect Control Hosp Epidemiol. 1999;20(9):620-3.
- 21. Bonten MJ, Bergmans DC, Speijer H, Stobberingh EE. Characteristics of polyclonal endemicity of Pseudomonas aeruginosa colonization in intensive care units. Implications for infection control. Am J Respir Crit Care Med. 1999;160(4):1212-9.
- 22. Prashanth K, Singh SK, Kanungo R, Sharma S, Shashikala P, Joshi S, et al. Correlations between genotyping and antibiograms of clinical isolates of Pseudomonas aeruginosa from three different south Indian hospitals. Indian J Med Microbiol. 2010;28(2):130-7.
- 23. Kropec A, Huebner J, Riffel M, Bayer U, Benzing A, Geiger K, et al. Exogenous or endogenous reservoirs of nosocomial Pseudomonas aeruginosa and Staphylococcus aureus infections in a surgical intensive care unit. Intensive Care Med. 1993;19(3):161-5.
- 24. Trautmann M, Lepper PM, Haller M. Ecology of Pseudomonas aeruginosa in the intensive care unit and the evolving role of water outlets as a reservoir of the organism. Am J Infect Control. 2005;33(5):S41-9.
- 25. Moolenaar RL, Crutcher JM, San Joaquin VH, Sewell LV, Hutwagner LC, Carson LA, et al. A prolonged outbreak of Pseudomonas aeruginosa in a neonatal intensive care unit: did staff fingernails play a role in disease transmission? Infect Control Hosp Epidemiol. 2000;21(2):80-5.
- 26. Crivaro V, Di Popolo A, Caprio A, Lambiase A, Di Resta M, Borriello T, et al. Pseudomonas aeruginosa in a neonatal intensive care unit: molecular epidemiology and infection control measures. BMC Infect Dis. 2009;9(1):70.
- 27. Burns JL, Saiman L, Whittier S, Krzewinski J, Liu Z, Larone D et al. Comparison of two commercial systems (Vitek and MicroScan-WalkAway) for antimicrobial susceptibility testing of Pseudomonas aeruginosa isolates from cystic fibrosis patients. Diagn Microbiol Infect Dis. 2001;39(4):257-60.
- 28. Zhao WH, Hu ZQ. Beta-lactamases identified in clinical isolates of Pseudomonas aeruginosa Crit Rev Microbiol. 2010;36(3):245-58.
- 29. van Belkum A. DNA fingerprinting of medically important microorganisms by use of PCR. Clin Microbiol Rev. 1994;7(2):174-84.
- 30. Lau YJ, Liu PY, Hu BS, Shyr JM, Shi ZY, Tsai WS et al. DNA fingerprinting of Pseudomonas aeruginosa serotype O11 by enterobacterial repetitive intergenic consensus-polymerase chain reaction and pulsed-field gel electrophoresis. J Hosp Infect. 1995;31(1):61-6.
- 31. Gillings M, Holley M. Amplification of anonymous DNA fragments using pairs of long primers generates reproducible DNA fingerprints that are sensitive to genetic variation. Electrophoresis. 1997;18(9):1512-8.
- 32. Tenover FC, Arbeit RD, Goering RV, Mickelsen PA, Murray BE, Persing DH, et al. Interpreting chromosomal DNA restriction patterns produced by pulsed-field gel electrophoresis: criteria for bacterial strain typing. J Clin Microbiol. 1995;33(9):2233-9.
Publication Dates
-
Publication in this collection
06 Dec 2011 -
Date of issue
2011
History
-
Received
16 Nov 2010 -
Accepted
16 Mar 2011