Abstracts
A new species of Moenkhausia is described from tributaries of the rio Paraguay, Brazil. The new species is diagnosed from congeners by characters related to body coloration, the number of lateral line scales, the degree of poring of the lateral line, and number of scales rows above and below the lateral line. Molecular analyses using partial sequences of the mitochondrial gene Cytochrome Oxidase I from specimens of the new species and specimens belonging to morphologically similar species demonstrated that the new species is easily differentiated by their high genetic distance and by their position in the phylogenetic hypothesis obtained through the Maximum Parsimony methodology. The analyses of three samples of M. oligolepis also revealed that they have high genetic distances and belong to different monophyletic groups suggesting that this species corresponds to a species complex rather than a single species.
Taxonomy; Systematic; Molecular Phylogeny; COI gene; Moenkhausia sanctaefilomenae
Uma nova espécie de Moenkhausia é descrita de tributários do rio Paraguai, Brasil. Essa nova espécie é diagnosticada de seus congêneres por caracteres relacionados ao padrão de colorido do corpo, número de escamas da linha lateral, grau de desenvolvimento dos poros sensoriais na linha lateral e número de séries de escamas acima e abaixo da linha lateral. Um análise molecular usando sequências parciais do gene mitocondrial Citocromo Oxidase I de espécimes representativos da nova espécie e espécimes pertencentes a espécies morfologicamente similares demonstrou que a nova espécie é facilmente diferenciada por sua elevada distância genética e por sua posição na hipótese filogenética obtida pelo método de máxima parcimônia. A análise de três amostras de M. oligolepis também revelou que estas apresentam distâncias genéticas elevadas e pertencem a grupos monofiléticos distintos, sugerindo que esta espécie corresponda a um complexo de espécies e não uma única espécie.
New species of Moenkhausia Eigenmann, 1903 (Characiformes: Characidae) with comments on the Moenkhausia oligolepis species complex
Ricardo C. Benine; Tatiane C. Mariguela; Claudio Oliveira
Laboratório de Biologia e Genética de Peixes, Departamento de Morfologia, IBB-UNESP, Campus de Botucatu, Botucatu, SP 18618-000, Brazil. rbenine@ibb.unesp.br (RCB); tatimariguela@gmail.com (TCM); claudio@ibb.unesp.br (CO)
ABSTRACT
A new species of Moenkhausia is described from tributaries of the rio Paraguay, Brazil. The new species is diagnosed from congeners by characters related to body coloration, the number of lateral line scales, the degree of poring of the lateral line, and number of scales rows above and below the lateral line. Molecular analyses using partial sequences of the mitochondrial gene Cytochrome Oxidase I from specimens of the new species and specimens belonging to morphologically similar species demonstrated that the new species is easily differentiated by their high genetic distance and by their position in the phylogenetic hypothesis obtained through the Maximum Parsimony methodology. The analyses of three samples of M. oligolepis also revealed that they have high genetic distances and belong to different monophyletic groups suggesting that this species corresponds to a species complex rather than a single species.
Key words: Taxonomy, Systematic, Molecular Phylogeny, COI gene, Moenkhausia sanctaefilomenae.
RESUMO
Uma nova espécie de Moenkhausia é descrita de tributários do rio Paraguai, Brasil. Essa nova espécie é diagnosticada de seus congêneres por caracteres relacionados ao padrão de colorido do corpo, número de escamas da linha lateral, grau de desenvolvimento dos poros sensoriais na linha lateral e número de séries de escamas acima e abaixo da linha lateral. Um análise molecular usando sequências parciais do gene mitocondrial Citocromo Oxidase I de espécimes representativos da nova espécie e espécimes pertencentes a espécies morfologicamente similares demonstrou que a nova espécie é facilmente diferenciada por sua elevada distância genética e por sua posição na hipótese filogenética obtida pelo método de máxima parcimônia. A análise de três amostras de M. oligolepis também revelou que estas apresentam distâncias genéticas elevadas e pertencem a grupos monofiléticos distintos, sugerindo que esta espécie corresponda a um complexo de espécies e não uma única espécie.
Introduction
The distributed characid genus Moenkhausia was proposed by Eigenmann (1903) based mainly on the combination of the presence of premaxillary teeth in two rows, an inner premaxillary tooth row with five or more teeth, a completely pored lateral line without a downward curve anteriorly, and a sheath of scales covering the base of caudal-fin lobes, characters still used to substantiate the generic allocation of new species to Moenkhausia (see Benine et al., 2007; Lima et al., 2007; Lucinda et al., 2007), considering the nonexistence of a phylogenetic definition. Currently allocated as incertae sedis in Characidae, Moenkhausia comprises, at present, 65 valid species widely distributed in the Neotropical Cis-Andean river basins, except for those in Patagonia, with its greatest diversity occurring in the basins of the Amazon and Guianas (Lima et al., 2003).
Albeit the genus Moenkhausia cannot be defined as monophyletic at this time, a group of species within Moenkhausia shares an identical or, at least, very similar color pattern consisting of an overall reticulate pattern formed by dark borders on the scales, a vertically-elongate humeral blotch, a conspicuous dark blotch on the caudal peduncle preceded by a lighter area, and frequently a deep red eye in life. This group consists of M. oligolepis Günther, M. sanctaefilomenae Steindachner, M. cotinho Eigenmann, and M. pyrophthalma Costa, as suggested by Costa (1994). Lima & Toledo-Piza (2001) discussed a putatively close relationship of M. dyktiota Lima & Toledo-Piza with these cited species also on the basis of its overall similar colour pattern, albeit without the characteristic round, dark, blotch on the caudal peduncle.
Despite the similar color pattern, these species are easily differentiated from each other on the basis of number of lateral line scales and number of series of scales above and below lateral line. Nonetheless, misidentifications between Moenkhausia oligolepis and M. sanctaefilomenae are common. According to Lima et al. (2003), M. oligolepis occurs in the Guianas and in the Amazon River basin, whereas M. sanctaefilomenae is distributed in the Parnaíba, São Francisco, upper Paraná, Paraguay and Uruguay River systems. Detailed examination revealed that not does only Moenkhausia oligolepis occur in the rio Paraguay basin but also that new similar species occur in sympatry revealing a species complex. In the present paper, we describe a new species of the M. oligolepis species complex and provide an analysis of its phylogenetic position based on molecular data.
Material and Methods
Material examined in this study is deposited in the California Academy of Sciences (CAS), San Francisco; Laboratório de Biologia e Genética de Peixes do Departamento de Morfologia da Universidade Estadual Paulista (LBP), Botucatu, SP; Museu de Ciências e Tecnologia, Pontifícia Universidade Católica do Rio Grande do Sul (MCP), Porto Alegre, RS; and Museu de Zoologia da Universidade de São Paulo (MZUSP), São Paulo, SP. Vouchers of the material used in the molecular analysis are deposited at LBP.
Morphometric and meristic data were taken following Fink & Weitzman (1974), except for head depth (HD), which was measured in the vertical through the middle of the orbit. Counts are followed by their frequency in parentheses. Asterisks indicate values for the holotype. Vertebrae of the Weberian apparatus were counted as four elements and the fused PU1+U1 as a single element. Vertebra and supraneural counts were taken from two cleared and stained (c&s) specimens prepared following the method of Taylor & Van Dyke (1985).
Total DNA was extracted from ethanol-preserved muscle tissues with the Wizard Genomic DNA Purification Kit (Promega). Partial sequences of COI gene were amplified with the primers Fish F1 and Fish R1 (Ward et al., 2005). Primer final concentrations were 5 mM. Amplifications were performed by the polymerase chain reaction (PCR) in a total volume of 25 ml for 35 cycles (30 s at 95 ºC, 60 s at 50-60 ºC, and 120 s at 72 ºC). The PCR products were identified in a 1% agarose gel. The amplified segments were extracted from the gel with the kit GFXTM PCR DNA and Gel Purification (Amersham Pharmacy Biotech Inc). The sequencing reactions were done with the kit DYEnamic ET Terminator Cycle Sequencing (Amersham Pharmacy Biotech Inc.) and analyzed in an automated sequencer ABI PrismTM 377 DNA Sequencer (Perking-Elmer). All sequences were read at least twice (forward and reverse). Individual sequences of each species were initially analyzed using BioEdit 5.0.9 (Hall, 1999) and a consensus sequence was obtained for each DNA segment for each species. All sequences were then aligned with DAMBE (Xia & Xie, 2001). Genetic distances (Kimura, 1980) were examined using MEGA 4.0 (Kumar et al., 2004).
Phylogenetic analyses based on the maximum-parsimony method (Swofford & Berlocher, 1987) were performed using PAUP* beta version 4.0b10 (Swofford, 2002) with heuristic searches using random addition of sequences and the tree bisection and reconnection (TBR) algorithm. In all analyses the character-state optimization method employed was the accelerated transformation (ACCTRAN). Parsimony trees were generated using Ti/Tv ratio of 1:1 and considering terminal gaps as missing data. Bootstrap resampling (Felsenstein, 1985) was applied to assess support for individual nodes using 1000 replicates with random additions and TBR branch swapping.
Moenkhausia forestii, new species
Holotype. MZUSP 97827, 34.0 mm SL, undetermined sex, Brazil, Mato Grosso State, Cáceres, rio Sepotuba, rio Paraguay basin,15º46'07"S 57º38'54"W, 3-4 Mar 2002, H. A. Britski, O. Fröehlich, A. Catella & F. Marques.
Paratypes. MZUSP 90270, 14, 24.0-36.4 mm SL, (2 c&s, 25.6-33.1 mm SL), same data as the holotype. MZUSP 19111, 10, 30.7-35.0 mm SL, Brazil, Mato Grosso State, Descalvados, rio Paraguay, 16º46'S 57º44'W, 9 Aug 1980, R. M. C. Castro & H. Ortega. LBP 3793, 10, 28.3-33.4 mm SL, Brazil, Mato Grosso do Sul State, Aquidauana, rio Negro, rio Paraguay basin, 19º34'02.3"S 56º14'09.1"W, 2 Aug 2006, C. Oliveira & L. H. G. Pereira.
Non-types. LBP 5074, 10, 25.7-32.1 mm SL, Brazil, Mato Grosso State, Cáceres, Baía do Caiçara, 16º03'11.3"S 57º48'32.0"W. LBP 3739, 10, 22.4 - 32.5 mm SL, Brazil, Mato Grosso do Sul State, Aquidauana, rio Negro, rio Paraguay basin, 19º34'54.6"S 56º15'16.5"W. LBP 4655, 65, 21.9-32.9 mm SL, Brazil, Mato Grosso do Sul State, Batayporã, rio Baía, rio Paraná basin, 22º43'46.2"S 53º19'04.2"W. LBP 2630, 18, 23.0-37.9 mm SL, Brazil, Paraná State, Porto Rico, rio Paraná, 22º43'03.2"S 53º17'27.6"W. LBP 5225, 10, 25.0-32.8 mm SL, Brazil, Paraná State, Porto Rico, rio Paraná, 22º47'29"S 53º20'58"W.
Diagnosis. Moenkhausia forestii is readly distinguished from all congeners, except M. oligolepis, M. sanctaefilomenae, M. pyrophthalma, and M. diktyota, by the presence of a reticulated body pigmentation pattern, and a conspicuous dark blotch on the caudal peduncle extending to the base of caudal-fin rays preceded by a lighter area (vs. absence of a reticulated body pigmentation pattern, and a conspicuous dark blotch on the caudal peduncle extending to the base of caudal-fin rays preceded by a lighter area). Moenkhausia forestii is readily distinguished from M. oligolepis in the degree of poring of the lateral line (incomplete, vs. complete, respectively), and the total number of lateral line scales (23-26, vs. 28-31 scales, respectively). Moenkhausia forestii can be distinguished from M. sanctaefilomenae in the number of transverse series of scales above the lateral line (5, vs. 4 scales, respectively), by the number of transverse series of scales below the lateral line (4, vs. 3 scales, respectively). Moenkhausia forestii is distinguished from M. pyrophthalma by its higher body depth (38.3-45.2% in SL vs. 31.2-33.0% in SL, respectively), by its shorter lateral line (23-26 vs. 27-31 scales, respectively), and by the form of the humeral spot (an inconspicuous dark inverse triangular humeral blotch located on second and third lateral line scales vs. a conspicuous black inverse drop-shaped humeral blotch located on second to fourth lateral line scales, respectively). Moenkhausia forestii is distinguished from M. diktyota by the color pattern of the caudal peduncle (caudal peduncle with very widely scattered dark pigment resulting in conspicuous lighter area preceding a conspicuous caudal-fin blotch which extends between the dorsal to ventral margins of the proximal one-third of caudal-fin base vs. caudal peduncle with elongate black stripe extending from the vertical through the last anal-fin ray to the tips of middle caudal fin rays, respectively).
Description. Morphometric data summarized in Table 1. Overall size small (maximum of 36.4 mm SL). Greatest body depth at origin of dorsal fin. Dorsal profile of head straight or slightly convex. Dorsal profile of body sligthly convex from posterior tip of supraocciptal to end of dorsal-fin base; slightly convex from rear of dorsal-fin base to end of adipose-fin base. Caudal peduncle profile slightly concave both dorsally and ventrally. Ventral profile of body convex from tip of lower jaw to caudal peduncle origin. Prepelvic region transversely flattened, flattening more pronounced proximate to pelvic-fin insertion. Postpelvic median keel extending from pelvic-fin insertion to anal-fin origin.
Mouth terminal, with lower jaw as long as, to somewhat longer than, upper jaw. Premaxillary teeth in two rows; outer row teeth 3-5* (mode = 5, n = 35), with 3-5 cusps, central cusp longer than other cusps; inner row teeth 5, with 4-5 cusps and rarely with lateral-most tooth with 3 cusps. Maxillary teeth 1-2* (mode = 2, n = 35), each with 5 cusps. Dentary with 4 larger teeth anteriorly, each with 4-5 cusps with central cusp longest (n =35). Larger anterior teeth followed by 5-7 small teeth with 1-3 cusps in two c&s paratypes (Fig. 2).
Dorsal-fin rays ii,9. Dorsal-fin origin at middle body length, slightly posterior to vertical through pelvic-fin origin. Posterior margin of dorsal fin slightly concave. Anal-fin rays iv,17-23, iv,22* (mode = iv,22, n = 35). Distal margin of anal fin straight to somewhat concave with 4th unbranched and anterior 1st-3rd branched fin ray longer. Remaining anal-fin rays decreasing gradually in length rearward. Pectoral-fin rays i,11-12* (mode = i,12, n = 35). Pectoral fin pointed, 1st-2nd branched rays longer, lateral and medial margins straight, posterior margin oblique and straight. Tip of adpressed pectoral fin extends posterior of mid-length of adpressed pelvic fin. Pelvic-fin rays i,7*. Pelvic fin pointed, lateral and medial margins straight, posterior margin oblique and straight.Tip of pelvic fin reaches anal-fin origin. Adipose fin origin at vertical through insertion point of antepenultimate branched anal fin. Principal caudal-fin rays i,9,8,i. Caudal-fin lobes equal.
Scales cycloids. Lateral line with 7-11 (10*; mode = 9, n = 34) pored scales and 23-26 (25*; mode = 25, n = 34) total scales. Lateral line slightly ventrally curved anteriorly; with 5 (n = 35) series of scales above and 4 (n = 35) series of scales below. Scales around caudal peduncle 7-9* (mode = 9, n = 35). Single row of scales overlaying basal portion of anterior rays of anal fin. Sheet of scales covering proximal one-third of upper caudal-fin lobe and proximal one-half of lower caudal-fin lobe.
First gill arch with 6-7* (mode = 7, n = 35) gill-rakers on upper limb and 10-11* (mode = 11, n = 28) gill-rakers on lower limb. Precaudal vertebrae 13, caudal vertebrae 17 (n = 2). Supraneurals 4 (n = 2).
Color in alcohol. Overall ground coloration dark silver or yellowish tan (depending on degree of retention of guanine). Dark chromatophores concentrated on distal margin of scales resulting in conspicuous reticulated pattern. Mid-dorsal region darker than flanks. Humeral region with dark inverse triangle-shaped blotch located on second to third lateral line scales, extending 3-4 scales vertically, including lateral line. Few dark chromatophores scattered on infraorbitals and opercle. Dark thin stripe extending along horizontal septum on posterior half of body. Caudal fin with conspicuous dark blotch extending from its anterior portion onto proximal one-third of caudal-fin rays. Caudal peduncle with very widely scattered dark pigment resulting in conspicuous lighter area preceding caudal-fin blotch. Dorsal fin with scattered dark pigmentation, more concentrated on anterior half. Anal fin with scattered dark pigmentation. Paired fins hyaline with scattered dark pigmentation, more so on unbranched rays.
Sexual dimorphism. One paratype (30.9 mm SL, MZUSP 90270) has small hooks on the segments of each anal-fin ray (up to four hooks per segment in each side of the anal fin). Additional non-type material from the upper rio Paraná system (LBP 5225) collected during the reproductive season included males with the same pattern of hooks. No further dimorphic characters were observed.
Distribution. Moenkhausia forestii is known from tributaries from rio Paraguay drainage (Mato Grosso and Mato Grosso do Sul States), from rio Baía (upper rio Paraná system at Batayporã, Mato Grosso do Sul State) and from rio Paraná (at Porto Rico, Paraná State) (Fig. 3).
Etymology. The specific epithet, forestii, is in honour of Fausto Foresti for his contributions to our knowledge of fish genetics.
Molecular Analyses. DNA sequences were obtained from: Moenkhausia forestii LBP 3739, LBP 4655 and LBP 5074; Moenkhausia oligolepis LBP 4098; LBP 1498 and LBP 5073; Moenkhausia sanctaefilomenae LBP 4695.
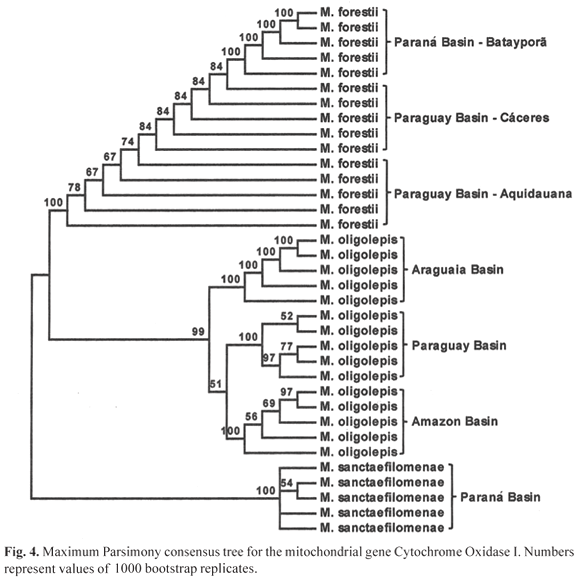
The sequences obtained in this study have been deposited in GenBank (Table 2). Genetic distances (Kimura, 1980) range from zero among specimens of Moenkhausia forestii from Paraná basin to 0.200 ± 0.021 between M. oligolepis from the rio Araguaia Basin and M. sanctaefilomenae from the upper rio Paraná system (Table 2). The combined sequence data of the 35 specimens resulted in a matrix with 611 base pairs (bp), from which 419 were conserved sites and 181 were phylogenetically informative. The MP consensus tree obtained from the analysis of 1000 bootstrap replicates is presented in Fig. 4.
The MP analyses showed that there are six monophyletic groups supported by values of equal or higher than 99% (Fig. 4). Three monophyletic groups corresponding to the species Moenkhausia sanctaefilomenae, M. oligolepis and M. forestii and three monophyletic groups correspond to the different samples of M. oligolepis. The relationship between M. oligolepis from the Amazon and Paraguay river basins was weakly supported but they are apparently different from the sample from the rio Araguaia basin.
Discussion
According to Eigenmann (1907), the degree of development of the lateral-line (number and position of perforated scales) distinguishes Moenkhausia (complete lateral line) from Hemigrammus (incomplete lateral line) and Psellogrammus (interrupted lateral line). This purported difference was recently discussed by Costa (1994), Lima & Toledo-Piza (2001), who highlighted the weakness of the phylogenetic information in this feature. These authors described their new species in Moenkhausia, even though those forms demonstrated incompletely pored lateral-lines and, thus failed to conform to the traditional definition of the genus. They based their generic assignments on other evidence, specifically overall similarity with other species of Moenkhausia that they felt justified this generic assignment. Such procedure was equally adopted herein.
Eigenmann (1917: 84) included specimens with incomplete, interrupted, and complete lateral line in a single species, Moenkhausia sanctaefilomenae. In fact, this author raised the potential of the existence of more than one species in those samples, noting that the forms with incomplete lateral line would correspond to his M. australe, and those with both interrupted and complete lateral line would correspond to M. sanctaefilomenae. The basis for Eigenmann's actions was unclear, but he presumably included samples with different degrees of development of the lateral line in a single species in the absence of broad samples at his disposal and consequently placed M. australe in the synonymy of M. sanctaefilomenae. Moenkhausia australe was, however, described by Eigenmann (1908) from two specimens collected at two different localities in Paraguay, Arroyo Trementina and Arroyo Chagalalina. Those specimens were examined as part of this study and, contrary to Eigenmann (1917), these specimens do not have an incompletely pored lateral line and present a higher upper jaw length in comparison with Moenkhausia forestii (48.9-52.6% vs. 39.9-43.7% in HL, respectively), thus, M. australe is not equivalent to the new species described herein. Additional material from rio Uruguay (MCP 25198 and MCP 27464) with a complete lateral line has a comparable upper jaw length (47.2-52.4% in HL) with the syntypes of M. australe.
The observed values of genetic distances (Kimura, 1980) among species were 0.182 ± 0.019 between M. sanctaefilomenae and M. forestii, 0.183 ± 0.017 between M. oligolepis and M. forestii , and 0.191 ± 0.017 between M. sanctaefilomenae and M. oligolepis. These values are higher than those obtained in the similar analyses of within-genus COI sequences (0.0993) of marine fishes from Australia (Ward et al., 2005) and of freshwater fishes from Canada (Hubert et al., 2008) and from Mexico and Guatemala (Valdez-Moreno et al., 2009). The observed values of genetic distances within-species were 0.003 ± 0.001 for M. sanctaefilomenae, 0.004 ± 0.001 for M. forestii, and 0.092 ± 0.009 for M. oligolepis. This high value observed for M. oligolepis leads us to hypothesize that M. oligolepis still represents a species complex rather than a single species which will be explored in a further study.
Searches through the collections at MZUSP revealed that Moenkhausia forestii has been only recently collected in the upper portions of the rio Paraná system, most probably as a consequence of movement of the species through the "Piracema Channel" (a fish ladder constructed around Itaipu Hydroelectric). This fish ladder linked the middle and upper portions of rio Paraná, that were originally separated by the natural barrier formed by the Sete Quedas falls. Several other examples of similar establishment of species in the upper Paraná as a consequence of movement via this fish ladder are cited by Graça & Pavanelli (2007). Due to this secondary establishment, the specimens from rio Paraná drainage were included as non-type material. Non-types also includes dissected, alcohol-fixed, and poorly preserved specimens.
Comparative material. Moenkhausia australe: CAS 70818, syntype, 38.9 mm SL, arroyo Trementina, rio Paraguay, Paraguay; CAS 70819, syntype, 29.7 mm SL, arroyo Chagalalina, rio Paraguay, Paraguay; MCP 25198, 5, 44.0-54.8 mm SL, rio Uruguay, Rio Grande do Sul State; MCP 27464, 5, 34.1-49.0 mm SL, rio Uruguay, Rio Grande do Sul State, Brazil. Moenkhausia diktyota: MZUSP 64107, 2,06 paratypes, 35.4-44.8 mm SL, rio Tiquié, Amazonas State, Brazil. Moenkhausia oligolepis: LBP 5365, 1, 37.3 mm SL, rio Jari, Laranjal do Jari, Amapá State, Brazil; LBP 4098, 9, 34.8-47.7 mm SL, rio Japiim, rio Amazonas basin, Mâncio Lima, Acre State, Brazil; LBP 1498, 10, 46.1-65.1 mm SL, córrego Fundo, rio Araguaia basin, Barra do Garças, Mato Grosso State, Brazil; LBP 5073, 29, 39.0-52.5 mm SL, rio Paraguay Basin, Cáceres, Mato Grosso State, Brazil. Moenkhausia pyrophthalma: MZUSP 45290, 7, paratypes, 19.8-29.8 mm SL, rio das Mortes, between Água Boa and Cocalinho, Mato Grosso State, Brazil; LBP 4922, 5, 27.6-34.0 mm SL, rio das Mortes, Barra do Garças, Mato Grosso State, Brazil. Moenkhausia sanctaefilomenae: LBP 4695, 8, 25.1-61.3 mm SL, rio Paraná Basin, Marapoama, São Paulo State, Brazil; LBP 5541, 44, 31.6 - 47.1 mm SL, MCP 42016, 5, 32.6-45.6 mm SL, rio Parnaíba, Santa Filomena, Piauí State, Brazil; MZUSP 94090, 5, 25.9-43.2 mm, SL, rio Parnaíba, Santa Filomena, Piauí State, Brazil.
Acknowledgements
The authors are indebted to Osvaldo T. Oyakawa (MZUSP) for loan of material and curatorial assistance, to Dave Catania (CAS) and Zilda M. Lucena (MCP) for loan of material. Richard P. Vari (USNM) provided valuable suggestions and comments on various drafts of the manuscript. Waldo P. Troy provided specimens for molecular analysis. Guilherme A. M. Lopes assisted with Fig. 1. RCB and TCM are financially supported by FAPESP (proc. 06/00545-3; 06/04551-8, respectively) and CO by FAPESP and CNPq (proc. 306054/2006-0). This study is part of the FAPESP Thematic Project "Phylogenetic relationships in the Characidae (Ostariophysi: Characiformes) (proc. 04/09219-6).
Literature Cited
Accepted April 2009
Published June 17, 2009
- Benine, R. C., R. M. C. Castro & A. C. A. Santos. 2007. A new Moenkhausia Eigenmann, 1903 (Characiformes: Characidae) from Chapada Diamantina, rio Paraguaçu basin, Bahia, Northeastern Brazil. Neotropical Ichthyology, 5(3): 259-262.
- Costa, W. J. E. M. 1994. Description of two new species of the genus Moenkhausia (Characiformes: Characidae) from the central Brazil. Zoologischer Anzeiger, 232: 21-29.
- Eigenmann, C. H. 1903. New genera of South America fresh-water fishes, and new names for some old genera. Smithsonian Miscellaneous Collection, 45: 144-148.
- Eigenmann, C. H. 1907. On further collections of fishes from Paraguay. Annals of the Carnegie Museum, 4: 108-157.
- Eigenmann, C. H. 1908. Preliminary descriptions of new genera and species of tetragonopterid characins. (Zoölogical Results of the Thayer Brazilian Expedition). Bulletin of the Museum of Comparative Zoology, 52: 91-106.
- Eigenmann, C. H. 1917. The American Characidae. Part I. Memoirs of the Museum of Comparative Zoology, 43: 1-102.
- Felsenstein, J. 1985. Confidence limits on phylogenies: an approach using the bootstrap. Evolution, 39: 783-791.
- Fink, W. L. & S. H. Weitzman. 1974. The so-called Cheirodontin fishes of Central America with descriptions of two new species (Pisces: Characidae). Smithsonian Contributions to Zoology, 172: 1-46.
- Graça, W. J. & C. S. Pavanelli. 2007. Peixes da planície de inundação do alto rio Paraná e áreas adjacentes. Maringá, EDUEM, 241p.
- Hall, T. A. 1999. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symposium Series, 41: 95-98.
- Hubert, N., R. Hanner, E. Holm, N. E. Mandrak, E. Taylor, M. Burridge, D. Watkinson, P. Dumont, A. Curry, P. Bentzen, J. Zhang, J. April & L. Bernatchez. 2008. Identifying canadian freshwater fishes through DNA barcodes. PLoS ONE, 3: e2490.
- Kimura, M. 1980. A simple method of estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. Journal of Molecular Evolution, 16: 111-120.
- Kumar, S., K. Tamura & M. Nei. 2004. MEGA3: Integrated software for Molecular Evolutionary Genetics Analysis and sequence alignment. Briefings in Bioinformatics, 5: 150-163.
- Lima, F. C. T., H. A. Britski & F. A. Machado. 2007. A new Moenkhausia (Characiformes: Characidae) from central Brazil, with comments on the area relationship between the upper rio Tapajós and upper rio Paraguai systems. Aqua International Journal of Ichthyology, 13(2): 45-54.
- Lima, F. C. T., L. R. Malabarba, P. A. Buckup, J. F. P, Silva, R. P. Vari, A. Harold, R. Benine, O. T. Oyakawa, C. S. Pavanelli, N. A. Menezes, C. A. S. Lucena, R. E. Reis, F. Langeani, L. Casatti, V. A. Bertaco, C. R. Moreira & P. H. F. Lucinda. 2003. Genera Incertae Sedis in Characidae. Pp. 106-169. In: Reis, R. E., S. O. Kullander & C. J. Ferraris Jr. (Eds.). Check List of the Freshwater fishes of South and Central America. Porto Alegre, Edipucrs, 729p.
- Lima, F. C. T. & M. Toledo-Piza. 2001. New species of Moenkhausia (Characiformes: Characidae) from the rio Negro of Brazil. Copeia, 2004(4): 1058-1063.
- Lucinda, P. H. F., L. R. Malabarba & R. C. Benine. 2007. On a new species of the genus Moenkhausia Eigenmann (Ostariophysi: Characidae). Zootaxa, 1525: 61-68.
- Swofford, D. L. 2002. PAUP* - Phylogenetic analysis using parsimony (*and other methods). Version 4b10. Sinauer, Sunderland.
- Swofford, D. L. & S. H. Berlocher. 1987. Inferring evolutionary trees from gene frequency data under the principle of maximum parsimony. Systematic Zoology, 36(3): 293-325.
- Taylor, W. R. & G. C. Van Dyke. 1985. Revised procedures for staining and clearing small fishes and other vertebrates for bone and cartilage. Cybium, 9(2): 107-119.
- Valdez-Moreno, M., N. V. Ivanova, M. Elias-Gutierrez, S. Contreras-Balderas & P. D. N. Hebert. 2009. Probing diversity in freshwater fishes from Mexico and Guatemala with DNA barcodes. Journal of Fish Biology, 74(2): 377-402.
- Ward, R. D., T. S. Zemlak, B. H. Innes, P. R. Last & P. D. N. Hebert. 2005. DNA barcoding Australia's fish species. Philosophical Transactions of Royal Society, 359: 1847-1857.
- Xia, X. & Z. Xie. 2001. DAMBE: Data analysis in molecular biology and evolution. Journal of Heredity, 92: 371-373.
Publication Dates
-
Publication in this collection
15 July 2009 -
Date of issue
June 2009
History
-
Accepted
17 June 2009 -
Received
Apr 2009