ABSTRACT
Characiformes is the most cytogenetically studied group of freshwater Actinopterygii, but karyotypical data of several taxa remain unknown. This is the case of Nematocharax , regarded as a monotypic genus and characterized by marked sexual dimorphism. Therefore, we provide the first cytogenetic report of allopatric populations of Nematocharax venustus based on distinct methods of chromosomal banding and fluorescence in situ hybridization (FISH) with repetitive DNA probes (18S and 5S rDNA). The karyotype macrostructure was conserved in all specimens and populations, independently on sex, since they shared a diploid number (2n) of 50 chromosomes divided into 8m+26sm+14st+2a. The heterochromatin was mainly distributed at pericentromeric regions and base-specific fluorochrome staining revealed a single pair bearing GC-rich sites, coincident with nucleolar organizer regions (NORs). On the other hand, interpopulation variation in both number and position of repetitive sequences was observed, particularly in relation to 5S rDNA. Apparently, the short life cycles and restricted dispersal of small characins, such as N. venustus , might have favored the divergence of repetitive DNA among populations, indicating that this species might encompass populations with distinct evolutionary histories, which has important implications for conservation measures.
Keywords:
18S rDNA; 5S rDNA; Coastal drainages; FISH; Population cytogenetics
RESUMO
Characiformes é o grupo de Actinopterygii de água doce mais estudado citogeneticamente, porém dados cariotípicos de vários taxa permanecem desconhecidos. Este é o caso de Nematocharax , considerado um gênero monotípico e caracterizado pelo acentuado dimorfismo sexual. Em vista disso, nós fornecemos a primeira descrição citogenética de populações alopátricas de Nematocharax venustus , baseada em métodos distintos de bandamento cromossômico e hibridação fluorescente in situ (FISH) com sondas de DNA repetitivo (DNAr 18S e 5S). A macroestrutura cariotípica mostrou-se conservada em todos os espécimes e populações, independentemente do sexo, uma vez que compartilharam um número diploide (2n) de 50 cromossomos dividido em 8m+26sm+14st+2a. A heterocromatina distribuiu-se principalmente nas regiões pericentroméricas e a coloração com fluorocromos base-específicos revelou um único par portador de sítios GC-ricos, coincidentes com as regiões organizadoras de nucléolo (RONs). Por outro lado, foi observada uma variação interpopulacional no número e na posição das sequências repetitivas, especialmente em relação ao DNAr 5S. Aparentemente, ciclos de vida curtos e dispersão restrita dos pequenos caracídeos, tal como N. venustus , podem ter favorecido a divergência do DNA repetitivo entre as populações, indicando que essa espécie pode englobar populações com distintas histórias evolutivas, o que tem implicações importantes para medidas de conservação.
Introduction
The Neotropical freshwater ichthyofauna is remarkable by their richness, complexity and high endemism (Vari & Malabarba, 1998Vari, R. P. &L. R. Malabarba. 1998. Neotropical Ichthyology: an overview. Pp. 1-11. In: Malabarba, L. R., R. E. Reis, R. P. Vari, Z. M. S. Lucena & C. A. S. Lucena (Eds.). Phylogeny and classification of Neotropical fishes. Porto Alegre, Edipucrs, 603p.). A great majority of fish species in this region belongs to the family Characidae, which includes at least 1,107 valid species (Eschmeyer & Fong, 2015Eschmeyer, W. N. & J. D. Fong. 2015. Species by family/subfamily. Species by family/subfamily in the catalog of fishes. Available from: Available from: http://researcharchive.calacademy.org/research/ichthyology/catalog/speciesbyfamily.asp
. (05 February 2016).
http://researcharchive.calacademy.org/re...
). However, the phylogenetic relationships and systematics of genera in this family are controversial, being several taxa placed as incertae sedis (Weitzman & Fink, 1983Weitzman, S. H. & W. L. Fink. 1983. Relationships of the neon tetras, a group of South American freshwater fishes (Teleostei, Characidae), with comments on the phylogeny of New World Characiforms. Bulletin of the Museum of Comparative Zoology, 150: 339-395.). In addition, the identification of species complexes or cryptic species hinders a precise characterization of the diversity in this group (Kavalco et al., 2009Kavalco, K. F., K. O. Brandão, R. Pazza & L. F. A. Toledo. 2009. Astyanax hastatus Myers, 1928 (Teleostei, Characidae): a new species complex within the genus Astyanax ?. Genetics and Molecular Biology, 32: 477-483.; Castro et al., 2014aCastro, J. P., M. O. Moura, O. Moreira Filho, O. A. Shibatta, M. H. Santos, V. Nogaroto, M. R. Vicari, M. C. Almeida & R. F. Artoni. 2014a. Diversity of the Astyanax scabripinnis species complex (Teleostei: Characidae) in the Atlantic Forest, Brazil: species limits and evolutionary inferences. Reviews in Fish Biology and Fisheries, 25: 231-244.).
Accordingly, cytogenetic analyses have been carried out to address this issue, since chromosomal markers have proved to be useful for both systematic and evolutionary inferences in Neotropical fish (Blanco et al., 2010Blanco, D. R., R. L. Lui, L. A. C. Bertollo, V. P. Margarido & O. Moreira Filho. 2010. Karyotypic diversity between allopatric populations of the group Hoplias malabaricus (Characiformes: Erythrinidae): evolutionary and biogeographic considerations. Neotropical Ichthyology, 8: 361-368.; Almeida et al., 2013Almeida, J. S., P. R. A. M. Affonso, D. Diniz, P. L. S. Carneiro & A. L. Dias. 2013. Chromosomal variation in the tropical armored catfish Callichthys callichthys (Siluriformes, Callichthyidae): implications for conservation and taxonomy in a species complex from a Brazilian hotspot. Zebrafish, 10: 451-458.; Piscor et al., 2015Piscor, D., A. L. Alves & P. P. P. Maltempi. 2015. Chromosomal microstructure diversity in three Astyanax (Characiformes, Characidae) species: comparative analysis of the chromosomal locations of the 18S and 5S rDNAs. Zebrafish, 12: 81-90.), including taxonomically problematic groups (Bellafronte et al., 2010Bellafronte, E., M. O. Schemberger, O. Moreira Filho, M. C. Almeida, R. F. Artoni, V. P. Margarido & M. R. Vicari. 2010. Chromosomal markers in Parodontidae: an analysis of new and reviewed data with phylogenetic inferences. Reviews in Fish Biology and Fisheries, 21: 559-570.; Mendes et al., 2011Mendes, M. M., R. Rosa, L. G. Caetano & A. L. Dias. 2011. Karyotype diversity of four species of the incertae sedis group (Characidae) from different hydrographic basins: analysis of AgNORs, CMA3 and 18S rDNA. Genetics and Molecular Research, 10: 3596-3608.). Moreover, the availability of refined cytogenetic methods have allowed identifying population polymorphism and unique evolutionary units, even within morphologically similar groups (Bitencourt et al., 2011Bitencourt, J. A., P. R. A. M. Affonso, L. Giuliano-Caetano & A. L. Dias. 2011. Identification of distinct evolutionary units in allopatric populations of Hypostomus cf. wuchereri Günther, 1864 (Siluriformes: Loricariidae): karyotypic evidence. Neotropical Ichthyology, 9: 317-324.; Utsunomia et al., 2014Utsunomia, R., J. C. P. Alves, G. J. C. Silva, F. F. Mendonça, P. C. Scacchetti, C. Oliveira & F. Foresti. 2014. Molecular and cytogenetic analyses of cryptic species within the Synbranchus marmoratus Bloch, 1795 (Synbranchiformes: Synbranchidae) grouping: species delimitations, karyotypic evolution and intraspecific diversification. Neotropical Ichthyology, 12: 903-911.).
Nematocharax Weitzman, Menezes & Britski represents one of the several genera in Characidae whose taxonomic status is still under debate. This genus has been considered monotypic since the validation of Nematocharax venustusWeitzman, Menezes & Britski (1986Weitzman, S. H., N. A. Menezes & H. A. Britski. 1986. Nematocharax venustus , a new genus and species of fish from the rio Jequitinhonha, Minas Gerais, Brazil (Teleostei: Characidae). Proceedings of the Biological Society of Washington, 99: 335-346.), originally described for the Jequitinhonha River basin, eastern Brazil. Recently, Bragança et al. (2013Bragança, P. H. N., M. A. Barbosa & J. L. Mattos. 2013. A new Nematocharax species from the middle Contas River basin, Northeastern Brazil (Characiformes: Characidae). Vertebrate Zoology, 63: 3-8.) reported a second species, named N. costai , for the Contas River basin, northeastern Brazil, which was not widely accepted among ichthyologists. Hence, Menezes et al . (2015Menezes, N. A., A. M. Zanata & P. Camelier. 2015. Nematocharax costai Bragança, Barbosa & Mattos a junior synonym of Nematocharax venustus Weitzman, Menezes & Britski (Teleostei: Characiformes: Characidae). Zootaxa, 3920: 453-462.) carried out a detailed analysis of meristic, morphometric, osteological and coloration patterns of specimens from several hydrographic basins throughout the range of Nematocharax . According to these authors, N. costai is actually a junior synonym of N. venustus once overlapped morphological features and polymorphism of secondary sex traits in males were observed for both putative species, thereby supporting Nematocharax as a monotypic genus.
Based on the complex systematics and morphological variation of N. venustus , we performed comparative chromosomal analyses in allopatric populations of this species, including those previously recognized as N. costai in order to elucidate the taxonomic status of this fish group and infer biogeographic aspects in ichthyofauna from coastal basins of eastern Brazil. Therefore, distinct cytogenetic methodologies, ranging from basic banding techniques up to fluorescence in situ hybridization (FISH) experiments with repetitive DNA probes, were used to identify the karyotypic structure and possible population differences through the range of this species.
Material and Methods
A total of 71 males and 70 females of Nematocharax venustus were collected in seven localities along the Contas, Almada and Jequitinhonha River basins in the states of Bahia and Minas Gerais (Brazil) (Fig. 1). The data about collection sites and samples are shown in Table 1.
Map of Brazil (a) and collection sites (b) of Nematocharax venustus along the Almada (Almada), Contas (Upper Contas, Gongogi 1, 2, and 3), and Jequitinhonha River basins (Jequitinhonha 1 and 2) in the states of Bahia and Minas Gerais.
Collection sites and sampling of Nematocharax venustus from the Almada, Contas, and Jequitinhonha River basins. ♂ = males and ♀ = females.
The collection license was provided by Instituto Chico Mendes de Conservação da Biodiversidade (ICMBio; license number SISBIO 39728-1). Voucher specimens were stored in the fish collection from the Museu de Zoologia da Universidade Federal da Bahia (MZUFBA), Brazil (UFBA 7953, 7954, 8016, 8017, 8018, 8019, and 8020).
The specimens were transported to aerated tanks, and mitotic chromosomes were obtained from kidney cells according to Netto et al. (2007Netto, M. R. C. B., E. Pauls & P. R. A. M. Affonso. 2007. A standard protocol for obtaining fish chromosomes under post-mortem conditions. Micron, 38: 214-217.) and Blanco et al. (2012Blanco, D. R., L. A. C. Bertollo, R. L. Lui, M. R. Vicari, V. P. Margarido, R. F. Artoni & O. Moreira Filho. 2012. A new technique for obtaining mitotic chromosome spreads from fishes in the field. Journal of Fish Biology, 81: 351-357.) after mitotic stimulation for 48 h (Lee & Elder, 1980Lee, M. R. & F. F. B. Elder. 1980. Yeast stimulation of bone marrow mitosis for cytogenetic investigations. Cytogenetics and Cell Genetics, 26: 36-40.). All experimental procedures and euthanasia of specimens were previously authorized by the Comitê de Ética em Uso de Animais da Universidade Estadual do Sudoeste da Bahia (CEUA/UESB, number 32/2013).
For karyotyping, the chromosomes were classified into metacentric (m), submetacentric (sm), subtelocentric (st) and acrocentric (a), according to Levan et al. (1964Levan, A., K. Fredga & A. A. Sandberg. 1964. Nomenclature for centromeric position on chromosomes. Hereditas, 52: 201-220.). The active nucleolar organizer regions (Ag-NORs) were detected by silver nitrate staining (Howell & Black, 1980Howell, W. M. & D. A. Black. 1980. Controlled silver-staining of nucleolus organizer regions with a protective colloidal developer: a 1-step method. Experientia, 36: 1014-1015.) and heterochromatin segments were visualized by C-banding (Sumner, 1972Sumner, A. T. 1972. A simple technique for demonstrating centromeric heterochromatin. Experimental Cell Research, 75: 304-306.). Base-specific fluorochrome staining using Chromomycin A3 (CMA3) and 4′6-diamidino-2-phenylindole (DAPI) were applied to detected GC- and AT-rich regions, respectively (Schweizer, 1980Schweizer, D. 1980. Simultaneous fluorescent staining of R bands and specific heterochromatic regions (DA-DAPI bands) in human chromosomes. Cytogenetics and Cell Genetics, 27: 190-193.).
Two repetitive DNA sequences isolated from Hoplias malabaricus (Bloch, 1794) were used as probes in FISH. The first probe comprised the 5S rDNA, which included 120 base pairs (bp) of coding sequence and 200 bp of non-transcribed spacer (Martins et al., 2006Martins, C., I. A. Ferreira, C. Oliveira, F. Foresti & P. M. Galetti Júnior. 2006. A tandemly repetitive centromeric DNA sequence of the fish Hoplias malabaricus (Characiformes: Erythrinidae) is derived from 5S rDNA. Genetica, 127: 133-141.). The second probe corresponded to a segment of 1,400 bp of 18S rDNA obtained by Polymerase Chain Reaction (PCR) (Cioffi et al., 2009aCioffi, M. B., C. Martins & L. A. C. Bertollo. 2009a. Comparative chromosome mapping of repetitive sequences. Implications for genomic evolution in the fish, Hoplias malabaricus . BMC Genetics, 10: 34(11p.).). Both probes were cloned in plasmidial vectors and propagated in competent cells of Escherichia coli DH5α (Invitrogen, San Diego, CA, USA). The 5S and 18S rDNA probes were labeled by nick translation with biotin-16-dUTP and digoxigenin-11-dUTP, respectively, following manufacturer's instructions (Roche, Mannheim, Germany).
The FISH experiments were performed as described by Pinkel et al. (1986Pinkel, D., T. Straume & J. W. Gray. 1986. Cytogenetic analysis using quantitative, high-sensitivity, fluorescence hybridization. Proceedings of the National Academy of Sciences of the United States of America, 83: 2934-2938.), using both probes simultaneously (double-FISH) under high stringency conditions (77%). The 5S rDNA probe was detected by conjugated fluorescein isothiocyanate-avidin (avidin-FITC) (Sigma, St. Louis, MO, USA) while the 18S rDNA signals were detected by anti-digoxigenin-Rhodamine conjugate (Roche, Mannheim, Germany). The chromosomes were counterstained with DAPI (1.2 µg/ml) in antifading solution (Vector, Burlingame, CA, USA) and analyzed under an epifluorescent microscope Olympus BX51 (Olympus Corporation, Ishikawa, Japan). The images were captured using the software CoolSNAP-Pro (Media Cybernetics).
Results
All specimens, independently on collection site or sex, shared a diploid number of 2n=50 with karyotypes composed of 8m+26sm+14st+2a (Fig. 2). The heterochromatin was preferentially distributed at pericentromeric regions of submetacentric and subtelocentric chromosomes in most populations (Fig. 3a). However, the samples from the Upper Contas River were characterized by the presence of C-bands at terminal regions of pairs 11 and 18 (Fig. 3b).
Representative C-banded karyotypes of Nematocharax venustus from (a) Almada, Gongogi 1, Gongogi 2, Gongogi 3, Jequitinhonha 1, Jequitinhonha 2 and (b) Upper Contas. Bar = 5 µm.
Invariably, Ag-NORs in N. venustus were detected at terminal regions on short arms of a single pair. Yet, the NOR-bearing pair differed among populations, since the Ag-NORs in specimens from the Upper Contas River were interspersed with C-bands on pair 18 (st) while the remaining samples revealed active ribosomal regions on pair 8 (sm) (Fig. 4). Likewise, GC-rich regions (CMA3 + and DAPI-) were observed in a single pair, being equivalent to Ag-NORs (Fig. 4).
The FISH with ribosomal probes confirmed the occurrence of a single NOR system in most populations. The only exception refers to one sample in the Jequitinhonha River basin (named Jequitinhonha 2), which presented additional 18S rDNA sites on long arms in one homologous from pair 8 and on short arms of a single chromosomes from pair 10. This procedure was also informative in revealing four distribution patterns of 18S and 5S rDNA in N. venustus (Fig. 4).
The first pattern, shared by specimens from the Almada River and some populations from the Contas (Gongogi 1) and Jequitinhonha River basins (Jequitinhonha 1), includes 18S rRNA genes at terminal region of short arms of a sm pair (equivalent to Ag-NORs) and 5S rRNA genes at interstitial region on short arms of two pairs (16 - sm, and 21 - st). The population from Gongogi 3 differs from this pattern by presenting heteromorphic 5S rDNA cistrons in pair 16, located at interstitial position on short arms of one chromosome and at pericentromeric region in the homologous (Fig. 4). The second pattern was similar that abovementioned, being observed in the population named as Gongogi 2. The only difference in this case refers to the presence of a single pair (16) bearing 5S rDNA at interstitial position on short arms (Fig. 4).
Chromosomes of distinct populations of Nematocharax venustus after silver nitrate staining (Ag-NORs), base-specific fluorochrome staining (CMA3/DA/DAPI) and FISH with 18S (magenta) and 5S rDNA probes (green). Bar = 5 µm.
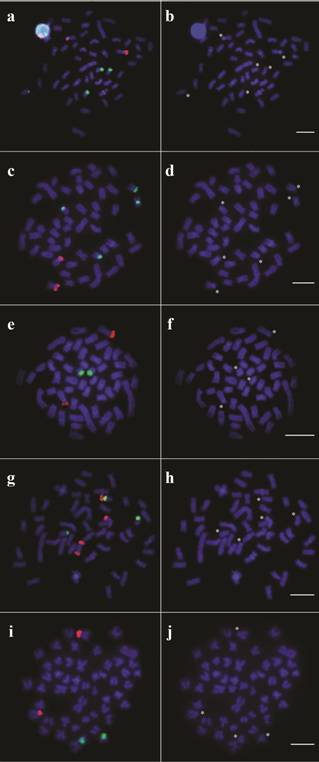
The population identified as Jequitinhonha 2 was characterized by a unique distribution of ribosomal genes, including bitelomeric 18S rDNA signals in one chromosome from pair 8. Moreover, this sample presented syntenic location of 18S and 5S rDNAs on short arms of one homologous from pair 10, being the latter closer to the centromeric region (Fig. 4). Another differential pattern was verified in the population from the Upper Contas River, since the specimens from this locality showed 18S rDNA sites on short arms of pair 18 while 5S rDNA regions were located interstitially on short arms of a single sm pair (16) (Fig. 4). The double-FISH metaphases for each different pattern observed, followed by the same metaphases stained with DAPI only, are shown in Figure 5.
Representative metaphases for each different pattern reported in populations of Nematocharax venustus showing the hybridization with 18S (magenta) and 5S rDNA probes (green) (double-FISH) and only DAPI staining for (a, b) Almada, Gongogi 1, and Jequitinhonha 1; (c, d) Gongogi 3; (e, f) Gongogi 2; (g, h) Jequitinhonha 2; and (i, j) Upper Contas. The asterisks indicate the chromosomes marked by FISH. Bars = 5 µm.
Discussion
In spite of being the most cytogenetically studied group of Neotropical ichthyofauna (Nirchio et al., 2014Nirchio, M., A. R. Rossi, F. Foresti & C. Oliveira. 2014. Chromosome evolution in fishes: a new challenging proposal from Neotropical species. Neotropical Ichthyology, 12: 761-770.), karyotypic data are still absent for many species and genera of Characiformes, such as Nematocharax . The karyotype macrostructure of N. venustus , including the modal diploid number of 2n=50, high number of biarmed chromosomes and a large metacentric pair (1st pair), is observed in several species of Characidae. For instance, this pattern has been reported in Astyanax scabripinnis (Jenyns, 1842) (Moreira Filho & Bertollo, 1991Moreira Filho, O. & L. A. C. Bertollo. 1991. Astyanax scabripinnis (Pisces, Characidae): a species complex. Revista Brasileira de Genética, 142: 331-357.), Hasemania nana (Lütken, 1875) (Moreira et al ., 2007Moreira, P. W. A., L. A. C. Bertollo & O. Moreira Filho. 2007. Comparative cytogenetics between three Characidae fish species from the São Francisco River basin. Caryologia, 60: 64-68.), Hyphessobrycon anisitsi (Eigenmann, 1907) (Mendes et al., 2011Mendes, M. M., R. Rosa, L. G. Caetano & A. L. Dias. 2011. Karyotype diversity of four species of the incertae sedis group (Characidae) from different hydrographic basins: analysis of AgNORs, CMA3 and 18S rDNA. Genetics and Molecular Research, 10: 3596-3608.), Hollandichthys multifasciatus (Eigenmann & Norris, 1900) (Balen et al., 2013Balen, R. E., R. B. Noleto, M. R. Vicari, R. F. Artoni & M. M. Cestari. 2013. Comparative cytogenetics among populations of Hollandichthys multifasciatus (Teleostei: Characidae). Zoological Science, 30: 105-109.) and Rhoadsia altipinna Fowler, 1911 (Romero et al., 2015Romero, O. S., C. Q. Abad, P. Q. Cordero, V. F. Sene, M. Nirchio & C. Oliveira. 2015. First description of the karyotype and localization of major and minor ribosomal genes in Rhoadsia altipinna Fowler, 1911 (Characiformes, Characidae) from Ecuador. Comparative Cytogenetics, 9: 271-280.), being considered a plesiomorphic condition of characins (Scheel, 1973Scheel, J. J. 1973. Fish chromosomes and their evolution. Charlottenlund, Internal report of Danmarks Akvarium. 22p.; Morelli et al., 1983Morelli, S., L. A. C. Bertollo, F. Foresti, O. Moreira Filho & S. A. Toledo Filho. 1983. Cytogenetic considerations on the genus Astyanax (Pisces, Characidae). I. Kariotypic variability. Caryologia, 36: 235-244.). The presence of distinct types of chromosomes (m, sm, st, and a) without variation in diploid number suggests the occurrence of pericentric inversions, another common rearrangement in characins (Medrado et al., 2015Medrado, A. S., P. R. A. M. Affonso, P. L. S. Carneiro, M. R. Vicari, R. F. Artoni & M. A. Costa. 2015. Allopatric divergence in Astyanax aff. fasciatus Cuvier, 1819 (Characidae, Incertae sedis ) inferred from DNA mapping and chromosomes. Zoologischer Anzeiger, 257: 119-129.).
In the case of taxa with similar chromosomal number and morphology, the analysis of heterochromatin distribution might be helpful to provide cytotaxonomic or population markers, as previously shown in other Characiformes (Galetti Júnior et al., 1991Galetti Júnior, P. M., A. C. G. Cesar & P. C. Venere. 1991. Heterochromatin and NORs variability in Leporinus fish (Anostomidae, Characiformes). Caryologia, 44: 287-292.; Mantovani et al., 2000Mantovani, M., L. D. S. Abel, C. A. Mestriner & O. Moreira Filho. 2000. Accentuated polymorphism of heterochromatin and nucleolar organizer regions in Astyanax scabripinnis (Pisces, Characidae): tools for understanding karyotypic evolution. Genetica, 109: 161-168.; Jacobina et al., 2009Jacobina, U. P., P. R. A. M. Affonso, P. L. S. Carneiro & J. A. Dergam. 2009. Biogeography and comparative cytogenetics between two populations of Hoplias malabaricus (Bloch, 1794) (Ostariophysi: Erythrinidae) from coastal basins in the State of Bahia, Brazil. Neotropical Ichthyology, 7: 617-622.). However, the C-bands were restricted to pericentromeric region and NORs in most populations of N. venustus , following a common trend in fish (Imai, 1991Imai, H. T. 1991. Mutability of constitutive heterochromatin (C-bands) during eukaryotic chromosomal evolution and their cytological meaning. The Japanese Journal of Genetics, 66: 635-661.; Aguilar & Galetti Júnior, 2008Aguilar, C. T. & P. M. Galetti Júnior. 2008. Chromosome mapping of 5S rRNA genes differentiates Brazilian populations of Leporellus vittatus (Anostomidae, Characiformes). Genetics and Molecular Biology, 31: 188-194.; Rosa et al., 2009Rosa, R., M. Rubert, L. R. Malabarba, I. C. M. Santos & L. G. Caetano. 2009. Cytogenetic analysis of Astyanax laticeps (Cope, 1894) (Ostariophysi: Characidae) from the laguna dos Patos system. Neotropical Ichthyology, 7: 601-605.). The only exception was the population from the Upper Contas River, which presented terminal heterochromatic segments in two pairs (Fig. 3b). This unique C-banding pattern indicates that this population is genetically divergent, as further discussed.
On the other hand, microstructural interpopulation differences were observed by mapping the ribosomal genes in N. venustus , particularly in relation to the samples from Jequitinhonha 2 and Upper Contas. Even though most populations presented active rDNA sites interspersed with GC-rich sites (CMA3 +), a common feature in fish (e.g. , Verma et al., 2011Verma, J., W. S. Lakra, B. Kushwaha, M. Sirajuddin, N. S. Nagpure & R. Kumar. 2011. Characterization of two freshwater silurid catfish using conventional and molecular cytogenetic techniques. Journal of Genetics, 90: 319-322.; Santos et al., 2012Santos, A. R., M. Rubert, L. G. Caetano & A. L. Dias. 2012. Sympatric occurrence of four cytotypes and one extra chromosome in Bryconamericus ecai (Characidae): 18S rDNA polymorphism and heterochromatin composition. Hereditas, 149: 24-33.), additional 18S rDNA cistrons were identified by FISH in specimens from Jequitinhonha 2, characterizing a multiple NOR system. In the case of individuals from Upper Contas, the NORs were located on the first st pair while submetacentric NOR-bearing pairs were observed in the other populations of N. venustus (Fig. 4). Thus, the distribution of 18S rDNA cistrons apparently represents cytogenetic markers for the populations from the Contas River basin.
These results corroborate the dynamic evolution of NORs in Characidae (Galetti Júnior, 1998Galetti Júnior, P. M. 1998. Chromosome diversity in Neotropical fishes: NOR studies. Italian Journal of Zoology, 65(S1): 53-56.). Indeed, even though some genera of Characidae share single NORs, a basal condition in teleosteans (Gornung, 2013Gornung, E. 2013. Twenty years of physical mapping of major ribosomal RNA genes across the Teleosts: a review of research. Cytogenetic and Genome Research, 141: 90-102.), as reported in Tetragonopterus Cuvier (Alberdi & Fenocchio, 1997Alberdi, A. J. & A. S. Fenocchio. 1997. Karyotypes of five Tetragonopterinae species (Pisces, Characidae) from Argentina. Cytologia, 62: 171-176.), Moenkhausia Eigenmann (Castro & Júlio Júnior, 2002Castro, A. L. B. P. & H. F. Júlio Júnior. 2002. Karyotype relationships among species of the subfamily Tetragonopterinae (Pisces, Characidae): cytotaxonomy and evolution aspects. Cytologia, 67: 329-336.), Bryconamericus Eigenmann (Marques et al., 2003Marques, T. R. P., L. G. Caetano & A. L. Dias. 2003. Cytogenetic characterization of a population of Bryconamericus aff. iheringii (Characidae, Tetragonopterinae). Genetics and Molecular Biology, 26: 145-149.) and Piabina Reinhardt (Pazian et al., 2012Pazian, M. F., L. H. G. Pereira, C. K. S. Dias, C. Oliveira & F. Foresti. 2012. Cytogenetic and molecular markers reveal the complexity of the genus Piabina Reinhardt, 1867 (Characiformes: Characidae). Neotropical Ichthyology, 10: 329-340.), most species in this family present multiple NORs (Kavalco & Moreira Filho, 2003Kavalco, K. F. & O. Moreira Filho. 2003. Cytogenetical analyses in four species of the genus Astyanax (Pisces, Characidae) from Paraíba do Sul River basin. Caryologia, 56: 453-461.).
The remarkable variation of both position and number of ribosomal sites in characins and other Characiformes might reflect the polyphyletism of this fish group (Peres et al., 2008Peres, W. A. M., L. A. C. Bertollo & O. Moreira Filho. 2008. Physical mapping of the 18S and 5S ribosomal genes in nine Characidae species (Teleostei, Characiformes). Genetics and Molecular Biology, 31(S1): 222-226.) and/or the association of rDNA with transposable elements (TE) that mobilize adjacent ribosomal sequences (Cioffi et al., 2010Cioffi, M. B., C. Martins & L. A. C. Bertollo. 2010. Chromosome spreading of associated transposable elements and ribosomal DNA in the fish Erythrinus erythrinus . Implications for genome change and karyoevolution in fish. BMC Evolutionary Biology, 10: 271 (9p.).). Reinforcing the polymorphic nature of rRNA genes in Characidae, the mapping of 18S and 5S rDNA by FISH revealed clear interpopulation cytogenetic divergence in N. venustus .
Usually, the 18S and 5S rDNA sites in Neotropical fish are located on distinct chromosomal pairs (Galetti Júnior & Martins, 2004Galetti Júnior, P. M. & C. Martins. 2004. Contribuição da hibridização in situ para o conhecimento dos cromossomos de peixes. Pp. 61-88. In: Guerra, M. (Ed.). FISH: conceitos e aplicações na Citogenética. Ribeirão Preto, Sociedade Brasileira de Genética.). This pattern was observed in most populations but Jequitinhonha 2, since double-FISH revealed syntenic 18S and 5S rDNA cistrons in a homologous from pair 10 (Fig. 4). However, this characteristic may be an intrapopulation variation found in Jequitinhonha 2, since the double-FISH was obtained for a single individual from this collection site. Other reports of synteny between both families of ribosomal genes are available in some genera of Characiformes, like Astyanax Baird & Girard (Castro et al., 2014bCastro, J. P., M. O. Moura, O. Moreira Filho, O. A. Shibatta, M. H. Santos, V. Nogaroto, M. R. Vicari, M. C. Almeida & R. F. Artoni. 2014b. Evidence of incipient speciation in Astyanax scabripinnis species complex (Teleostei: Characidae). Neotropical Ichthyology, 12: 429-438.), Prochilodus Agassiz (Vicari et al., 2006Vicari, M. R., M. C. Almeida, L. A. C. Bertollo, O. Moreira Filho & R. F. Artoni. 2006. Cytogenetic analysis and chromosomal characteristics of the polymorphic 18S rDNA in the fish Prochilodus lineatus (Characiformes, Prochilodontidae). Genetics and Molecular Biology, 29: 621-625.), Triportheus Cope (Diniz et al., 2009Diniz, D., A. Laudicina & L. A. C. Bertollo. 2009. Chromosomal location of 18S and 5S rDNA sites in Triportheus fish species (Characiformes, Characidae). Genetics and Molecular Biology, 32: 37-41.), and Hoplias Gill (Cioffi et al., 2009bCioffi, M. B., C. Martins, L. Centofante, U. Jacobina & L. A. C. Bertollo. 2009b. Chromosomal variability among allopatric populations of Erythrinidae fish Hoplias malabaricus: mapping of three classes of repetitive DNAs. Cytogenetic and Genome Research, 125: 132-141.). It should be pointed out that active Ag-NORs in the population from Jequitinhonha 2 were observed only in pair 8. Nonetheless, syntenic ribosomal genes were detected by FISH in one chromosome from pair 10. The lack of hybridization signal in the other homologous could indicate differences in the number of copies for this gene between chromosomes of pair 10 possibly related to unequal exchanges.
In addition, the location of 18S rDNA sites at terminal chromosomal regions is widespread in teleosteans. Apparently, this behavior favors the rearrangement of NORs without interfering with other genetic linkage groups (Hanson et al., 1996Hanson, R. E., M. N. Islam-Faridi, E. A. Percival, C. F. Crane, Y. Ji, T. D. McKnight, D. M. Stelly & H. J. Price. 1996. Distribution of 5S and 18S-28S rDNA loci in a tetraploid cotton (Gossypium hirsutum L.) and its putative diploid ancestors. Chromosoma, 105: 55-61.), besides promoting the redistribution of ribosomal genes by the proximity of telomeres in the interphasic nuclei (Gornung, 2013Gornung, E. 2013. Twenty years of physical mapping of major ribosomal RNA genes across the Teleosts: a review of research. Cytogenetic and Genome Research, 141: 90-102.). Such distribution of 18S rRNA genes in N. venustus could account for the putative transposition of some copies of this rDNA in one homologous of pair 8 in the population from Jequitinhonha 2, giving rise to the bitelomeric NORs (Fig. 4). Again, the presence of 18S rDNA sites in both telomeres have already been reported in other Characiformes characterized by accentuated chromosomal variation, such as A. fasciatus (Cuvier, 1819) (Fernandes et al., 2009Fernandes, C. A., I. C. Martins Santos & D. Bailly. 2009. Karyotype analysis and mapping of the 18S and 5S ribosomal genes in Astyanax fasciatus (Teleostei, Characiformes) from Paranapanema river basin. Cytologia, 74: 295-300.), A. scabripinnis (Mantovani et al., 2005Mantovani, M., L. D. S. Abel & O. Moreira Filho. 2005. Conserved 5S and variable 45S rDNA chromosomal localisation revealed by FISH in Astyanax scabripinnis (Pisces, Characidae). Genetica, 123: 211-216.), A. hastatus Myers, 1928 (Kavalco et al., 2009Kavalco, K. F., K. O. Brandão, R. Pazza & L. F. A. Toledo. 2009. Astyanax hastatus Myers, 1928 (Teleostei, Characidae): a new species complex within the genus Astyanax ?. Genetics and Molecular Biology, 32: 477-483.), and H. malabaricus (Cioffi et al., 2009aCioffi, M. B., C. Martins & L. A. C. Bertollo. 2009a. Comparative chromosome mapping of repetitive sequences. Implications for genomic evolution in the fish, Hoplias malabaricus . BMC Genetics, 10: 34(11p.).).
Differently from 18S rDNA, the 5S rDNA sites in fish are usually located at interstitial and euchromatic chromosomal regions (Martins & Galetti Júnior, 2001Martins, C. & P. M. Galetti Júnior. 2001. Two 5S rDNA arrays in Neotropical fish species: is it a general rule for fishes?. Genetica, 111: 439-446.), as reported in N. venustus . This condition seems to reduce the genomic dispersal and reorganization of 5S rRNA genes when compared to NORs (Martins & Wasko, 2004Martins, C. & A. P. Wasko. 2004. Organization and evolution of 5S ribosomal DNA in the fish genome. Pp. 335-363. In: Williams, C. R. (Ed.). Focus on genome research. Hauppauge, Nova Science Publishers.). Nonetheless, a possible heterozygous pericentric inversion, encompassing the 5S rDNA cluster, is suggested for the heteromorphic pair 16 in individuals from Gongogi 3 (Fig. 4), thereby explaining the differential position of 5S rDNA signals between homologous. Moreover, numerical variation of 5S rDNA signals was also detected in the present study. While most populations of N. venustus shared two pairs bearing 5S rRNA genes, as commonly reported in characins (Kavalco et al., 2011Kavalco, K. F., R. Pazza, K. O. Brandão, C. Garcia & L. F. A. Toledo. 2011. Comparative cytogenetics and molecular phylogeography in the group Astyanax altiparanae - Astyanax aff. bimaculatus (Teleostei, Characidae). Cytogenetic and Genome Research, 134: 108-119.), the individuals from Gongogi 2 and Upper Contas were characterized by a single 5S rDNA-bearing pair. These data show the importance of refined methodologies of chromosomal mapping in revealing chromosomal rearrangements that otherwise would remain imperceptible, mainly due to the homogeneous macrokaryotypic structure in this species.
Along with the cytogenetic particularities, the specimens from the Upper Contas River, in Diamantina Plateau, are also morphologically distinctive once they present slender bodies and short rays in pelvic fins of males (data not shown), being likely to correspond to a new and undescribed species of Nematocharax . Reproductive isolation and speciation in the Upper Contas River are favored by the geological processes in the eastern region of Brazil, which includes the Diamantina Plateau and the Espinhaço Hills (Derby, 1906Derby, O. A. 1906. The Serra do Espinhaço, Brazil. The Journal of Geology, 14: 374-401.). In fact, divergent species associated to the isolation promoted by these biogeographic barriers have been reported even in animals of high dispersal potential, like birds (Vasconcelos et al., 2012Vasconcelos, M. F., A. V. Chaves & F. R. Santos. 2012. First record of Augastes scutatus for Bahia refines the location of a purported barrier promoting speciation in the Espinhaço range, Brazil. Revista Brasileira de Ornitologia, 20: 443-446.).
Furthermore, karyotypic novelties can evolve independently within relatively short periods in populations of small characins, like N. venustus , because their restricted dispersal and short life cycles (Castro, 1999Castro, R. M. C. 1999. Evolução da ictiofauna de riachos sul-americanos: padrões gerais e possíveis processos causais. Pp. 139-155. In: Caramaschi, E. P., R. Mazzoni & P. R. Peres Neto (Eds.). Ecologia de peixes de riachos. Rio de Janeiro, Programa de Pós-Graduação em Ecologia, Instituto de Biologia, Universidade Federal do Rio de Janeiro. (Oecologia Brasiliensis, v. 6).) lead to reduced gene flow and rapid establishment of chromosomal rearrangements (Centofante et al., 2006Centofante, L., L. A. C. Bertollo & O. Moreira Filho. 2006. Chromosomal differentiation between populations of Oligosarcus hepsetus (Teleostei, Characidae) from small tributaries at opposite margins of the Paraiba do Sul River (Brazil). Brazilian Archives of Biology and Technology, 49: 981-987.; Pazza & Kavalco, 2007Pazza, R. & K. F. Kavalco. 2007. Chromosomal evolution in the Neotropical characin Astyanax (Teleostei, Characidae). The Nucleus, 50: 519-543.). Therefore, isolated and small populations in headwaters, like the one from the Upper Contas River, are more susceptible to genetic drift effects, potentially leading to the fixation of unique chromosomal features at random. Similarly, the populations of N. venustus from the Jequitinhonha River basin are endangered due to the increased human activities (Menezes & Lima, 2008Menezes, N. A. & F. C. T. Lima. 2008. Nematocharax venustus Weitzman, Menezes & Britski, 1986. Pp. 85-86. In: Machado, A. B. M., G. M. Drummond & A. P. Paglia (Eds.). Livro Vermelho da Fauna Brasileira Ameaçada de Extinção. Brasília, MMA. v. 2, 888p.), which might have reduced their effective population sizes. Thus, inbreeding and genetic drift effects related to bottleneck events might have increased the probability of fixing distinct chromosomal variants (Hedrick, 1981Hedrick, P. W. 1981. The establishment of chromosomal variants. Evolution, 322-332.).
In conclusion, the present results increase the karyotypic reports in characins from coastal basins in Eastern Atlantic, a region recognized by high levels of endemism combined with severe human impacts (Camelier & Zanata, 2014Camelier, P. & A. M. Zanata. 2014. Biogeography of freshwater fishes from the Northeastern Mata Atlântica freshwater ecoregion: distribution, endemism, and area relationships. Neotropical Ichthyology, 12: 683-698.). The comparative cytogenetic analysis provided evidence of allopatric chromosomal variation in N. venustus along its range, suggesting the presence of populations with distinct evolutionary histories that should be further investigated using other genetic markers. Otherwise, species or structured populations can be lost because they lack appropriate conservation management plans.
Acknowledgements
The authors are grateful to Fundação de Amparo à Pesquisa do Estado da Bahia (FAPESB) for the financial support (grants PNE0019/2011, and RED0009/2013) and Instituto Chico Mendes de Conservação da Biodiversidade (ICMBio) for authorizing the collection of specimens.
References
- Aguilar, C. T. & P. M. Galetti Júnior. 2008. Chromosome mapping of 5S rRNA genes differentiates Brazilian populations of Leporellus vittatus (Anostomidae, Characiformes). Genetics and Molecular Biology, 31: 188-194.
- Alberdi, A. J. & A. S. Fenocchio. 1997. Karyotypes of five Tetragonopterinae species (Pisces, Characidae) from Argentina. Cytologia, 62: 171-176.
- Almeida, J. S., P. R. A. M. Affonso, D. Diniz, P. L. S. Carneiro & A. L. Dias. 2013. Chromosomal variation in the tropical armored catfish Callichthys callichthys (Siluriformes, Callichthyidae): implications for conservation and taxonomy in a species complex from a Brazilian hotspot. Zebrafish, 10: 451-458.
- Balen, R. E., R. B. Noleto, M. R. Vicari, R. F. Artoni & M. M. Cestari. 2013. Comparative cytogenetics among populations of Hollandichthys multifasciatus (Teleostei: Characidae). Zoological Science, 30: 105-109.
- Bellafronte, E., M. O. Schemberger, O. Moreira Filho, M. C. Almeida, R. F. Artoni, V. P. Margarido & M. R. Vicari. 2010. Chromosomal markers in Parodontidae: an analysis of new and reviewed data with phylogenetic inferences. Reviews in Fish Biology and Fisheries, 21: 559-570.
- Bitencourt, J. A., P. R. A. M. Affonso, L. Giuliano-Caetano & A. L. Dias. 2011. Identification of distinct evolutionary units in allopatric populations of Hypostomus cf. wuchereri Günther, 1864 (Siluriformes: Loricariidae): karyotypic evidence. Neotropical Ichthyology, 9: 317-324.
- Blanco, D. R., L. A. C. Bertollo, R. L. Lui, M. R. Vicari, V. P. Margarido, R. F. Artoni & O. Moreira Filho. 2012. A new technique for obtaining mitotic chromosome spreads from fishes in the field. Journal of Fish Biology, 81: 351-357.
- Blanco, D. R., R. L. Lui, L. A. C. Bertollo, V. P. Margarido & O. Moreira Filho. 2010. Karyotypic diversity between allopatric populations of the group Hoplias malabaricus (Characiformes: Erythrinidae): evolutionary and biogeographic considerations. Neotropical Ichthyology, 8: 361-368.
- Bragança, P. H. N., M. A. Barbosa & J. L. Mattos. 2013. A new Nematocharax species from the middle Contas River basin, Northeastern Brazil (Characiformes: Characidae). Vertebrate Zoology, 63: 3-8.
- Camelier, P. & A. M. Zanata. 2014. Biogeography of freshwater fishes from the Northeastern Mata Atlântica freshwater ecoregion: distribution, endemism, and area relationships. Neotropical Ichthyology, 12: 683-698.
- Castro, A. L. B. P. & H. F. Júlio Júnior. 2002. Karyotype relationships among species of the subfamily Tetragonopterinae (Pisces, Characidae): cytotaxonomy and evolution aspects. Cytologia, 67: 329-336.
- Castro, J. P., M. O. Moura, O. Moreira Filho, O. A. Shibatta, M. H. Santos, V. Nogaroto, M. R. Vicari, M. C. Almeida & R. F. Artoni. 2014a. Diversity of the Astyanax scabripinnis species complex (Teleostei: Characidae) in the Atlantic Forest, Brazil: species limits and evolutionary inferences. Reviews in Fish Biology and Fisheries, 25: 231-244.
- Castro, J. P., M. O. Moura, O. Moreira Filho, O. A. Shibatta, M. H. Santos, V. Nogaroto, M. R. Vicari, M. C. Almeida & R. F. Artoni. 2014b. Evidence of incipient speciation in Astyanax scabripinnis species complex (Teleostei: Characidae). Neotropical Ichthyology, 12: 429-438.
- Castro, R. M. C. 1999. Evolução da ictiofauna de riachos sul-americanos: padrões gerais e possíveis processos causais. Pp. 139-155. In: Caramaschi, E. P., R. Mazzoni & P. R. Peres Neto (Eds.). Ecologia de peixes de riachos. Rio de Janeiro, Programa de Pós-Graduação em Ecologia, Instituto de Biologia, Universidade Federal do Rio de Janeiro. (Oecologia Brasiliensis, v. 6).
- Centofante, L., L. A. C. Bertollo & O. Moreira Filho. 2006. Chromosomal differentiation between populations of Oligosarcus hepsetus (Teleostei, Characidae) from small tributaries at opposite margins of the Paraiba do Sul River (Brazil). Brazilian Archives of Biology and Technology, 49: 981-987.
- Cioffi, M. B., C. Martins & L. A. C. Bertollo. 2009a. Comparative chromosome mapping of repetitive sequences. Implications for genomic evolution in the fish, Hoplias malabaricus . BMC Genetics, 10: 34(11p.).
- Cioffi, M. B., C. Martins & L. A. C. Bertollo. 2010. Chromosome spreading of associated transposable elements and ribosomal DNA in the fish Erythrinus erythrinus . Implications for genome change and karyoevolution in fish. BMC Evolutionary Biology, 10: 271 (9p.).
- Cioffi, M. B., C. Martins, L. Centofante, U. Jacobina & L. A. C. Bertollo. 2009b. Chromosomal variability among allopatric populations of Erythrinidae fish Hoplias malabaricus: mapping of three classes of repetitive DNAs. Cytogenetic and Genome Research, 125: 132-141.
- Derby, O. A. 1906. The Serra do Espinhaço, Brazil. The Journal of Geology, 14: 374-401.
- Diniz, D., A. Laudicina & L. A. C. Bertollo. 2009. Chromosomal location of 18S and 5S rDNA sites in Triportheus fish species (Characiformes, Characidae). Genetics and Molecular Biology, 32: 37-41.
- Eschmeyer, W. N. & J. D. Fong. 2015. Species by family/subfamily. Species by family/subfamily in the catalog of fishes. Available from: Available from: http://researcharchive.calacademy.org/research/ichthyology/catalog/speciesbyfamily.asp (05 February 2016).
» http://researcharchive.calacademy.org/research/ichthyology/catalog/speciesbyfamily.asp - Fernandes, C. A., I. C. Martins Santos & D. Bailly. 2009. Karyotype analysis and mapping of the 18S and 5S ribosomal genes in Astyanax fasciatus (Teleostei, Characiformes) from Paranapanema river basin. Cytologia, 74: 295-300.
- Galetti Júnior, P. M. 1998. Chromosome diversity in Neotropical fishes: NOR studies. Italian Journal of Zoology, 65(S1): 53-56.
- Galetti Júnior, P. M., A. C. G. Cesar & P. C. Venere. 1991. Heterochromatin and NORs variability in Leporinus fish (Anostomidae, Characiformes). Caryologia, 44: 287-292.
- Galetti Júnior, P. M. & C. Martins. 2004. Contribuição da hibridização in situ para o conhecimento dos cromossomos de peixes. Pp. 61-88. In: Guerra, M. (Ed.). FISH: conceitos e aplicações na Citogenética. Ribeirão Preto, Sociedade Brasileira de Genética.
- Gornung, E. 2013. Twenty years of physical mapping of major ribosomal RNA genes across the Teleosts: a review of research. Cytogenetic and Genome Research, 141: 90-102.
- Hanson, R. E., M. N. Islam-Faridi, E. A. Percival, C. F. Crane, Y. Ji, T. D. McKnight, D. M. Stelly & H. J. Price. 1996. Distribution of 5S and 18S-28S rDNA loci in a tetraploid cotton (Gossypium hirsutum L.) and its putative diploid ancestors. Chromosoma, 105: 55-61.
- Hedrick, P. W. 1981. The establishment of chromosomal variants. Evolution, 322-332.
- Howell, W. M. & D. A. Black. 1980. Controlled silver-staining of nucleolus organizer regions with a protective colloidal developer: a 1-step method. Experientia, 36: 1014-1015.
- Imai, H. T. 1991. Mutability of constitutive heterochromatin (C-bands) during eukaryotic chromosomal evolution and their cytological meaning. The Japanese Journal of Genetics, 66: 635-661.
- Jacobina, U. P., P. R. A. M. Affonso, P. L. S. Carneiro & J. A. Dergam. 2009. Biogeography and comparative cytogenetics between two populations of Hoplias malabaricus (Bloch, 1794) (Ostariophysi: Erythrinidae) from coastal basins in the State of Bahia, Brazil. Neotropical Ichthyology, 7: 617-622.
- Kavalco, K. F., K. O. Brandão, R. Pazza & L. F. A. Toledo. 2009. Astyanax hastatus Myers, 1928 (Teleostei, Characidae): a new species complex within the genus Astyanax ?. Genetics and Molecular Biology, 32: 477-483.
- Kavalco, K. F. & O. Moreira Filho. 2003. Cytogenetical analyses in four species of the genus Astyanax (Pisces, Characidae) from Paraíba do Sul River basin. Caryologia, 56: 453-461.
- Kavalco, K. F., R. Pazza, K. O. Brandão, C. Garcia & L. F. A. Toledo. 2011. Comparative cytogenetics and molecular phylogeography in the group Astyanax altiparanae - Astyanax aff. bimaculatus (Teleostei, Characidae). Cytogenetic and Genome Research, 134: 108-119.
- Lee, M. R. & F. F. B. Elder. 1980. Yeast stimulation of bone marrow mitosis for cytogenetic investigations. Cytogenetics and Cell Genetics, 26: 36-40.
- Levan, A., K. Fredga & A. A. Sandberg. 1964. Nomenclature for centromeric position on chromosomes. Hereditas, 52: 201-220.
- Mantovani, M., L. D. S. Abel, C. A. Mestriner & O. Moreira Filho. 2000. Accentuated polymorphism of heterochromatin and nucleolar organizer regions in Astyanax scabripinnis (Pisces, Characidae): tools for understanding karyotypic evolution. Genetica, 109: 161-168.
- Mantovani, M., L. D. S. Abel & O. Moreira Filho. 2005. Conserved 5S and variable 45S rDNA chromosomal localisation revealed by FISH in Astyanax scabripinnis (Pisces, Characidae). Genetica, 123: 211-216.
- Marques, T. R. P., L. G. Caetano & A. L. Dias. 2003. Cytogenetic characterization of a population of Bryconamericus aff. iheringii (Characidae, Tetragonopterinae). Genetics and Molecular Biology, 26: 145-149.
- Martins, C., I. A. Ferreira, C. Oliveira, F. Foresti & P. M. Galetti Júnior. 2006. A tandemly repetitive centromeric DNA sequence of the fish Hoplias malabaricus (Characiformes: Erythrinidae) is derived from 5S rDNA. Genetica, 127: 133-141.
- Martins, C. & P. M. Galetti Júnior. 2001. Two 5S rDNA arrays in Neotropical fish species: is it a general rule for fishes?. Genetica, 111: 439-446.
- Martins, C. & A. P. Wasko. 2004. Organization and evolution of 5S ribosomal DNA in the fish genome. Pp. 335-363. In: Williams, C. R. (Ed.). Focus on genome research. Hauppauge, Nova Science Publishers.
- Medrado, A. S., P. R. A. M. Affonso, P. L. S. Carneiro, M. R. Vicari, R. F. Artoni & M. A. Costa. 2015. Allopatric divergence in Astyanax aff. fasciatus Cuvier, 1819 (Characidae, Incertae sedis ) inferred from DNA mapping and chromosomes. Zoologischer Anzeiger, 257: 119-129.
- Mendes, M. M., R. Rosa, L. G. Caetano & A. L. Dias. 2011. Karyotype diversity of four species of the incertae sedis group (Characidae) from different hydrographic basins: analysis of AgNORs, CMA3 and 18S rDNA. Genetics and Molecular Research, 10: 3596-3608.
- Menezes, N. A. & F. C. T. Lima. 2008. Nematocharax venustus Weitzman, Menezes & Britski, 1986. Pp. 85-86. In: Machado, A. B. M., G. M. Drummond & A. P. Paglia (Eds.). Livro Vermelho da Fauna Brasileira Ameaçada de Extinção. Brasília, MMA. v. 2, 888p.
- Menezes, N. A., A. M. Zanata & P. Camelier. 2015. Nematocharax costai Bragança, Barbosa & Mattos a junior synonym of Nematocharax venustus Weitzman, Menezes & Britski (Teleostei: Characiformes: Characidae). Zootaxa, 3920: 453-462.
- Moreira, P. W. A., L. A. C. Bertollo & O. Moreira Filho. 2007. Comparative cytogenetics between three Characidae fish species from the São Francisco River basin. Caryologia, 60: 64-68.
- Moreira Filho, O. & L. A. C. Bertollo. 1991. Astyanax scabripinnis (Pisces, Characidae): a species complex. Revista Brasileira de Genética, 142: 331-357.
- Morelli, S., L. A. C. Bertollo, F. Foresti, O. Moreira Filho & S. A. Toledo Filho. 1983. Cytogenetic considerations on the genus Astyanax (Pisces, Characidae). I. Kariotypic variability. Caryologia, 36: 235-244.
- Netto, M. R. C. B., E. Pauls & P. R. A. M. Affonso. 2007. A standard protocol for obtaining fish chromosomes under post-mortem conditions. Micron, 38: 214-217.
- Nirchio, M., A. R. Rossi, F. Foresti & C. Oliveira. 2014. Chromosome evolution in fishes: a new challenging proposal from Neotropical species. Neotropical Ichthyology, 12: 761-770.
- Pazian, M. F., L. H. G. Pereira, C. K. S. Dias, C. Oliveira & F. Foresti. 2012. Cytogenetic and molecular markers reveal the complexity of the genus Piabina Reinhardt, 1867 (Characiformes: Characidae). Neotropical Ichthyology, 10: 329-340.
- Pazza, R. & K. F. Kavalco. 2007. Chromosomal evolution in the Neotropical characin Astyanax (Teleostei, Characidae). The Nucleus, 50: 519-543.
- Peres, W. A. M., L. A. C. Bertollo & O. Moreira Filho. 2008. Physical mapping of the 18S and 5S ribosomal genes in nine Characidae species (Teleostei, Characiformes). Genetics and Molecular Biology, 31(S1): 222-226.
- Pinkel, D., T. Straume & J. W. Gray. 1986. Cytogenetic analysis using quantitative, high-sensitivity, fluorescence hybridization. Proceedings of the National Academy of Sciences of the United States of America, 83: 2934-2938.
- Piscor, D., A. L. Alves & P. P. P. Maltempi. 2015. Chromosomal microstructure diversity in three Astyanax (Characiformes, Characidae) species: comparative analysis of the chromosomal locations of the 18S and 5S rDNAs. Zebrafish, 12: 81-90.
- Romero, O. S., C. Q. Abad, P. Q. Cordero, V. F. Sene, M. Nirchio & C. Oliveira. 2015. First description of the karyotype and localization of major and minor ribosomal genes in Rhoadsia altipinna Fowler, 1911 (Characiformes, Characidae) from Ecuador. Comparative Cytogenetics, 9: 271-280.
- Rosa, R., M. Rubert, L. R. Malabarba, I. C. M. Santos & L. G. Caetano. 2009. Cytogenetic analysis of Astyanax laticeps (Cope, 1894) (Ostariophysi: Characidae) from the laguna dos Patos system. Neotropical Ichthyology, 7: 601-605.
- Santos, A. R., M. Rubert, L. G. Caetano & A. L. Dias. 2012. Sympatric occurrence of four cytotypes and one extra chromosome in Bryconamericus ecai (Characidae): 18S rDNA polymorphism and heterochromatin composition. Hereditas, 149: 24-33.
- Scheel, J. J. 1973. Fish chromosomes and their evolution. Charlottenlund, Internal report of Danmarks Akvarium. 22p.
- Schweizer, D. 1980. Simultaneous fluorescent staining of R bands and specific heterochromatic regions (DA-DAPI bands) in human chromosomes. Cytogenetics and Cell Genetics, 27: 190-193.
- Sumner, A. T. 1972. A simple technique for demonstrating centromeric heterochromatin. Experimental Cell Research, 75: 304-306.
- Utsunomia, R., J. C. P. Alves, G. J. C. Silva, F. F. Mendonça, P. C. Scacchetti, C. Oliveira & F. Foresti. 2014. Molecular and cytogenetic analyses of cryptic species within the Synbranchus marmoratus Bloch, 1795 (Synbranchiformes: Synbranchidae) grouping: species delimitations, karyotypic evolution and intraspecific diversification. Neotropical Ichthyology, 12: 903-911.
- Vari, R. P. &L. R. Malabarba. 1998. Neotropical Ichthyology: an overview. Pp. 1-11. In: Malabarba, L. R., R. E. Reis, R. P. Vari, Z. M. S. Lucena & C. A. S. Lucena (Eds.). Phylogeny and classification of Neotropical fishes. Porto Alegre, Edipucrs, 603p.
- Vasconcelos, M. F., A. V. Chaves & F. R. Santos. 2012. First record of Augastes scutatus for Bahia refines the location of a purported barrier promoting speciation in the Espinhaço range, Brazil. Revista Brasileira de Ornitologia, 20: 443-446.
- Verma, J., W. S. Lakra, B. Kushwaha, M. Sirajuddin, N. S. Nagpure & R. Kumar. 2011. Characterization of two freshwater silurid catfish using conventional and molecular cytogenetic techniques. Journal of Genetics, 90: 319-322.
- Vicari, M. R., M. C. Almeida, L. A. C. Bertollo, O. Moreira Filho & R. F. Artoni. 2006. Cytogenetic analysis and chromosomal characteristics of the polymorphic 18S rDNA in the fish Prochilodus lineatus (Characiformes, Prochilodontidae). Genetics and Molecular Biology, 29: 621-625.
- Weitzman, S. H. & W. L. Fink. 1983. Relationships of the neon tetras, a group of South American freshwater fishes (Teleostei, Characidae), with comments on the phylogeny of New World Characiforms. Bulletin of the Museum of Comparative Zoology, 150: 339-395.
- Weitzman, S. H., N. A. Menezes & H. A. Britski. 1986. Nematocharax venustus , a new genus and species of fish from the rio Jequitinhonha, Minas Gerais, Brazil (Teleostei: Characidae). Proceedings of the Biological Society of Washington, 99: 335-346.
Data availability
Data citations
Eschmeyer, W. N. & J. D. Fong. 2015. Species by family/subfamily. Species by family/subfamily in the catalog of fishes. Available from: Available from: http://researcharchive.calacademy.org/research/ichthyology/catalog/speciesbyfamily.asp (05 February 2016).
Publication Dates
-
Publication in this collection
2016
History
-
Received
14 Sept 2015 -
Accepted
11 Apr 2016