ABSTRACT
Pacific and Atlantic-slope rivers of the Isthmus of Tehuantepec in Mexico contain two cichlid species of the genus Vieja relevant to the current study, V. zonata and V. coatlicue, respectively. The Atlantic-slope species was initially considered a population of uncertain taxonomic status and recently described as a distinct species based on three putatively diagnostic morphological characters. The objective of this study was to combine new and existing genetic data, along with reanalysis of morphological characters, to test the distinctiveness of V. coatlicue. Little genetic variability and no geographic structuring were recovered. Additionally, putatively diagnostic morphological character states were present across both species, failing to separate the forms. The synthesis of molecular and morphological data supports the recognition of V. coatlicue as a junior synonym of V. zonata.
Keywords:
Cichlid; Freshwater; Mexico; River; Taxonomy
RESUMEN
Dos especies del género Vieja se encuentran distribuidas a lo largo Istmo de Tehuantepec en México; V. zonata se distribuye en los ríos de la vertiente del Pacífico y V. coatlicue en los ríos de la vertiente del Atlántico. La especie que se distribuye en la vertiente del Atlántico era considerada como una población de estado taxonómico incierto, pero recientemente fue descrita como nueva especie en base a tres caracteres morfológicos diagnósticos. El objetivo de este estudio fue utilizar datos moleculares nuevos con datos moleculares previamente publicados en combinación con una reanálisis de los caracteres morfológicos para testear la validez taxonómica de la especie V. coatlicue. Nuestros análisis moleculares no recobran estructura geográfica y además muestran baja variabilidad genética. Además, los tres caracteres morfológicos diagnósticos se encontraron presentes en individuos de ambas especies, nuestro trabajo sugiere que estos caracteres morfológicos diagnósticos no proven información que permita separar a las formas que se distribuyen en la vertiente del Pacífico y la vertiente del Atlántico. Nuestra síntesis de datos moleculares y morfológicos provee evidencia para reconocer a V. coatlicue como una sinónima mas reciente de V. zonata.
Palabras clave:
Agua dulce; Cíclido; México; Río; Taxonomía
Introduction
The genus Vieja Fernández-Yépez, 1969 is a clade of predominantly Northern Middle American cichlids, with highest diversity and endemism occurring in southern Mexico and Guatemala. One species in the genus, Vieja zonata (Meek, 1905), is endemic to Mexico and found in rivers along the Isthmus of Tehuantepec from the Río Tequisistlán in Oaxaca west of Tehuantepec to Río Tapantepec near the Chiapas border likely near the town of San Pedro Tapantepec (Kullander, 2003Kullander SO. Family Cichlidae (Cichlids). In: Reis RE, Kullander SO, Ferraris CJ Jr., organizers. Check list of the freshwater fishes of South and Central America. Porto Alegre: Edipucrs; 2003. p.605-54.; Miller et al., 2005Miller RR, Minckley WL, Norris SM. Freshwater fishes of México. Chicago (IL): University of Chicago Press; 2005.; Matamoros et al., 2015Matamoros WA, McMahan CD, Chakrabarty P, Albert JS, Schaefer JF. Derivation of the freshwater fish fauna of Central America revisited: Myers’s hypothesis in the twenty-first century. Cladistics. 2015; 31(2):177-88.). In addition, an Atlantic slope population of this species exists in the Río Coatzacoalcos drainage (Conkel, 1993Conkel D. Cichlids of North and Central America. New Jersey: T.F.H. Publications; 1993.; Miller et al., 2005Miller RR, Minckley WL, Norris SM. Freshwater fishes of México. Chicago (IL): University of Chicago Press; 2005.) that Miller et al. (2005Miller RR, Minckley WL, Norris SM. Freshwater fishes of México. Chicago (IL): University of Chicago Press; 2005.) claimed did not represent V. zonata but a “population of undetermined taxonomic status”. Del Moral-Flores et al. (2018del Moral-Flores LF, López-Segovia E, Hernández-Arellano T. Vieja coatlicue sp. nov., una nueva especie de cíclido (Actinopterygii: Cichlidae) de la Cuenca del Río Coatzacoalcos, México. Rev Zool, Universidad Nacional Autónoma de México. 2018; 29:1-12.) recognized the Río Coatzacoalcos population as a distinct species, describing it as V. coatlicue based on three diagnostic morphological characters. Compared to V. zonata, the species was diagnosed based on an elongate body, dorsal-fin origin equal or posterior to opercular border, and a separation between the ends of non-filamentous rays of the pelvic fins and the anus.
In a molecular systematic study of Vieja, McMahan et al. (2010McMahan CD, Geheber AD, Piller KR. Molecular systematics of the enigmatic Middle American genus Vieja (Teleostei: Cichlidae). Mol Phylogenet Evol . 2010; 57(3):1293-300.) recovered low genetic differentiation among individuals of sister species V. zonata and V. guttulata (Günther, 1864) in their phylogeny of the genus. Species-level analyses were beyond the scope of that study; however, Atlantic slope (Río Coatzacoalcos) sequences of V. zonata (currently V. coatlicue) were included in their molecular phylogeny and all populations were recovered in the same clade. Given the lack of genetic distinctiveness of the population in Río Coatzacoalcos, the objective of this study was to synthesize genetic data and reassess the morphological characters provided by del Moral-Flores et al. (2018del Moral-Flores LF, López-Segovia E, Hernández-Arellano T. Vieja coatlicue sp. nov., una nueva especie de cíclido (Actinopterygii: Cichlidae) de la Cuenca del Río Coatzacoalcos, México. Rev Zool, Universidad Nacional Autónoma de México. 2018; 29:1-12.) to test morphological distinctiveness of V. coatlicue in the Río Coatzacoalcos drainage.
Material and Methods
Molecular data. Sequence data for V. coatlicue and several populations of V. zonata have been published in previous studies (e.g. McMahan et al., 2010McMahan CD, Geheber AD, Piller KR. Molecular systematics of the enigmatic Middle American genus Vieja (Teleostei: Cichlidae). Mol Phylogenet Evol . 2010; 57(3):1293-300.; Říčan et al., 2016Říčan O, Piálek L, Dragová K, Novák J. Diversity and evolution of the Middle American cichlid fishes (Teleostei: Cichlidae) with revised classification. Vertebr Zool. 2016; 66(1):1-102.). We sequenced three additional individuals to provide data from other localities for both species (Fig. 1; Tab. 1). Whole genomic DNA was extracted from tissue samples (muscle or fin clip) using the DNeasy kit (Qiagen, Inc.). The mitochondrial cytochrome b (cyt b) gene was amplified and sequenced for samples following protocols described in Perdices et al. (2002Perdices A, Bermingham E, Montilla A, Doadrio I. Evolutionary history of the genus Rhamdia (Teleostei: Pimelodidae) in Central America. Mol Phylogenet Evol . 2002; 25(1):172-89.). Cytochrome b is an appropriate marker to address the main objective of this study as it has been quite useful to provide resolution at intra- and interspecific levels within Middle American cichlids (López-Fernández et al., 2010López-Fernández H, Winemiller KO, Honeycutt RL. Multilocus phylogeny and rapid radiations in Neotropical cichlid fishes (Perciformes: Cichlidae: Cichlinae). Mol Phylogenet Evol. 2010; 55(3):1070-86.; McMahan et al., 2010McMahan CD, Geheber AD, Piller KR. Molecular systematics of the enigmatic Middle American genus Vieja (Teleostei: Cichlidae). Mol Phylogenet Evol . 2010; 57(3):1293-300., 2017aMcMahan CD, Ginger L, Cage M, David KT, Chakrabarty P, Johnston M, Matamoros WA. Pleistocene to Holocene expansion of the black-belt cichlid in Central America, Vieja maculicauda (Teleostei: Cichlidae). PLoS ONE . 2017a; 12(5):e0178439.; Harrison et al., 2014Harrison E, Trexler JC, Collins TM, Vasquez-Domínguez E, Razo-Mendivil U, Matamoros WA, Barrientos C. Genetic evidence for multiple sources of the non-native fish Cichlasoma urophthalmus (Günther; Mayan cichlids) in Southern Florida. PLoS ONE. 2014; 9(9):e104173.; Barrientos-Villalobos et al., 2018Barrientos-Villalobos J, Schmitter-Soto JJ, Espinosa de los Monteros AJ. Several subspecies or phenotypic plasticity? A geometric morphometric and molecular analysis of variability of the Mayan cichlid Mayaheros urophthalmus in the Yucatan. Copeia . 2018; 106(2):268-78.). Additionally, various nuclear markers have failed to delineate taxa at this level (López-Fernández et al., 2010López-Fernández H, Winemiller KO, Honeycutt RL. Multilocus phylogeny and rapid radiations in Neotropical cichlid fishes (Perciformes: Cichlidae: Cichlinae). Mol Phylogenet Evol. 2010; 55(3):1070-86.; McMahan et al., 2010McMahan CD, Geheber AD, Piller KR. Molecular systematics of the enigmatic Middle American genus Vieja (Teleostei: Cichlidae). Mol Phylogenet Evol . 2010; 57(3):1293-300.; Říčan et al., 2016Říčan O, Piálek L, Dragová K, Novák J. Diversity and evolution of the Middle American cichlid fishes (Teleostei: Cichlidae) with revised classification. Vertebr Zool. 2016; 66(1):1-102.). Amplification products were visualized on a 1% agarose gel, followed by purification with ExoSap-IT, and then sequenced on an ABI 3730 in the Pritzker Laboratory for Molecular Systematics and Evolution at the Field Museum of Natural History. An individual of V. guttulata was sequenced as an additional ingroup taxon to test monophyly of V. zonata and V. coatlicue. Vieja fenestrata (Günther, 1860) was used as an outgroup following phylogenetic relationships of McMahan et al. (2010McMahan CD, Geheber AD, Piller KR. Molecular systematics of the enigmatic Middle American genus Vieja (Teleostei: Cichlidae). Mol Phylogenet Evol . 2010; 57(3):1293-300.) and Říčan et al. (2016Říčan O, Piálek L, Dragová K, Novák J. Diversity and evolution of the Middle American cichlid fishes (Teleostei: Cichlidae) with revised classification. Vertebr Zool. 2016; 66(1):1-102.). All new sequences were deposited in GenBank (Tab. 1).
Map of Mexico displaying localities for specimens (triangles) examined and tissue samples (circles) for Vieja coatlicue (green) and Vieja zonata (black) used in this study. Star indicates locality for Vieja guttulata sample.
Specimens/tissues examined. No. refers to number by tip in phylogeny (Fig. 2). Acronyms follow Sabaj (2016Sabaj MH, editor. Standard symbolic codes for institutional resource collections in herpetology and ichthyology: an online reference. Version 6.5 [Internet]. American Society of Ichthyologists and Herpetologists; 2018 [updated 16 Aug 2016]. Available from: https://asih.org/standard-symbolic-codes
https://asih.org/standard-symbolic-codes... ).
Sequence data were visually inspected and aligned using the software Geneious v.9.1 (Kearse et al., 2012Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T, Ashton B, Meintjes P, Drummond A. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics. 2012; 28(12):1647-49.). A multiple sequence alignment was generated using the Muscle algorithm (Edgar, 2004Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004; 32(5):1792-97.) implemented in Geneious using default parameters. The final alignment consisted of 1,050 base pairs. Partition scheme and model selection were performed using the software PartionFinder (Lanfear et al., 2012Lanfear R, Calcott B, Ho SYW, Guindon S. PartitionFinder: combined selection of partitioning schemes and substitution models for phylogenetic analyses. Mol Biol Evol . 2012; 29(6):1695-701.) using a greedy search. The best partition scheme and substitution model were selected using the Akaike Information Criterion (AIC).
Phylogenetic hypotheses were inferred under a Bayesian framework using the software Mr. Bayes v.3.2.6 (Hulsenbeck et al., 2001Hulsenbeck JP, Ronquist F, Nielsen R, Bollback JP. Bayesian inference of phylogeny and its impact on evolutionary biology. Science. 2001; 294(5550):2310-14.) implemented in the CIPRES portal (Miller et al., 2010Miller MA, Pfeiffer W, Schwartz T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. In: Proceedings of the Gateway Computing Environments workshop (GCE), 2010; New Orleans, LA. 2010. p. 1-8.). Three independent analyses of 12,000,000 generations of two replicated runs each were executed, with tree sampling every 6,000 generations. Sampling stationarity was assessed via visual inspection in Tracer v.1.5 (Rambaut et al., 2018Rambaut A, Drummond AJ, Xie D, Baele G, Suchard MA. Posterior summarization in Bayesian phylogenetics using Tracer 1.7. Syst Biol. 2018; 67(5):901-04.) and observed average standard deviations of split frequencies (<0.01). The initial 25% of sampled trees were discarded as burn-in. Bayesian posterior probabilities were calculated using the post burn-in trees, and the three independent runs were compared to evaluate topological congruence.
Due to missing sequence data in one sample (KU854740), estimation of genetic diversity and reconstruction of a haplotype network relied on a slightly reduced dataset (1,011 base pairs). Nucleotide (π; Nei, Li, 1979Nei M, Li WH. Mathematical model for studying genetic variation in terms of restriction endonucleases. PNAS. 1979; 76(10):5269-73.), genetic (θw; Watterson, 1975Watterson GA. On the number of segregating sites in genetical models without recombination. Theor Popul Biol. 1975; 7(2):256-76.), and haplotype (Hd) diversity indices were calculated in the software DnaSP v.5.10 (Librado, Rozas, 2009Librado P, Rozas J. DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics . 2009; 25(11):1451-52.). Matrilineal haplotypes were reconstructed using statistical parsimony analysis in the software TCS (Templeton et al., 1992Templeton AR, Crandall KA, Sing CF. A cladistics analysis of phenotypic association with haplotypes inferred from restriction endonuclease mapping and DNA sequence data. III. Cladogram estimation. Genetics. 1992; 132(2):619-33.) with a 95% confidence limit for connected haplotypes. The recovered network with TCS was used to infer gene genealogies using a median-joining network (Bandelt et al., 1999Bandelt HJ, Foster P, Röhl A. Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol. 1999; 16(1):37-48.) implemented in the software POPART (Leigh, Bryant, 2015Leigh JW, Bryant D. PopART: Full-feature software for haplotype network construction. Methods Ecol Evol. 2015; 6(9):1110-16.).
Morphological data. Material examined covered four localities for V. zonata (N= 29) along Pacific drainages in the Isthmus of Tehuantepec, including the holotype and paratype, and three populations of V. coatlicue (N= 22) in the Río Coatzacoalcos (Tab. 1; Fig. 1). Our goal was to reassess the putative morphological distinctiveness of V. coatlicue relative to V. zonata; thus, characters examined herein focused on the three described by del Moral-Flores et al. (2018del Moral-Flores LF, López-Segovia E, Hernández-Arellano T. Vieja coatlicue sp. nov., una nueva especie de cíclido (Actinopterygii: Cichlidae) de la Cuenca del Río Coatzacoalcos, México. Rev Zool, Universidad Nacional Autónoma de México. 2018; 29:1-12.) to diagnose V. coatlicue as a distinct species from V. zonata. Standard length (SL) and maximum body depth (BD) were measured for each specimen. The ratio of SL:BD, also known as the fineness ratio, was calculated and used as a measure of body elongation, with greater values indicating more elongate bodies (Aguirre et al., 2016Aguirre WE, Navarrete R, Malato G, Calle P, Loh MK, Vital WF, Valadez G, Vu V, Shervette VR, Granda JC. Body shape variation and population genetic structure of Rhoadsia altipinna (Characidae: Rhoadsiinae) in Southwestern Ecuador. Copeia. 2016; 104(2):554-69.). Additionally, data on anterior insertion of the dorsal fin relative to posterior of the operculum (anterior/posterior to or in-line with) and presence/absence of a space between non-filamentous portions of the pelvic rays and the anus were recorded. These data were assessed to determine if the proposed diagnostic characters could separate the two species and if morphological differences were congruent with any observed patterns of genetic differentiation.
Results
Molecular data. A total of 10 individuals of V. coatlicue (four localities) and seven individuals of V. zonata (five localities) were analyzed based on new and previously published sequences. The best partition scheme was by codon with independent substitution models (1st codon: HKY, 2nd codon: F81, 3rd codon: GTR+gamma). A single clade containing V. zonata and V. coatlicue was recovered based on phylogenetic analysis of cyt b data, with an average 0.2% sequence divergence observed across all individuals of both species. We recovered no geographic structuring within this clade (Fig. 2). Vieja guttulata was recovered as sister to the clade containing V. zonata and V. coatlicue, with an average sequence divergence of 2.2%. This result is consistent with McMahan et al. (2015McMahan CD, Matamoros WA, Piller KR, Chakrabarty P. Taxonomy and systematics of the herichthyins (Cichlidae: Tribe Heroini), with the description of eight new Middle American Genera. Zootaxa . 2015; 3999(2):211-34.) and Říčan et al. (2016Říčan O, Piálek L, Dragová K, Novák J. Diversity and evolution of the Middle American cichlid fishes (Teleostei: Cichlidae) with revised classification. Vertebr Zool. 2016; 66(1):1-102.). Sequence divergence between the outgroup (V. fenestrata) and ingroup was 2.7%.
Bayesian phylogeny based on cytochrome b sequence data. Number above clades is posterior probability.
We recovered four haplotypes in the network generated by statistical parsimony (Fig. 3). One haplotype was common and widely distributed across the Coatzacoalcos and Tehuantepec river basins and shared between individuals of V. coatlicue and V. zonata (Fig. 3). Finally, all samples from the clade containing V. coatlicue and V. zonata showed low levels of nucleotide (π=0.493, standard deviation 0.131), genetic (θw =0.0016, standard deviation 0.0078), and haplotype (Hd=0.0031, standard deviation 0.00139) diversity across populations and river basins.
Haplotype network recovered from analysis of the cyt b dataset for populations of Vieja coatlicue (Atlantic) and Vieja zonata (Pacific). Dashes lines represent mutational steps.
Morphological data. The average fineness ratio for V. coatlicue was 2.11 (N=22, standard deviation 0.11, minimum 1.87, maximum 2.31), compared to 2.28 (N=29, standard deviation 0.1, minimum 2.04, maximum 2.45) for V. zonata (Tab. 2). This reveals a slightly more streamlined or elongate overall body shape in V. zonata, inconsistent with the purported elongate body diagnostic of V. coatlicue. Across both species we found that most individuals possessed a dorsal-fin origin marginally anterior to- or in-line with the posterior margin of the operculum, although a moderately higher number of individuals of V. zonata exhibited a dorsal-fin origin slightly posterior to the operculum (Tab. 3; Fig. 4). Dorsal-fin origin was only vaguely anterior to the posterior end of the operculum in individuals with this character state and could easily be considered in-line with posterior of the operculum. Additionally, we observed these character states to be skewed in bent specimens, depending on if gathering data from the right or left sides. Absence of a space between non-filamentous rays of the pelvic fin and the anus was observed across the majority of individuals of both V. coatlicue and V. zonata (Tab. 3; Fig. 5).
Dorsal-fin origin relative to operculum in Vieja coatlicue (top; UMMZ 178548) and Vieja zonata (bottom; FMNH 3776 (holotype), UMMZ 184746), illustrating variability within and between species. Scale bar equals 1cm.
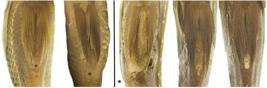
Pelvic fin relative to anus in Vieja coatlicue (left of vertical line; UMMZ 178548) and Vieja zonata (right of vertical line; FMNH 3776 (holotype), UMMZ 184746), illustrating variability within and between species. Asterisk indicates holotype of V. zonata.
Number of individuals of Vieja coatlicue and Vieja zonata with examined character states dorsal-fin origin relative to operculum and presence/absence of space between non-filamentous ends of the pelvic fins and anus.
Discussion
We found a lack of monophyly for V. zonata and V. coatlicue, as well as absence of any reasonable level of genetic divergence across species or populations. Additionally, reanalysis of putatively diagnostic morphological characters fails to unambiguously separate the two species, with all character states being observed across the range of the two species. Fineness ratios show the opposite trend from description of V. coatlicue, with slightly more elongate bodies in populations of V. zonata. The origin of the dorsal fin relative to posterior of the operculum exhibits variability between the two species; however, in nearly all specimens the origin is very close to in-line with the posterior edge of the operculum. Few individuals, across both species, exhibit a clearly anterior or posterior condition. We found that specimens of either species rarely possess a clearly defined space or gap between tips of the (non-filamentous) pelvic fin and the anus. High levels of variability were consistently observed for this character across all material examined. Indeed, the holotype of V. zonata possesses the most evident space between the non-filamentous pelvic rays and the anus (Fig. 5). The variability in these morphological characters is high and consistent across populations of both species. Thus, the combination of morphological and genetic data supports the recognition of V. coatlicue as a junior synonym of V. zonata.
Many studies have demonstrated high levels of intraspecific morphological variability within cichlid fishes (McMahan et al., 2017bMcMahan CD, Kutz J, Murray CM, Chakrabarty P, Geheber AD, Elias D. Objectively measuring subjectively described traits: geographic variation in body shape and caudal coloration pattern within Vieja melanura (Teleostei: Cichlidae). Rev Biol Trop. 2017b; 65(2):623-31.; Barrientos-Villalobos et al., 2018Barrientos-Villalobos J, Schmitter-Soto JJ, Espinosa de los Monteros AJ. Several subspecies or phenotypic plasticity? A geometric morphometric and molecular analysis of variability of the Mayan cichlid Mayaheros urophthalmus in the Yucatan. Copeia . 2018; 106(2):268-78.; Gómez-González et al., 2018Gómez-González AE, Alvarez F, Matamoros WA, Velázquez-Velázquez E, Schmitter-Soto JJ, Gonzalez-Diaz AA, McMahan CD. Redescription of Vieja hartwegi (Taylor & Miller 1980) (Teleostei: Cichlidae) from the Grijalva River basin, Mexico and Guatemala, with description of a rheophilic morph. Zootaxa. 2018; 4375(3):371-91.). While genetic data are certainly not a required component of detection and description of new species, these independent sources of data can offer additional ways to test hypotheses regarding species delineation. While McMahan et al. (2010McMahan CD, Geheber AD, Piller KR. Molecular systematics of the enigmatic Middle American genus Vieja (Teleostei: Cichlidae). Mol Phylogenet Evol . 2010; 57(3):1293-300.) provided sequence data from populations of V. zonata from the Río Tehuantepec and Río Coatzacoalcos, admittedly the authors did not discuss this in text given it was outside the scope of that study. However, locality data and sequences for these specimens were published and deposited in GenBank. Říčan et al. (2016Říčan O, Piálek L, Dragová K, Novák J. Diversity and evolution of the Middle American cichlid fishes (Teleostei: Cichlidae) with revised classification. Vertebr Zool. 2016; 66(1):1-102.) later found the same lack of genetic distinctiveness between V. zonata from both slopes of the Isthmus of Tehuantepec. Anzueto-Calvo et al. (2016Anzueto-Calvo MJ, Velázquez-Velázquez E, Gómez-González AE. Peces de la Reserva de la Biosfera Selva El Ocote y presa Nezahualcóyotl (Malpaso) Chiapas, México. Rev Mex Biodivers. 2016; 87(3):972-79.) additionally report the occurrence of V. zonata in the Río Grijalva, which should be studied in future analyses of this and related species.
The Isthmus of Tehuantepec represents the narrowest geographic point between the Pacific and Atlantic basins in the northern portion of Middle America and has been demonstrated to be an important region for organismal dispersal across Mexico, permitting taxa to expand between rivers along both slopes (Quiroz-Martínez et al., 2014Quiroz-Martínez B, Álvarez F, Espinosa H, Salgado-Maldonado G. Concordant biogeographic patterns among multiple taxonomic groups in the Mexican freshwater biota. PLoS ONE . 2014; 9(8):e105510.). This hypothesis is additionally supported by occurrence of a population of V. zonata in the Atlantic Río Coatzacoalcos. Thus, V. zonata in the Atlantic represent a recently dispersed population and not a distinct lineage.
Acknowledgments
We are thankful to Susan Mochel (FMNH) and Kevin Swagel (FMNH) for invaluable help with logistics and specimen loans. Doug Nelson (UMMZ) and Hernán López-Fernández (UMMZ) kindly provided loans of specimens supporting this work. Many thanks to Adán Gómez González for valuable discussions and fieldwork assistance. This work was supported, in part, with a grant from the Negaunee Foundation to CDM. Molecular work was carried out in the Pritzker Laboratory for Molecular Systematics and Evolution at the Field Museum of Natural History, with support from the Pritzker Foundation.
References
- Aguirre WE, Navarrete R, Malato G, Calle P, Loh MK, Vital WF, Valadez G, Vu V, Shervette VR, Granda JC. Body shape variation and population genetic structure of Rhoadsia altipinna (Characidae: Rhoadsiinae) in Southwestern Ecuador. Copeia. 2016; 104(2):554-69.
- Anzueto-Calvo MJ, Velázquez-Velázquez E, Gómez-González AE. Peces de la Reserva de la Biosfera Selva El Ocote y presa Nezahualcóyotl (Malpaso) Chiapas, México. Rev Mex Biodivers. 2016; 87(3):972-79.
- Bandelt HJ, Foster P, Röhl A. Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol. 1999; 16(1):37-48.
- Barrientos-Villalobos J, Schmitter-Soto JJ, Espinosa de los Monteros AJ. Several subspecies or phenotypic plasticity? A geometric morphometric and molecular analysis of variability of the Mayan cichlid Mayaheros urophthalmus in the Yucatan. Copeia . 2018; 106(2):268-78.
- Conkel D. Cichlids of North and Central America. New Jersey: T.F.H. Publications; 1993.
- Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004; 32(5):1792-97.
- Gómez-González AE, Alvarez F, Matamoros WA, Velázquez-Velázquez E, Schmitter-Soto JJ, Gonzalez-Diaz AA, McMahan CD. Redescription of Vieja hartwegi (Taylor & Miller 1980) (Teleostei: Cichlidae) from the Grijalva River basin, Mexico and Guatemala, with description of a rheophilic morph. Zootaxa. 2018; 4375(3):371-91.
- Harrison E, Trexler JC, Collins TM, Vasquez-Domínguez E, Razo-Mendivil U, Matamoros WA, Barrientos C. Genetic evidence for multiple sources of the non-native fish Cichlasoma urophthalmus (Günther; Mayan cichlids) in Southern Florida. PLoS ONE. 2014; 9(9):e104173.
- Hulsenbeck JP, Ronquist F, Nielsen R, Bollback JP. Bayesian inference of phylogeny and its impact on evolutionary biology. Science. 2001; 294(5550):2310-14.
- Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T, Ashton B, Meintjes P, Drummond A. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics. 2012; 28(12):1647-49.
- Kullander SO. Family Cichlidae (Cichlids). In: Reis RE, Kullander SO, Ferraris CJ Jr., organizers. Check list of the freshwater fishes of South and Central America. Porto Alegre: Edipucrs; 2003. p.605-54.
- Lanfear R, Calcott B, Ho SYW, Guindon S. PartitionFinder: combined selection of partitioning schemes and substitution models for phylogenetic analyses. Mol Biol Evol . 2012; 29(6):1695-701.
- Leigh JW, Bryant D. PopART: Full-feature software for haplotype network construction. Methods Ecol Evol. 2015; 6(9):1110-16.
- Librado P, Rozas J. DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics . 2009; 25(11):1451-52.
- López-Fernández H, Winemiller KO, Honeycutt RL. Multilocus phylogeny and rapid radiations in Neotropical cichlid fishes (Perciformes: Cichlidae: Cichlinae). Mol Phylogenet Evol. 2010; 55(3):1070-86.
- Matamoros WA, McMahan CD, Chakrabarty P, Albert JS, Schaefer JF. Derivation of the freshwater fish fauna of Central America revisited: Myers’s hypothesis in the twenty-first century. Cladistics. 2015; 31(2):177-88.
- McMahan CD, Geheber AD, Piller KR. Molecular systematics of the enigmatic Middle American genus Vieja (Teleostei: Cichlidae). Mol Phylogenet Evol . 2010; 57(3):1293-300.
- McMahan CD, Matamoros WA, Piller KR, Chakrabarty P. Taxonomy and systematics of the herichthyins (Cichlidae: Tribe Heroini), with the description of eight new Middle American Genera. Zootaxa . 2015; 3999(2):211-34.
- McMahan CD, Ginger L, Cage M, David KT, Chakrabarty P, Johnston M, Matamoros WA. Pleistocene to Holocene expansion of the black-belt cichlid in Central America, Vieja maculicauda (Teleostei: Cichlidae). PLoS ONE . 2017a; 12(5):e0178439.
- McMahan CD, Kutz J, Murray CM, Chakrabarty P, Geheber AD, Elias D. Objectively measuring subjectively described traits: geographic variation in body shape and caudal coloration pattern within Vieja melanura (Teleostei: Cichlidae). Rev Biol Trop. 2017b; 65(2):623-31.
- Miller RR, Minckley WL, Norris SM. Freshwater fishes of México. Chicago (IL): University of Chicago Press; 2005.
- Miller MA, Pfeiffer W, Schwartz T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. In: Proceedings of the Gateway Computing Environments workshop (GCE), 2010; New Orleans, LA. 2010. p. 1-8.
- del Moral-Flores LF, López-Segovia E, Hernández-Arellano T. Vieja coatlicue sp. nov., una nueva especie de cíclido (Actinopterygii: Cichlidae) de la Cuenca del Río Coatzacoalcos, México. Rev Zool, Universidad Nacional Autónoma de México. 2018; 29:1-12.
- Nei M, Li WH. Mathematical model for studying genetic variation in terms of restriction endonucleases. PNAS. 1979; 76(10):5269-73.
- Perdices A, Bermingham E, Montilla A, Doadrio I. Evolutionary history of the genus Rhamdia (Teleostei: Pimelodidae) in Central America. Mol Phylogenet Evol . 2002; 25(1):172-89.
- Quiroz-Martínez B, Álvarez F, Espinosa H, Salgado-Maldonado G. Concordant biogeographic patterns among multiple taxonomic groups in the Mexican freshwater biota. PLoS ONE . 2014; 9(8):e105510.
- Rambaut A, Drummond AJ, Xie D, Baele G, Suchard MA. Posterior summarization in Bayesian phylogenetics using Tracer 1.7. Syst Biol. 2018; 67(5):901-04.
- Říčan O, Piálek L, Dragová K, Novák J. Diversity and evolution of the Middle American cichlid fishes (Teleostei: Cichlidae) with revised classification. Vertebr Zool. 2016; 66(1):1-102.
- Sabaj MH, editor. Standard symbolic codes for institutional resource collections in herpetology and ichthyology: an online reference. Version 6.5 [Internet]. American Society of Ichthyologists and Herpetologists; 2018 [updated 16 Aug 2016]. Available from: https://asih.org/standard-symbolic-codes
» https://asih.org/standard-symbolic-codes - Templeton AR, Crandall KA, Sing CF. A cladistics analysis of phenotypic association with haplotypes inferred from restriction endonuclease mapping and DNA sequence data. III. Cladogram estimation. Genetics. 1992; 132(2):619-33.
- Watterson GA. On the number of segregating sites in genetical models without recombination. Theor Popul Biol. 1975; 7(2):256-76.
Edited by
Data availability
Data citations
Sabaj MH, editor. Standard symbolic codes for institutional resource collections in herpetology and ichthyology: an online reference. Version 6.5 [Internet]. American Society of Ichthyologists and Herpetologists; 2018 [updated 16 Aug 2016]. Available from: https://asih.org/standard-symbolic-codes
Publication Dates
-
Publication in this collection
19 June 2019 -
Date of issue
2019
History
-
Received
22 Jan 2019 -
Accepted
15 Apr 2019