Abstract
Introduction
Streptococcus mutans (S. mutans) exhibits extensive genotypic diversity, but the role of this variation is poorly understood.
Objective
To evaluate the genotypic diversity of S. mutans in preschoolers with and without early dental care and to associate it with dental caries experience.
Material and method
Twenty children, aged five year old, with history of dental care since they were one year old (G1) and 20 children without history of dental care (G2) were included. Their oral health status was assessed by the dmft index. S. mutans samples were isolated from the saliva and analysed by polymerase chain reaction. A total of 339 S. mutans isolates (173 from G1 and 166 from G2) were genotyped by random amplified polymorphic DNA with arbitrary primers OPA-02 and OPA-13.
Result
The results revealed 75 distinct genotypes of S. mutans in G1 and 73 in G2. Furthermore, G1 and G2 subjects harboured one to eight and one to seven distinct genotypes of S. mutans, respectively. A statistically significant association (P=0.021) and a moderate correlation (r= 0.503) were observed between dental caries experience and genotypic diversity of S. mutans in G1.
Conclusion
From the limitations of the study design, we just can hypothesize that actions of early dental care carried out by an educative-preventive program can contribute to a distinct oral microbiota.
Descriptors:
Genotypic diversity; Streptococcus mutans; dental care; dental caries
Resumo
Introdução
Streptococcus mutans (S. mutans) apresentam ampla diversidade genotípica, mas o papel desta variação ainda é pouco compreendido.
Objetivo
Avaliar a diversidade genotípica de S. mutans em pré-escolares com e sem assistência odontológica precoce e associá-la à experiência de cárie dentária.
Material e método
Participaram da pesquisa 20 crianças, de cinco anos de idade, com história de assistência odontológica desde o primeiro ano de vida (G1) e 20 crianças sem história de assistência odontológica precoce (G2). A experiência de cárie dentária foi avaliada por meio do índice ceo-d. Amostras de S. mutans foram isoladas a partir da saliva dos participantes e analisadas pela Reação em Cadeia da Polimerase para identificação molecular da espécie. Um total de 339 isolados bacterianos (173 do G1 e 166 do G2) foram genotipados através da Amplificação ao Acaso de DNA Polimórfico, utilizando-se os primers OPA-02 e OPA-13.
Resultado
Os resultados revelaram 75 diferentes genótipos de S. mutans no G1 e 73 no G2. Além disso, as crianças do G1 e G2 abrigaram em sua cavidade bucal de um a oito e de um a sete genótipos distintos de S. mutans, respectivamente. Foram observadas uma associação estatisticamente significante (P=0.021) e uma moderada correlação (r=0,503) entre a diversidade genotípica de S. mutans e a experiência de cárie somente no G1.
Conclusão
A partir das limitações do desenho do estudo, os resultados apenas sugerem que as ações realizadas por meio de um atendimento odontológico precoce podem estar contribuindo para o estabelecimento de uma microbiota bucal distinta.
Descritores:
Diversidade genotípica; Streptococcus mutans; assistência odontológica; cárie dentária
INTRODUCTION
S. mutans exhibits extensive genotypic diversity, but the role of this variation is poorly understood11 Lembo FL, Longo PL, Ota-Tsuzuki C, Rodrigues CR, Mayer MP. Genotypic and phenotypic analysis of Streptococcus mutans from different oral cavity sites of caries-free and caries-active children. Oral Microbiol Immunol. 2007 Oct;22(5):313-9. http://dx.doi.org/10.1111/j.1399-302X.2007.00361.x. PMid:17803628.
http://dx.doi.org/10.1111/j.1399-302X.20...
because different strains may display different virulence mechanisms. In addition, the ability of specific genotypes of S. mutans to compete with other strains may be essential for their colonization22 Longo PL, Mattos-Graner RO, Mayer MP. Determination of mutacin activity and detection of mutA genes in Streptococcus mutans genotypes from caries-free and caries-active children. Oral Microbiol Immunol. 2003 Jun;18(3):144-9. http://dx.doi.org/10.1034/j.1399-302X.2003.00042.x. PMid:12753464.
http://dx.doi.org/10.1034/j.1399-302X.20...
and consequently, some genotypes may be able to colonize the host and induce dental caries better than others11 Lembo FL, Longo PL, Ota-Tsuzuki C, Rodrigues CR, Mayer MP. Genotypic and phenotypic analysis of Streptococcus mutans from different oral cavity sites of caries-free and caries-active children. Oral Microbiol Immunol. 2007 Oct;22(5):313-9. http://dx.doi.org/10.1111/j.1399-302X.2007.00361.x. PMid:17803628.
http://dx.doi.org/10.1111/j.1399-302X.20...
,33 Alaluusua S, Mättö J, Grönroos L, Innilä S, Torkko H, Asikainen S, et al. Oral colonization by more than one clonal type of mutans streptococcus in children with nursing-bottle dental caries. Arch Oral Biol. 1996 Feb;41(2):167-73. http://dx.doi.org/10.1016/0003-9969(95)00111-5. PMid:8712973.
http://dx.doi.org/10.1016/0003-9969(95)0...
.
The investigation of S. mutans genotypic characteristics and the use of molecular biology techniques have allowed for significant development in the understanding of the process of dental caries44 Liu Y, Zou J, Shang R, Zhou XD. Genotypic diversity of Streptococcus mutans in 3- to 4-year-old Chinese nursery children suggests horizontal transmission. Arch Oral Biol. 2007 Sep;52(9):876-81. http://dx.doi.org/10.1016/j.archoralbio.2007.03.004. PMid:17466259.
http://dx.doi.org/10.1016/j.archoralbio....
and the improvement of caries risk assessments and treatment options55 Momeni SS, Whiddon J, Moser SA, Childers NK. Transmission patterns of Streptococcus mutans demonstrated by a combined rep-PCR and MLST approach. Clin Oral Investig. 2018 Nov;22(8):2847-58. http://dx.doi.org/10.1007/s00784-018-2371-8. PMid:29476335.
http://dx.doi.org/10.1007/s00784-018-237...
. Several studies have shown that individuals can be colonized with more than one genetic type of S. mutans, through the use of different molecular typing methods44 Liu Y, Zou J, Shang R, Zhou XD. Genotypic diversity of Streptococcus mutans in 3- to 4-year-old Chinese nursery children suggests horizontal transmission. Arch Oral Biol. 2007 Sep;52(9):876-81. http://dx.doi.org/10.1016/j.archoralbio.2007.03.004. PMid:17466259.
http://dx.doi.org/10.1016/j.archoralbio....
,66 Emanuelsson I-MR, Wang X. Demonstration of identical strains of mutans streptococci within Chinese families by genotyping. Eur J Oral Sci. 1998 Jun;106(3):788-94. http://dx.doi.org/10.1046/j.0909-8836.1998.eos106305.x. PMid:9672101.
http://dx.doi.org/10.1046/j.0909-8836.19...
7 Napimoga MH, Kamiya RU, Rosa RT, Rosa EA, Höfling JF, Mattos-Graner R, et al. Genotypic diversity and virulence traits of Streptococcus mutans in caries-free and caries-active individuals. J Med Microbiol. 2004 Jul;53(Pt 7):697-703. http://dx.doi.org/10.1099/jmm.0.05512-0. PMid:15184543.
http://dx.doi.org/10.1099/jmm.0.05512-0...
8 Pieralisi FJ, Rodrigues MR, Segura VG, Maciel SM, Ferreira FB, Garcia JE, et al. Genotypic diversity of Streptococcus mutans in caries-free and caries-active preschool children. Int J Dent. 2010;2010:824976. http://dx.doi.org/10.1155/2010/824976. PMid:20351760.
http://dx.doi.org/10.1155/2010/824976...
-99 Zhou Q, Qin X, Qin M, Ge L. Genotypic diversity of Streptococcus mutans and Streptococcus sobrinus in 3-4-year-old children with severe caries or without caries. Int J Paediatr Dent. 2011 Nov;21(6):422-31. http://dx.doi.org/10.1111/j.1365-263X.2011.01145.x. PMid:21689176.
http://dx.doi.org/10.1111/j.1365-263X.20...
. Among those, random amplified polymorphic DNA (RAPD) technique have been considered a sufficiently sensible method capable of detecting polymorphisms among distinct isolates of S. mutans1010 Saarela M, Hannula J, Mättö J, Asikainen S, Alaluusua S. Typing of mutans streptococci by arbitrarily primed polymerase chain reaction. Arch Oral Biol. 1996 Aug-Sep;41(8-9):821-6. http://dx.doi.org/10.1016/S0003-9969(96)00049-0. PMid:9022919.
http://dx.doi.org/10.1016/S0003-9969(96)...
11 Li Y, Caufield PW. Arbitrarily primed polymerase chain reaction fingerprinting for the genotypic identification of mutans streptococci from humans. Oral Microbiol Immunol. 1998 Feb;13(1):17-22. http://dx.doi.org/10.1111/j.1399-302X.1998.tb00745.x. PMid:9573817.
http://dx.doi.org/10.1111/j.1399-302X.19...
12 Grönroos L, Alaluusua S. Site-specific oral colonization of mutans streptococci detected by arbitrarily primed PCR fingerprinting. Caries Res. 2000 Nov-Dec;34(6):474-80. http://dx.doi.org/10.1159/000016626. PMid:11093021.
http://dx.doi.org/10.1159/000016626...
13 Redmo Emanuelsson IM, Carlsson P, Hamberg K, Bratthall D. Tracing genotypes of mutans streptococci on tooth sites by random amplified polymorphic DNA (RAPD) analysis. Oral Microbiol Immunol. 2003 Feb;18(1):24-9. http://dx.doi.org/10.1034/j.1399-302X.2002.180104.x. PMid:12588455.
http://dx.doi.org/10.1034/j.1399-302X.20...
14 Hames-Kocabas EE, Uçar F, Kocatas Ersin N, Uzel A, Alpöz AR. Colonization and vertical transmission of Streptococcus mutans in Turkish children. Microbiol Res. 2008;163(2):168-72. http://dx.doi.org/10.1016/j.micres.2006.03.016. PMid:16735109.
http://dx.doi.org/10.1016/j.micres.2006....
-1515 Tabchoury CP, Sousa MC, Arthur RA, Mattos-Graner RO, Del Bel Cury AA, Cury JA. Evaluation of genotypic diversity of streptococcus mutans using distinct arbitrary primers. J Appl Oral Sci. 2008 Nov-Dec;16(6):403-7. http://dx.doi.org/10.1590/S1678-77572008000600009. PMid:19082399.
http://dx.doi.org/10.1590/S1678-77572008...
.
In Brazil, since 1984, were created Preventive Programs for Babies with the development of clinical evaluations, preventive maintenance procedures and educational/preventive actions to promote children oral health and prevention of dental caries. However, two questions remains as “why some children still experience dental caries, even with a history of dental care since they were one year old?” and “the caries experience may be associated with oral colonization by more than one S. mutans genotype?” In this context, the literature point that relationship between dental caries and the genotypic diversity of S. mutans is still controversial22 Longo PL, Mattos-Graner RO, Mayer MP. Determination of mutacin activity and detection of mutA genes in Streptococcus mutans genotypes from caries-free and caries-active children. Oral Microbiol Immunol. 2003 Jun;18(3):144-9. http://dx.doi.org/10.1034/j.1399-302X.2003.00042.x. PMid:12753464.
http://dx.doi.org/10.1034/j.1399-302X.20...
3 Alaluusua S, Mättö J, Grönroos L, Innilä S, Torkko H, Asikainen S, et al. Oral colonization by more than one clonal type of mutans streptococcus in children with nursing-bottle dental caries. Arch Oral Biol. 1996 Feb;41(2):167-73. http://dx.doi.org/10.1016/0003-9969(95)00111-5. PMid:8712973.
http://dx.doi.org/10.1016/0003-9969(95)0...
-44 Liu Y, Zou J, Shang R, Zhou XD. Genotypic diversity of Streptococcus mutans in 3- to 4-year-old Chinese nursery children suggests horizontal transmission. Arch Oral Biol. 2007 Sep;52(9):876-81. http://dx.doi.org/10.1016/j.archoralbio.2007.03.004. PMid:17466259.
http://dx.doi.org/10.1016/j.archoralbio....
,77 Napimoga MH, Kamiya RU, Rosa RT, Rosa EA, Höfling JF, Mattos-Graner R, et al. Genotypic diversity and virulence traits of Streptococcus mutans in caries-free and caries-active individuals. J Med Microbiol. 2004 Jul;53(Pt 7):697-703. http://dx.doi.org/10.1099/jmm.0.05512-0. PMid:15184543.
http://dx.doi.org/10.1099/jmm.0.05512-0...
,88 Pieralisi FJ, Rodrigues MR, Segura VG, Maciel SM, Ferreira FB, Garcia JE, et al. Genotypic diversity of Streptococcus mutans in caries-free and caries-active preschool children. Int J Dent. 2010;2010:824976. http://dx.doi.org/10.1155/2010/824976. PMid:20351760.
http://dx.doi.org/10.1155/2010/824976...
,1616 Bowden GH. Does assessment of microbial composition of plaque/saliva allow for diagnosis of disease activity of individuals? Community Dent Oral Epidemiol. 1997 Feb;25(1):76-81. http://dx.doi.org/10.1111/j.1600-0528.1997.tb00902.x. PMid:9088695.
http://dx.doi.org/10.1111/j.1600-0528.19...
,1717 Mattos-Graner RO, Li Y, Caufield PW, Duncan M, Smith DJ. Genotypic diversity of mutans streptococci in Brazilian nursery children suggests horizontal transmission. J Clin Microbiol. 2001 Jun;39(6):2313-6. http://dx.doi.org/10.1128/JCM.39.6.2313-2316.2001.
http://dx.doi.org/10.1128/JCM.39.6.2313-...
Therefore, the purpose of this study was to evaluate the genotypic diversity of S. mutans in preschoolers with and without early dental care and to associate it with dental caries experience.
MATERIAL AND METHOD
The protocol for this study was approved by the Research Ethics Committee of the Faculty of Health Science of the University of North Paraná, Londrina, Paraná, Brazil (Protocol 281/07) and written informed consent was obtained from all the parents or legal guardians of minors.
The study population consisted of two groups. Group 1 (G1) comprised 20 randomly selected preschoolers, aged five year old of both genders, and who participated in an educative-preventive program since they were one year old (Oral Health Program for Babies of the University of Londrina, Londrina, Paraná, Brazil) In that program, the assistance involves an initial educational action to encourage families to self-care and routine dental visits for professional cleaning with pumice cup and topical application of 2% sodium fluoride gel, according to assessment on risk of dental caries development. Exclusion criteria were presence of systemic diseases or syndromes and consumption of any form of medication in the last month that might modify the oral microbiota before starting the study. Group 2 (G2) comprised 20 preschoolers without early dental care selected from children attending a nursery class. After eligibility criteria, age and gender were paired among the groups.
Dental caries experience in both groups was assessed using the World Health Organization criteria1818 World Health Organization – WHO. Oral health surveys, basic methods. 4th ed. Geneva: WHO; 1997. to determine the dmft index (decayed/missing/filled deciduous teeth), under artificial light with the aid of a dental mirror and explorer, when necessary, to eliminate debris. Only one examiner that had been previously trained and calibrated (K=0.97) conducted dental examinations. The children with dmft=0 were defined as caries-free and with dmft ≥1 as caries-active, considering at minimum one decayed teeth (white spots and/or cavities)
Microbial samples of saliva were collected using a sterile wooden spatula from subjects. Each side of the spatula was pressed directly onto mitis salivarius bacitracin sucrose agar plates, a medium selective for mutans streptococci. The plates were incubated in 5% CO2 at 37°C for 48 hours. After growth, ten representative morphological types of S. mutans colonies per plate were selected for subjects. Each single colony was transferred aseptically to a microtube with brain heart infusion broth (DIFCO Laboratories, Detroit, USA) and incubated in 5% CO2 at 37°C for 24 hours.
Chromosomal DNA from 400 S. mutans colonies isolates was extracted, as previously described by Saarela et al.1010 Saarela M, Hannula J, Mättö J, Asikainen S, Alaluusua S. Typing of mutans streptococci by arbitrarily primed polymerase chain reaction. Arch Oral Biol. 1996 Aug-Sep;41(8-9):821-6. http://dx.doi.org/10.1016/S0003-9969(96)00049-0. PMid:9022919.
http://dx.doi.org/10.1016/S0003-9969(96)...
, with some modifications. For species confirmation, the isolates were analysed by polymerase chain reaction (PCR) with specific primers for gtfB, encoding glucosyltransferase B (5’-ACTACACTTTCGGGTGGCTTGG-3’ and 5’-CAGTATAAGCGCCAGTTTCATC-3’)1919 Oho T, Yamashita Y, Shimazaki Y, Kushiyama M, Koga T. Simple and rapid detection of Streptococcus mutans and Streptococcus sobrinus in human saliva by polimerase chain reaction. Oral Microbiol Immunol. 2000 Aug;15(4):258-62. http://dx.doi.org/10.1034/j.1399-302x.2000.150408.x. PMid:11154412.
http://dx.doi.org/10.1034/j.1399-302x.20...
. After identification, S. mutans isolates were genotyped by RAPD with two arbitrary primers OPA-02 (5’-TGCCGAGCTG-3’)1111 Li Y, Caufield PW. Arbitrarily primed polymerase chain reaction fingerprinting for the genotypic identification of mutans streptococci from humans. Oral Microbiol Immunol. 1998 Feb;13(1):17-22. http://dx.doi.org/10.1111/j.1399-302X.1998.tb00745.x. PMid:9573817.
http://dx.doi.org/10.1111/j.1399-302X.19...
and OPA-13 (5’-CAGCACCCAC-3’)1010 Saarela M, Hannula J, Mättö J, Asikainen S, Alaluusua S. Typing of mutans streptococci by arbitrarily primed polymerase chain reaction. Arch Oral Biol. 1996 Aug-Sep;41(8-9):821-6. http://dx.doi.org/10.1016/S0003-9969(96)00049-0. PMid:9022919.
http://dx.doi.org/10.1016/S0003-9969(96)...
. In every reaction, S. mutans ATCC 25175 chromosomal DNA and ultrapure water were supplied as positive and negative controls, respectively.
The amplifications were performed in a thermocycler (Multi Gene II - Labnet International, São Paulo, Brazil) and the products were separated by electrophoresis (80 Volts) on 2% agarose gels in Tris-borate-EDTA running buffer (0,5X - pH 8,0). In each gel, 100-bp DNA Ladder (Invitrogen Life Technologies, São Paulo, Brazil) was run in one of the columns as a molecular weight marker. Ethidium bromide-stained (5 μg/mL) gel images were visualized under ultraviolet light (Model ENF - 280C, Spectroline Long Life, New York, USA) and captured with a digital camera. The RAPD fingerprints were analysed by side-by-side visual comparison and regarded as similar when the major bands were identical. The data were entered into a binary matrix and a similarity matrix was constructed with the arithmetical complement of Dice similarity coefficient. The genetic dissimilarities were represented in a dendrogram generated by the UPGMA (unweighted pair group method with arithmetic mean) with the program NTSYS-PC (Numerical Taxonomy System, Applied Biostatistics, Setauket, New York, USA).
After the determination of the mean distinct genotypes of S. mutans from each oral cavity, the preschoolers were categorized as having lower genotypic diversity (below the group mean) or greater genotypic diversity (above the group mean). Statistical analysis was performed using the Statistical Package for the Social Sciences (SPSS) 17.0 software (SPSS Inc., Chicago, Ill., USA). The Mann-Whitney test was used to compare dmft index values between G1 and G2. The chi-square test and the Spearman rank correlation test were used to analyse the associations and correlations between the results of RAPD and dental caries experience, respectively. The level of statistical significance was set at P < .05 and the confidence interval at 95%.
RESULT
In our study, S. mutans colonies were isolated from 10 caries-free subjects and 10 caries-active in both groups. (G1 mean age = 65 months and dmft index = 1.75; 95 percent confidence interval = 0.98 to 2.38; G2 mean age = 69 months and dmft index = 2.1; 95 percent confidence interval = 1.34 to 3.02). As the data related to dental caries experience did not present a normal distribution, we used the Mann-Whitney test to compare the populations, which did not show significant differences between the two groups (P = 0.082).
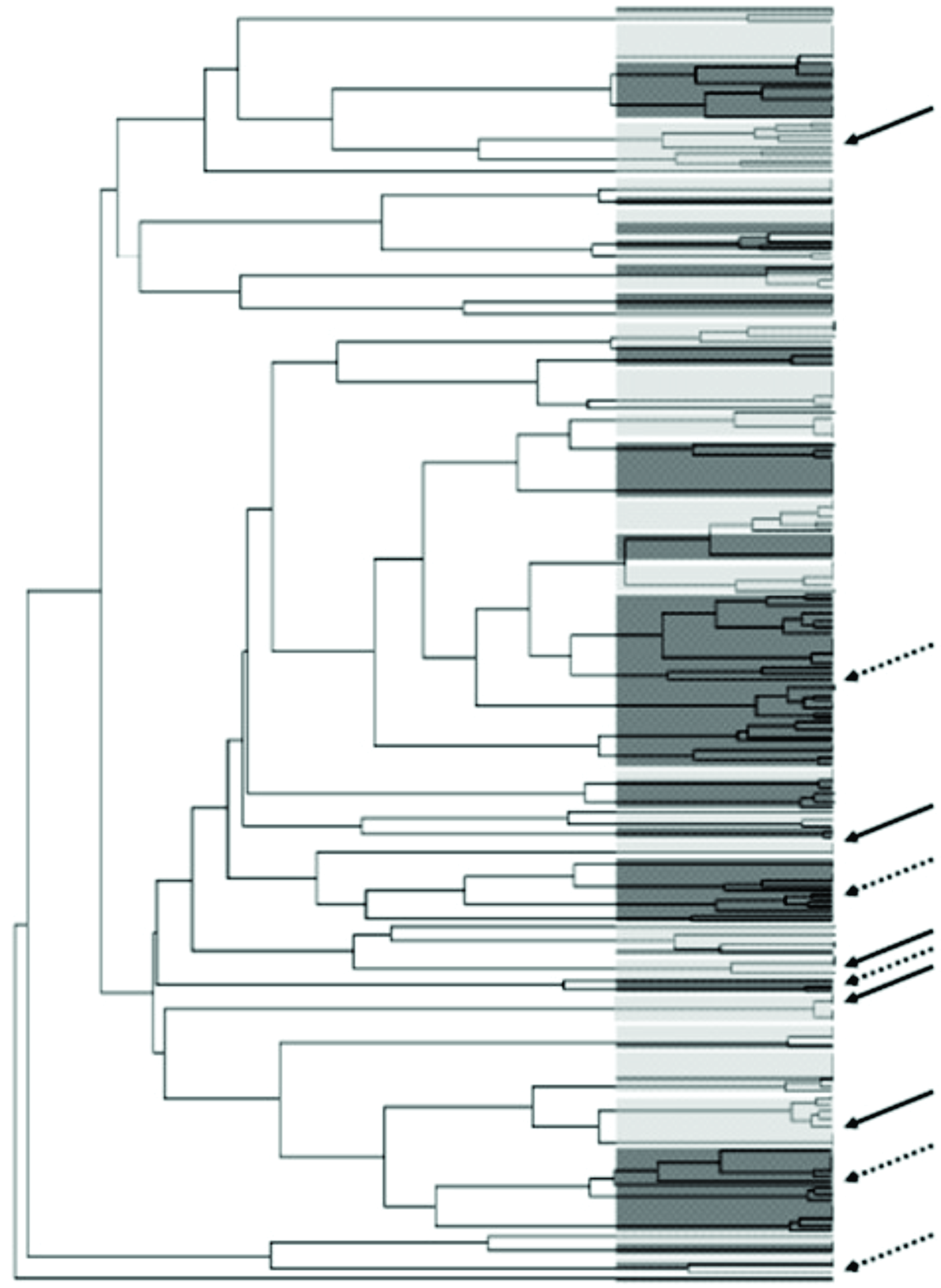
From 400 presumptive colonies, a total of 339 isolates (173 from G1 and 166 from G2) were identified as species S. mutans by PCR, being five to ten isolates from each subject. The results of RAPD revealed 75 distinct genotypes of S. mutans in G1 and 73 in G2, when the two primers were combined. Through analysis of the comparative dendrograms, we observed 22 (29%) genotypes exclusive to G1 and 51 (70%) that were exclusive to G2 (Figure 1). Figure 2 represents the genotypic diversity of S. mutans found in G1 subject with dental caries based on the profile of amplifications generated by primers OPA-02 and OPA-13. Amplification of DNA from S. mutans isolates with the OPA-02 primer resulted in fragments ranging from 250 to 1400-bp in size, whereas primer OPA-13 generated fragments ranging from 300 to 1500-bp in size.
Dendrogram based on random amplified polymorphic DNA profiles, showing genotypic diversity of S. mutans between G1 and G2 subjects. Filled arrows indicate the segments containing exclusive genotypes of G1 and the dotted arrows indicate exclusive genotypes of G2.
Agarose gel electrophoresis (2%) of the RAPD profiles of S. mutans strains isolated from a G1 subject with dental caries. DNA polymorphism was obtained with primers OPA-02 (A) and OPA-13 (B). Designations of isolates are indicated on the top. RAPD: Random Amplified Polymorphic DNA, M: molecular weight marker (100-bp DNA Ladder), C+: control positive, NO: control negative.
G1 and G2 subjects harboured in their oral cavities one to eight (mean = 3.75) and one to seven (mean = 3.65) distinct genotypes of S. mutans, respectively. Based on these results, the preschoolers were categorized as lower genotypic diversity (one to four distinct genotypes of S. mutans) or greater genotypic diversity (five to eight distinct genotypes of S. mutans). The analysis between genotypic diversity of S. mutans and experience of dental caries is presented in Table 1. A statistically significant association (P = 0.021) and a moderate correlation (r = 0.503) were observed between dental caries experience and genotypic diversity of S. mutans in G1.
DISCUSSION
The maintenance of the oral health of the infantile population is a commitment of the current preventive dentistry. The application of the concept of early dental care provided for the development of dentistry for babies and therefore, the implantation of centres of attention for this population. However, the fact that some children still present dental caries has drawn attention to the necessity for studies about the risks and aetiologic factors associated with disease.
Genetic and molecular biology studies point to a wide genotypic diversity of S. mutans1515 Tabchoury CP, Sousa MC, Arthur RA, Mattos-Graner RO, Del Bel Cury AA, Cury JA. Evaluation of genotypic diversity of streptococcus mutans using distinct arbitrary primers. J Appl Oral Sci. 2008 Nov-Dec;16(6):403-7. http://dx.doi.org/10.1590/S1678-77572008000600009. PMid:19082399.
http://dx.doi.org/10.1590/S1678-77572008...
,1717 Mattos-Graner RO, Li Y, Caufield PW, Duncan M, Smith DJ. Genotypic diversity of mutans streptococci in Brazilian nursery children suggests horizontal transmission. J Clin Microbiol. 2001 Jun;39(6):2313-6. http://dx.doi.org/10.1128/JCM.39.6.2313-2316.2001.
http://dx.doi.org/10.1128/JCM.39.6.2313-...
. Our findings are similar to earlier reports that found up to eight distinct genotypes of S. mutans colonizing the human oral cavity11 Lembo FL, Longo PL, Ota-Tsuzuki C, Rodrigues CR, Mayer MP. Genotypic and phenotypic analysis of Streptococcus mutans from different oral cavity sites of caries-free and caries-active children. Oral Microbiol Immunol. 2007 Oct;22(5):313-9. http://dx.doi.org/10.1111/j.1399-302X.2007.00361.x. PMid:17803628.
http://dx.doi.org/10.1111/j.1399-302X.20...
,77 Napimoga MH, Kamiya RU, Rosa RT, Rosa EA, Höfling JF, Mattos-Graner R, et al. Genotypic diversity and virulence traits of Streptococcus mutans in caries-free and caries-active individuals. J Med Microbiol. 2004 Jul;53(Pt 7):697-703. http://dx.doi.org/10.1099/jmm.0.05512-0. PMid:15184543.
http://dx.doi.org/10.1099/jmm.0.05512-0...
,88 Pieralisi FJ, Rodrigues MR, Segura VG, Maciel SM, Ferreira FB, Garcia JE, et al. Genotypic diversity of Streptococcus mutans in caries-free and caries-active preschool children. Int J Dent. 2010;2010:824976. http://dx.doi.org/10.1155/2010/824976. PMid:20351760.
http://dx.doi.org/10.1155/2010/824976...
,1313 Redmo Emanuelsson IM, Carlsson P, Hamberg K, Bratthall D. Tracing genotypes of mutans streptococci on tooth sites by random amplified polymorphic DNA (RAPD) analysis. Oral Microbiol Immunol. 2003 Feb;18(1):24-9. http://dx.doi.org/10.1034/j.1399-302X.2002.180104.x. PMid:12588455.
http://dx.doi.org/10.1034/j.1399-302X.20...
,2020 Braga MP, Piovesan A, Valarini N, Maciel SM, Andrade FB, Poli-Frederico RC. Genotypic diversity and virulence factors of Streptococcus mutans in caries-free and caries-active individuals. Braz Arch Biol Technol. 2013 Apr;56(2):241-8. http://dx.doi.org/10.1590/S1516-89132013000200009.
http://dx.doi.org/10.1590/S1516-89132013...
. Redmo Emanuelsson et al.1313 Redmo Emanuelsson IM, Carlsson P, Hamberg K, Bratthall D. Tracing genotypes of mutans streptococci on tooth sites by random amplified polymorphic DNA (RAPD) analysis. Oral Microbiol Immunol. 2003 Feb;18(1):24-9. http://dx.doi.org/10.1034/j.1399-302X.2002.180104.x. PMid:12588455.
http://dx.doi.org/10.1034/j.1399-302X.20...
suggest that a greater number of isolates analysed would increase the possibility of detecting different genotypes. However, Napimoga et al.77 Napimoga MH, Kamiya RU, Rosa RT, Rosa EA, Höfling JF, Mattos-Graner R, et al. Genotypic diversity and virulence traits of Streptococcus mutans in caries-free and caries-active individuals. J Med Microbiol. 2004 Jul;53(Pt 7):697-703. http://dx.doi.org/10.1099/jmm.0.05512-0. PMid:15184543.
http://dx.doi.org/10.1099/jmm.0.05512-0...
showed no relationship between the number of strains found in each subject and the number of genotypes detected. In the present study, the number of isolates of S. mutans, evaluated by child, supports previous findings33 Alaluusua S, Mättö J, Grönroos L, Innilä S, Torkko H, Asikainen S, et al. Oral colonization by more than one clonal type of mutans streptococcus in children with nursing-bottle dental caries. Arch Oral Biol. 1996 Feb;41(2):167-73. http://dx.doi.org/10.1016/0003-9969(95)00111-5. PMid:8712973.
http://dx.doi.org/10.1016/0003-9969(95)0...
,1717 Mattos-Graner RO, Li Y, Caufield PW, Duncan M, Smith DJ. Genotypic diversity of mutans streptococci in Brazilian nursery children suggests horizontal transmission. J Clin Microbiol. 2001 Jun;39(6):2313-6. http://dx.doi.org/10.1128/JCM.39.6.2313-2316.2001.
http://dx.doi.org/10.1128/JCM.39.6.2313-...
which indicate five to ten isolates from each individual must be investigated.
Distinct clonal types of S. mutans detected within the oral cavity of one subject can have different phenotypic and genetic properties2121 Caufield PW. Dental caries - a transmissible and infectious disease revisited: a position paper. Pediatr Dent. 1997 Nov-Dec;19(8):491-8. PMid:9442545.. The identification of 22 exclusive genotypes in preschoolers with dental care since their first year demonstrates the necessity of future research efforts aimed at exploring these S. mutans strains and their virulence factors, such as water-insoluble glucan synthesis, final pH of cultures and sucrose-dependent adherence. Additionally, the presence of 51 exclusive genotypes of S. mutans in G2 suggests a lack of selectivity of microorganisms; consequently, strong selective pressure does not occur in this population compared with G1. Li et al.2222 Li Y, Lau PC, Lee JH, Ellen RP, Cvitkovitch DG. Natural genetic transformation of Streptococcus mutans growing in biofilms. J Bacteriol. 2001 Feb;183(3):897-908. http://dx.doi.org/10.1128/JB.183.3.897-908.2001.
http://dx.doi.org/10.1128/JB.183.3.897-9...
add that the adaptation to an environmental stress by genetic transformation is a very infrequent event; however, when it occurs, it can be highly significant if the transforming DNA, such as an antibiotic resistance gene or a virulence factor, provides a selective advantage to the recipient cells.
The possession of a particular genotype or genotypes of S. mutans by an individual could be related to dental caries activity and the extent of genotypic diversity may reflect the nature of the oral environment1616 Bowden GH. Does assessment of microbial composition of plaque/saliva allow for diagnosis of disease activity of individuals? Community Dent Oral Epidemiol. 1997 Feb;25(1):76-81. http://dx.doi.org/10.1111/j.1600-0528.1997.tb00902.x. PMid:9088695.
http://dx.doi.org/10.1111/j.1600-0528.19...
. Although the variety in genotypes of S. mutans in both populations was numerically similar, we also observed that the diversity from the molecular genetic point of view influenced the occurrence of the disease in different ways. An interesting result was found in relation to the association of genotypic diversity of S. mutans and dental caries experience in G1, wherein the children with a larger number of genotypes presented disease. In addition, as observed in our study, previous studies33 Alaluusua S, Mättö J, Grönroos L, Innilä S, Torkko H, Asikainen S, et al. Oral colonization by more than one clonal type of mutans streptococcus in children with nursing-bottle dental caries. Arch Oral Biol. 1996 Feb;41(2):167-73. http://dx.doi.org/10.1016/0003-9969(95)00111-5. PMid:8712973.
http://dx.doi.org/10.1016/0003-9969(95)0...
,77 Napimoga MH, Kamiya RU, Rosa RT, Rosa EA, Höfling JF, Mattos-Graner R, et al. Genotypic diversity and virulence traits of Streptococcus mutans in caries-free and caries-active individuals. J Med Microbiol. 2004 Jul;53(Pt 7):697-703. http://dx.doi.org/10.1099/jmm.0.05512-0. PMid:15184543.
http://dx.doi.org/10.1099/jmm.0.05512-0...
,99 Zhou Q, Qin X, Qin M, Ge L. Genotypic diversity of Streptococcus mutans and Streptococcus sobrinus in 3-4-year-old children with severe caries or without caries. Int J Paediatr Dent. 2011 Nov;21(6):422-31. http://dx.doi.org/10.1111/j.1365-263X.2011.01145.x. PMid:21689176.
http://dx.doi.org/10.1111/j.1365-263X.20...
,2020 Braga MP, Piovesan A, Valarini N, Maciel SM, Andrade FB, Poli-Frederico RC. Genotypic diversity and virulence factors of Streptococcus mutans in caries-free and caries-active individuals. Braz Arch Biol Technol. 2013 Apr;56(2):241-8. http://dx.doi.org/10.1590/S1516-89132013000200009.
http://dx.doi.org/10.1590/S1516-89132013...
,2222 Li Y, Lau PC, Lee JH, Ellen RP, Cvitkovitch DG. Natural genetic transformation of Streptococcus mutans growing in biofilms. J Bacteriol. 2001 Feb;183(3):897-908. http://dx.doi.org/10.1128/JB.183.3.897-908.2001.
http://dx.doi.org/10.1128/JB.183.3.897-9...
,2323 Momeni SS, Whiddon J, Cheon K, Ghazal T, Moser SA, Childers NK. Genetic diversity and evidence for transmission of Streptococcus mutans by DiversiLab rep-PCR. J Microbiol Methods. 2016 Sep;128:108-17. http://dx.doi.org/10.1016/j.mimet.2016.07.010. PMid:27432341.
http://dx.doi.org/10.1016/j.mimet.2016.0...
have reported higher genotypic diversity of S. mutans in caries-active subjects than in caries-free individuals.
Heavy colonization of S. mutans and growth of multiple genotypes in the same oral cavity are likely to be consequences of frequent consumption of fermentable carbohydrates2121 Caufield PW. Dental caries - a transmissible and infectious disease revisited: a position paper. Pediatr Dent. 1997 Nov-Dec;19(8):491-8. PMid:9442545.. Alaluusua et al.33 Alaluusua S, Mättö J, Grönroos L, Innilä S, Torkko H, Asikainen S, et al. Oral colonization by more than one clonal type of mutans streptococcus in children with nursing-bottle dental caries. Arch Oral Biol. 1996 Feb;41(2):167-73. http://dx.doi.org/10.1016/0003-9969(95)00111-5. PMid:8712973.
http://dx.doi.org/10.1016/0003-9969(95)0...
suggest that this consumption could be favourable for the acquisition of certain strains by children and that the simultaneous action of several genotypes with possibly differing cariogenic potentials further increases the risk of dental caries. According to Lembo et al.11 Lembo FL, Longo PL, Ota-Tsuzuki C, Rodrigues CR, Mayer MP. Genotypic and phenotypic analysis of Streptococcus mutans from different oral cavity sites of caries-free and caries-active children. Oral Microbiol Immunol. 2007 Oct;22(5):313-9. http://dx.doi.org/10.1111/j.1399-302X.2007.00361.x. PMid:17803628.
http://dx.doi.org/10.1111/j.1399-302X.20...
, S. mutans genotypes that are highly resistant to acid shock are more prevalent in caries-active children than in caries-free children, and this finding may help to explain the differences in dental caries experience observed among S. mutans infected children.
The knowledge that dental caries is a multifactorial, diet-dependent, fluoride-mediated disease with genetic and behavioral susceptibility components2424 Edelstein BL, Ureles SD, Smaldone A. Very high salivary Streptococcus mutans predicts caries progression in young children. Pediatr Dent. 2016;38(4):325-30. PMid:27557922. and that its prevention involves actions on the main predisposing factors helps us to understand why some children still experience dental caries, even with a history of dental care since their first year. Although some variables were not explored in this study, as dietary and behavioral factors, our results support the findings of genetic diversity of S. mutans in preschool children being associated with dental caries. The results emphasize the importance of the educative-preventive actions carried out by dental program and in their domiciles, mainly through the control of behavioural habits.
In addition to environmental and host factors, such as diet, the identification of specific pathogenic genotypes of S. mutans that may be more virulent colonizers might predict sites that are more susceptible to disease2525 Napimoga MH, Höfling JF, Klein MI, Kamiya RU, Gonçalves RB. Transmission, diversity and virulence factors of Streptococcus mutans genotypes. J Oral Sci. 2005 Jun;47(2):59-64. http://dx.doi.org/10.2334/josnusd.47.59. PMid:16050484.
http://dx.doi.org/10.2334/josnusd.47.59...
. We suggest an expansion of this study to include the analysis of the biofilm formed on dental caries lesions and research into the persistence of those genotypes in the children's oral cavities. The present study, a molecular biology investigation to evaluate genetically S. mutans isolated from a differentiated population, emphasizes the necessity of identification of individuals at high risk for dental caries and paves the way for new studies aimed at better understanding the genotypic diversity of S. mutans in the development of dental caries.
CONCLUSION
Although our results point higher genotypic diversity of S. mutans in children with dental caries experience and dental care since their first year, from the limitations of the study design, we just can hypothesize that the actions of early dental carried out by an educative-preventive program can contribute to the establishment of a distinct oral microbiota.
ACKNOWLEDGEMENT
This investigation was supported by grants from the National Foundation for the Development of Private Higher Education (FUNADESP, Brazil). We are grateful to all subjects for their essential participation and to Jefferson Rosa Cardoso for the statistical analysis.
-
How to cite: Cota ALS, Junkes JA, Poli-Frederico RC. Genotypic diversity of Streptococcus mutans isolated from preschoolers with and without early dental care. Rev Odontol UNESP. 2019;48:e20180080. https://dx.doi.org/10.1590/1807-2577.08018.
REFERENCES
-
1Lembo FL, Longo PL, Ota-Tsuzuki C, Rodrigues CR, Mayer MP. Genotypic and phenotypic analysis of Streptococcus mutans from different oral cavity sites of caries-free and caries-active children. Oral Microbiol Immunol. 2007 Oct;22(5):313-9. http://dx.doi.org/10.1111/j.1399-302X.2007.00361.x PMid:17803628.
» http://dx.doi.org/10.1111/j.1399-302X.2007.00361.x -
2Longo PL, Mattos-Graner RO, Mayer MP. Determination of mutacin activity and detection of mutA genes in Streptococcus mutans genotypes from caries-free and caries-active children. Oral Microbiol Immunol. 2003 Jun;18(3):144-9. http://dx.doi.org/10.1034/j.1399-302X.2003.00042.x PMid:12753464.
» http://dx.doi.org/10.1034/j.1399-302X.2003.00042.x -
3Alaluusua S, Mättö J, Grönroos L, Innilä S, Torkko H, Asikainen S, et al. Oral colonization by more than one clonal type of mutans streptococcus in children with nursing-bottle dental caries. Arch Oral Biol. 1996 Feb;41(2):167-73. http://dx.doi.org/10.1016/0003-9969(95)00111-5 PMid:8712973.
» http://dx.doi.org/10.1016/0003-9969(95)00111-5 -
4Liu Y, Zou J, Shang R, Zhou XD. Genotypic diversity of Streptococcus mutans in 3- to 4-year-old Chinese nursery children suggests horizontal transmission. Arch Oral Biol. 2007 Sep;52(9):876-81. http://dx.doi.org/10.1016/j.archoralbio.2007.03.004 PMid:17466259.
» http://dx.doi.org/10.1016/j.archoralbio.2007.03.004 -
5Momeni SS, Whiddon J, Moser SA, Childers NK. Transmission patterns of Streptococcus mutans demonstrated by a combined rep-PCR and MLST approach. Clin Oral Investig. 2018 Nov;22(8):2847-58. http://dx.doi.org/10.1007/s00784-018-2371-8 PMid:29476335.
» http://dx.doi.org/10.1007/s00784-018-2371-8 -
6Emanuelsson I-MR, Wang X. Demonstration of identical strains of mutans streptococci within Chinese families by genotyping. Eur J Oral Sci. 1998 Jun;106(3):788-94. http://dx.doi.org/10.1046/j.0909-8836.1998.eos106305.x PMid:9672101.
» http://dx.doi.org/10.1046/j.0909-8836.1998.eos106305.x -
7Napimoga MH, Kamiya RU, Rosa RT, Rosa EA, Höfling JF, Mattos-Graner R, et al. Genotypic diversity and virulence traits of Streptococcus mutans in caries-free and caries-active individuals. J Med Microbiol. 2004 Jul;53(Pt 7):697-703. http://dx.doi.org/10.1099/jmm.0.05512-0 PMid:15184543.
» http://dx.doi.org/10.1099/jmm.0.05512-0 -
8Pieralisi FJ, Rodrigues MR, Segura VG, Maciel SM, Ferreira FB, Garcia JE, et al. Genotypic diversity of Streptococcus mutans in caries-free and caries-active preschool children. Int J Dent. 2010;2010:824976. http://dx.doi.org/10.1155/2010/824976 PMid:20351760.
» http://dx.doi.org/10.1155/2010/824976 -
9Zhou Q, Qin X, Qin M, Ge L. Genotypic diversity of Streptococcus mutans and Streptococcus sobrinus in 3-4-year-old children with severe caries or without caries. Int J Paediatr Dent. 2011 Nov;21(6):422-31. http://dx.doi.org/10.1111/j.1365-263X.2011.01145.x PMid:21689176.
» http://dx.doi.org/10.1111/j.1365-263X.2011.01145.x -
10Saarela M, Hannula J, Mättö J, Asikainen S, Alaluusua S. Typing of mutans streptococci by arbitrarily primed polymerase chain reaction. Arch Oral Biol. 1996 Aug-Sep;41(8-9):821-6. http://dx.doi.org/10.1016/S0003-9969(96)00049-0 PMid:9022919.
» http://dx.doi.org/10.1016/S0003-9969(96)00049-0 -
11Li Y, Caufield PW. Arbitrarily primed polymerase chain reaction fingerprinting for the genotypic identification of mutans streptococci from humans. Oral Microbiol Immunol. 1998 Feb;13(1):17-22. http://dx.doi.org/10.1111/j.1399-302X.1998.tb00745.x PMid:9573817.
» http://dx.doi.org/10.1111/j.1399-302X.1998.tb00745.x -
12Grönroos L, Alaluusua S. Site-specific oral colonization of mutans streptococci detected by arbitrarily primed PCR fingerprinting. Caries Res. 2000 Nov-Dec;34(6):474-80. http://dx.doi.org/10.1159/000016626 PMid:11093021.
» http://dx.doi.org/10.1159/000016626 -
13Redmo Emanuelsson IM, Carlsson P, Hamberg K, Bratthall D. Tracing genotypes of mutans streptococci on tooth sites by random amplified polymorphic DNA (RAPD) analysis. Oral Microbiol Immunol. 2003 Feb;18(1):24-9. http://dx.doi.org/10.1034/j.1399-302X.2002.180104.x PMid:12588455.
» http://dx.doi.org/10.1034/j.1399-302X.2002.180104.x -
14Hames-Kocabas EE, Uçar F, Kocatas Ersin N, Uzel A, Alpöz AR. Colonization and vertical transmission of Streptococcus mutans in Turkish children. Microbiol Res. 2008;163(2):168-72. http://dx.doi.org/10.1016/j.micres.2006.03.016 PMid:16735109.
» http://dx.doi.org/10.1016/j.micres.2006.03.016 -
15Tabchoury CP, Sousa MC, Arthur RA, Mattos-Graner RO, Del Bel Cury AA, Cury JA. Evaluation of genotypic diversity of streptococcus mutans using distinct arbitrary primers. J Appl Oral Sci. 2008 Nov-Dec;16(6):403-7. http://dx.doi.org/10.1590/S1678-77572008000600009 PMid:19082399.
» http://dx.doi.org/10.1590/S1678-77572008000600009 -
16Bowden GH. Does assessment of microbial composition of plaque/saliva allow for diagnosis of disease activity of individuals? Community Dent Oral Epidemiol. 1997 Feb;25(1):76-81. http://dx.doi.org/10.1111/j.1600-0528.1997.tb00902.x PMid:9088695.
» http://dx.doi.org/10.1111/j.1600-0528.1997.tb00902.x -
17Mattos-Graner RO, Li Y, Caufield PW, Duncan M, Smith DJ. Genotypic diversity of mutans streptococci in Brazilian nursery children suggests horizontal transmission. J Clin Microbiol. 2001 Jun;39(6):2313-6. http://dx.doi.org/10.1128/JCM.39.6.2313-2316.2001
» http://dx.doi.org/10.1128/JCM.39.6.2313-2316.2001 -
18World Health Organization – WHO. Oral health surveys, basic methods. 4th ed. Geneva: WHO; 1997.
-
19Oho T, Yamashita Y, Shimazaki Y, Kushiyama M, Koga T. Simple and rapid detection of Streptococcus mutans and Streptococcus sobrinus in human saliva by polimerase chain reaction. Oral Microbiol Immunol. 2000 Aug;15(4):258-62. http://dx.doi.org/10.1034/j.1399-302x.2000.150408.x PMid:11154412.
» http://dx.doi.org/10.1034/j.1399-302x.2000.150408.x -
20Braga MP, Piovesan A, Valarini N, Maciel SM, Andrade FB, Poli-Frederico RC. Genotypic diversity and virulence factors of Streptococcus mutans in caries-free and caries-active individuals. Braz Arch Biol Technol. 2013 Apr;56(2):241-8. http://dx.doi.org/10.1590/S1516-89132013000200009
» http://dx.doi.org/10.1590/S1516-89132013000200009 -
21Caufield PW. Dental caries - a transmissible and infectious disease revisited: a position paper. Pediatr Dent. 1997 Nov-Dec;19(8):491-8. PMid:9442545.
-
22Li Y, Lau PC, Lee JH, Ellen RP, Cvitkovitch DG. Natural genetic transformation of Streptococcus mutans growing in biofilms. J Bacteriol. 2001 Feb;183(3):897-908. http://dx.doi.org/10.1128/JB.183.3.897-908.2001
» http://dx.doi.org/10.1128/JB.183.3.897-908.2001 -
23Momeni SS, Whiddon J, Cheon K, Ghazal T, Moser SA, Childers NK. Genetic diversity and evidence for transmission of Streptococcus mutans by DiversiLab rep-PCR. J Microbiol Methods. 2016 Sep;128:108-17. http://dx.doi.org/10.1016/j.mimet.2016.07.010 PMid:27432341.
» http://dx.doi.org/10.1016/j.mimet.2016.07.010 -
24Edelstein BL, Ureles SD, Smaldone A. Very high salivary Streptococcus mutans predicts caries progression in young children. Pediatr Dent. 2016;38(4):325-30. PMid:27557922.
-
25Napimoga MH, Höfling JF, Klein MI, Kamiya RU, Gonçalves RB. Transmission, diversity and virulence factors of Streptococcus mutans genotypes. J Oral Sci. 2005 Jun;47(2):59-64. http://dx.doi.org/10.2334/josnusd.47.59 PMid:16050484.
» http://dx.doi.org/10.2334/josnusd.47.59
Publication Dates
-
Publication in this collection
03 June 2019 -
Date of issue
2019
History
-
Received
16 Aug 2018 -
Accepted
04 Apr 2019