Abstract
Equine piroplasmosisis, a tick-borne disease caused by the intra-erythrocytic protozoans Babesia caballi and Theileria equi, has economic importance due to the international trade and the increased movement of horses all over the world. The goal of this study was to evaluate the occurrence of phylogenetic diversity of T. equi and B. caballi genotypes among infected equids from São Luís Island, state of Maranhão, northeastern Brazil. Between December of 2011 and June of 2012, EDTA-blood and serum samples were collected from 139 equids (90 donkeys, 39 horses and 10 mules). From 139 serum samples submitted to ELISA assay, IgG antibodies to T. equi and B. caballi were detected in 19.4% (27/139) and 25.2% (35/139), respectively. Among sampled animals, 21.6% (30/139) and 55.4% (77/139) were positive for cPCR assays for T. equi and B. caballi, based on ema-1 and rap-1 genes, respectively. Overall, the T. equi sequences (n=7) submitted to Maximum Likelihood analysis (based on a 18S rRNA fragment of 1700 bp after alignment) grouped into three main groups, which were subdivided in eight clusters. The present work showed that different genotypes of T. equi and B. caballi circulate among equids in Brazil.
Keywords:
Babesia caballi; Theileria equi; phylogeny; 18S rRNA
Resumo
A piroplasmose equina, uma doença transmitida por carrapatos e causada pelos protozoários intra-eritrocíticos Babesia caballi e Theileria equi, tem importância econômica devido ao comércio internacional e ao aumento do movimento de cavalos em todo o mundo. O objetivo do presente estudo foi mostrar a diversidade filogenética de T. equi e B. caballi infectando cavalos, burros e jumentos na Ilha de São Luís, Estado do Maranhão, Nordeste do Brasil. Entre dezembro de 2011 e junho de 2012, amostras de sangue com EDTA e soro de foram coletadas de 139 equídeos (90 jumentos, 39 cavalos e 10 burros). Dentre as 139 amostras de soro submetidas ao ensaio de ELISA, foram detectados anticorpos IgG contra T. equi e B. caballi em 19,4% (27/139) e 25,2% (35/139), respectivamente. Entre os animais amostrados, 21,6% (30/139) e 55,4% (77/139) foram positivos por meio dos ensaios de cPCR para T. equi e B. caballi, com base nos genes ema-1 e rap-1, respectivamente. No geral, as sequências T. equi (n = 7) submetidas à análise de Máxima Verossimilhança (baseada em um fragmento do 18S rRNA de 1700 pb, após o alinhamento) foram agrupadas em três grupos principais, os quais foram subdivididos em oito grupos. O presente trabalho mostrou que diferentes genótipos de T. equi e B. caballi circulam entre equídeos no Brasil.
Palavras-chave:
Babesia caballi; Theileria equi; filogenia; 18S rRNA
Introduction
Equine piroplasmosisis is a tick-borne disease caused by intra-erythrocytic protozoans, namely Babesia caballi and Theileria equi, which is associated with an illness characterized by fever, anemia, dyspnea, icterus, hepato-and splenomegaly, edema, intravascular hemolysis, petecheal hemorrhages in mucosal surfaces, hemoglobinuria, and even death (SCHEIN, 1988Schein E. Equine babesiosis. In: Ristic M, editors. Babesiosis of domestic animals and man. Boca Raton: CRS Press; 1988. p. 197-208.; UILENBERG, 2006Uilenberg G. Babesia--a historical overview. Vet Parasitol 2006; 138(1-2): 3-10. http://dx.doi.org/10.1016/j.vetpar.2006.01.035.
http://dx.doi.org/10.1016/j.vetpar.2006....
).
This disease has economic importance due to the international trade and the increased movement of horses all over the world (BASHIRUDDIN et al., 1999Bashiruddin JB, Cammà C, Rebêlo E. Molecular detection of Babesia equi and Babesia caballi in horse blood by PCR amplification of part of the 16S RNA gene. Vet Parasitol 1999; 84(1-2): 75-83. PMid:10435792. http://dx.doi.org/10.1016/S0304-4017(99)00049-7.
http://dx.doi.org/10.1016/S0304-4017(99)...
; NICOLAIEWSKY et al., 2001Nicolaiewsky TB, Richter MF, Lunge VR, Cunha CW, Delagostin O, Ikuta N, et al. Detection of Babesia equi (Laveran, 1901) by nested polymerase chain reaction. Vet Parasitol 2001; 101(1): 9-21. PMid:11587829. http://dx.doi.org/10.1016/S0304-4017(01)00471-X.
http://dx.doi.org/10.1016/S0304-4017(01)...
). Nowadays, some disease-free countries usually require serological test in horses in order to prevent seropositive animals from entering their territories (BÖSE et al., 1995Böse R, Jorgensen WK, Dalgliesh RJ, Friedhoff KT, de Vos AJ. Current state and future trends in the diagnosis of babesiosis. Vet Parasitol 1995; 57(1-3): 61-74. PMid:7597794. http://dx.doi.org/10.1016/0304-4017(94)03111-9.
http://dx.doi.org/10.1016/0304-4017(94)0...
; BRÜNING, 1996Brüning A. Equine piroplasmosis an update on diagnosis, treatment and prevention. Br Vet J 1996; 152(2): 139-151. PMid:8680838. http://dx.doi.org/10.1016/S0007-1935(96)80070-4.
http://dx.doi.org/10.1016/S0007-1935(96)...
). Serological techniques, such as indirect fluorescent antibody (IFA) test and enzyme-linked-immunosorbent assay (ELISA), have been considered as preferable tests to check the exposure of horses to piroplasmids for international trade by the World Organization trade for Animal Health (OIE, 2011World Organization for Animal Health – OIE. Manual of diagnostic tests and vaccines for terrestrial animals. Paris: OIE; 2011.).
In Brazil, T. equi and B. caballi are mainly transmitted by Rhipicephalus (Boophilus) microplus and Anocentor nitens, respectively (GUIMARÃES et al., 1998Guimarães AM, Lima JD, Ribeiro MFB. Sporogony and experimental transmission of Babesia equi by Boophilus microplus. Parasitol Res 1998; 84(4): 323-327. PMid:9580425. http://dx.doi.org/10.1007/s004360050404.
http://dx.doi.org/10.1007/s004360050404...
; BATTSETSEG et al., 2002Battsetseg B, Lucero S, Xuan X, Claveria FG, Inoue N, Alhassan A, et al. Detection of natural infection of Boophilus microplus with Babesia equi and Babesia caballi in Brazilian horses using nested polymerase chain reaction. Vet Parasitol 2002; 107(4): 351-357. PMid:12163246. http://dx.doi.org/10.1016/S0304-4017(02)00131-0.
http://dx.doi.org/10.1016/S0304-4017(02)...
). In this country, the prevalence among horses ranges from 22 to 100% for T. equi (TENTER & FRIEDHOFF, 1986Tenter AM, Friedhoff KT. Serodiagnosis of experimental and natural Babesia equi and B. caballi infections. Vet Parasitol 1986; 20(1-3): 49-61. PMid:3518217. http://dx.doi.org/10.1016/0304-4017(86)90092-0.
http://dx.doi.org/10.1016/0304-4017(86)9...
; PFEIFER BARBOSA et al., 1995Pfeifer Barbosa I, Böse R, Peymann B, Friedhoff KT. Epidemiological aspects of equine babesioses in a herd of horses in Brazil. Vet Parasitol 1995; 58(1-2): 1-8. PMid:7676590. http://dx.doi.org/10.1016/0304-4017(94)00704-G.
http://dx.doi.org/10.1016/0304-4017(94)0...
; RIBEIRO et al., 1999Ribeiro MFB, Costa JO, Guimarães AM. Epidemiological aspects of Babesia equi in Horses in Minas Gerais, Brazil. Vet Res Commun 1999; 23(6): 385-390. PMid:10543367. http://dx.doi.org/10.1023/A:1006389725759.
http://dx.doi.org/10.1023/A:100638972575...
; HEUCHERT et al., 1999Heuchert CM, Giulli V Jr, Athaide D, Böse R, Friedhoff KT. Seroepidemiologic studies on Babesia equi and Babesia caballi infections in Brazil. Vet Parasitol 1999; 85(1): 1-11. PMid:10447188. http://dx.doi.org/10.1016/S0304-4017(99)00108-9.
http://dx.doi.org/10.1016/S0304-4017(99)...
; HEIM et al., 2007Heim A, Passos LM, Ribeiro MF, Costa-Júnior LM, Bastos CV, Cabral DD, et al. Detection and molecular characterization of Babesia caballi and Theileria equi isolates from endemic areas of Brazil. Parasitol Res 2007; 102(1): 63-68. PMid:17828553. http://dx.doi.org/10.1007/s00436-007-0726-1.
http://dx.doi.org/10.1007/s00436-007-072...
; GOLYNSKI et al., 2008Golynski AA, Fernandes KR, Baldani CD, Golynski AL, Madeiro AS, Machado RZ, et al. Seroepidemiological studies on Babesia equi in horses from the State of Rio Grande do Sul determined by indirect immunoflourecence test and Elisa. Rev Bras Parasitol Vet 2008;17(Suppl 1): 317-321. PMid:20059868.; KERBER et al., 2009Kerber CE, Labruna MB, Ferreira F, De Waal DT, Knowles DP, Gennari SM. Prevalence of equine Piroplasmosis and its association with tick infestation in the State of São Paulo, Brazil. Rev Bras Parasitol Vet 2009; 18(4): 1-8. PMid:20040201. http://dx.doi.org/10.4322/rbpv.01804001.
http://dx.doi.org/10.4322/rbpv.01804001...
; BALDANI et al., 2010Baldani CD, Nakaghi ACH, Machado RZ. Occurrence of Theileria equi in horses raised in the Jaboticabal microregion, São Paulo state, Brazil. Rev Bras Parasitol Vet 2010; 19(4): 228-232. PMid:21184699. http://dx.doi.org/10.1590/S1984-29612010000400007.
http://dx.doi.org/10.1590/S1984-29612010...
; SALVAGNI et al., 2010Salvagni CA, Dagnone AS, Gomes TS, Mota JS, Andrade GM, Baldani CD, et al. Serologic evidence of equine granulocytic anaplasmosis in horses from central West Brazil. Rev Bras Parasitol Vet 2010; 19(3): 135-140. PMid:20943015. http://dx.doi.org/10.1590/S1984-29612010000300002.
http://dx.doi.org/10.1590/S1984-29612010...
; MACHADO et al., 2012Machado RZ, Toledo CZ, Teixeira MC, André MR, Freschi CR, Sampaio PH. Molecular and serological detection of Theileria equi and Babesia caballi in donkeys (Equus asinus) in Brazil. Vet Parasitol 2012; 186(3-4): 461-465. PMid:22186194. http://dx.doi.org/10.1016/j.vetpar.2011.11.069.
http://dx.doi.org/10.1016/j.vetpar.2011....
), and from 70 to 90% for B. caballi (PFEIFER BARBOSA et al., 1995Pfeifer Barbosa I, Böse R, Peymann B, Friedhoff KT. Epidemiological aspects of equine babesioses in a herd of horses in Brazil. Vet Parasitol 1995; 58(1-2): 1-8. PMid:7676590. http://dx.doi.org/10.1016/0304-4017(94)00704-G.
http://dx.doi.org/10.1016/0304-4017(94)0...
; HEIM et al., 2007Heim A, Passos LM, Ribeiro MF, Costa-Júnior LM, Bastos CV, Cabral DD, et al. Detection and molecular characterization of Babesia caballi and Theileria equi isolates from endemic areas of Brazil. Parasitol Res 2007; 102(1): 63-68. PMid:17828553. http://dx.doi.org/10.1007/s00436-007-0726-1.
http://dx.doi.org/10.1007/s00436-007-072...
; KERBER et al., 2009Kerber CE, Labruna MB, Ferreira F, De Waal DT, Knowles DP, Gennari SM. Prevalence of equine Piroplasmosis and its association with tick infestation in the State of São Paulo, Brazil. Rev Bras Parasitol Vet 2009; 18(4): 1-8. PMid:20040201. http://dx.doi.org/10.4322/rbpv.01804001.
http://dx.doi.org/10.4322/rbpv.01804001...
; MACHADO et al., 2012Machado RZ, Toledo CZ, Teixeira MC, André MR, Freschi CR, Sampaio PH. Molecular and serological detection of Theileria equi and Babesia caballi in donkeys (Equus asinus) in Brazil. Vet Parasitol 2012; 186(3-4): 461-465. PMid:22186194. http://dx.doi.org/10.1016/j.vetpar.2011.11.069.
http://dx.doi.org/10.1016/j.vetpar.2011....
), indicating that both parasites are endemic among horses.
Horses that recover from an acute phase of infection become reservoirs for both parasites (DE WAAL, 1992de Waal DT. Equine piroplasmosis: a review. Br Vet J 1992; 148(1): 6-14. PMid:1551016. http://dx.doi.org/10.1016/0007-1935(92)90061-5.
http://dx.doi.org/10.1016/0007-1935(92)9...
). On the other hand, donkeys usually show an asymptomatic form of piroplasmosis, showing a lower parasitaemia when compared to horses (KUMAR et al., 2009Kumar S, Kumar R, Sugimoto C. A perspective on Theileria equi infections in donkeys. Jpn J Vet Res 2009; 56(4): 171-180. PMid:19358444.; MACHADO et al., 2012Machado RZ, Toledo CZ, Teixeira MC, André MR, Freschi CR, Sampaio PH. Molecular and serological detection of Theileria equi and Babesia caballi in donkeys (Equus asinus) in Brazil. Vet Parasitol 2012; 186(3-4): 461-465. PMid:22186194. http://dx.doi.org/10.1016/j.vetpar.2011.11.069.
http://dx.doi.org/10.1016/j.vetpar.2011....
).
In the last years, molecular techniques, such as reverse line blot (RLB), hybridization (NAGORE et al., 2004Nagore D, García-Sanmartín J, Garcıa-Pérez AL, Juste RA, Hurtado A. Detection and identification of equine Theileria and Babesia species by reverse line blotting: epidemiological survey and phylogenetic analysis. Vet Parasitol 2004; 123(1-2): 41-54. PMid:15265570. http://dx.doi.org/10.1016/j.vetpar.2004.04.010.
http://dx.doi.org/10.1016/j.vetpar.2004....
) and quantitative real time PCR (BHOORA et al., 2010Bhoora R, Quan M, Franssen L, Butler CM, Van der Kolk JH, Guthrie AJ, et al. Development and evaluation of real-time PCR assays for the quantitative detection of Babesia caballi and Theileria equi infections in horses from South Africa. Vet Parasitol 2010; 168(3-4): 201-211. PMid:20031328. http://dx.doi.org/10.1016/j.vetpar.2009.11.011.
http://dx.doi.org/10.1016/j.vetpar.2009....
) have been developed for the diagnosis of equine piroplasmosis. The antigenic polymorphisms of merozoite surface antigens have been commonly observed between different isolates of hemoprotozoan parasites (CARCY et al., 2006Carcy B, Précigout E, Schetters T, Gorenflot A. Genetic basis for GPI-anchor merozoite surface antigen polymorphism of Babesia and resulting antigenic diversity. Vet Parasitol 2006; 138(1-2): 33-49. PMid:16551492. http://dx.doi.org/10.1016/j.vetpar.2006.01.038.
http://dx.doi.org/10.1016/j.vetpar.2006....
). Most of the studies of molecular occurrence of equid piroplasmids have targeted ema-1 gene for T. equi and rap-1gene for B. caballi. Additionally, 18S rRNA gene has been extensively used for broad phylogenetic classification (BATTSETSEG et al., 2002Battsetseg B, Lucero S, Xuan X, Claveria FG, Inoue N, Alhassan A, et al. Detection of natural infection of Boophilus microplus with Babesia equi and Babesia caballi in Brazilian horses using nested polymerase chain reaction. Vet Parasitol 2002; 107(4): 351-357. PMid:12163246. http://dx.doi.org/10.1016/S0304-4017(02)00131-0.
http://dx.doi.org/10.1016/S0304-4017(02)...
; UETI et al., 2003Ueti MW, Palmer GH, Kappmeyer LS, Scoles GA, Knowles DP. Expression of Equi Merozoite Antigen 2 during Development of Babesia equi in the Midgut and Salivary Gland of the Vector Tick Boophilus microplus. J Clin Microbiol 2003; 41(12): 5803-5809. PMid:14662988. http://dx.doi.org/10.1128/JCM.41.12.5803-5809.2003.
http://dx.doi.org/10.1128/JCM.41.12.5803...
).
Despite the endemicity of both parasites among horses in Brazil, the epidemiology of piroplasmosis remains poorly studied among donkeys and mules. The goal of this study was to evaluate the occurrence of phylogenetic diversity of T. equi and B. caballi genotypes among infected equids from São Luís Island, state of Maranhão, northeastern Brazil.
Materials and Methods
Animals and area of study
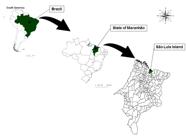
Between December of 2011 and June of 2012, a molecular and serological study was performed in São Luís Island (latitude -2° 31′ 48″ S, longitude - 44° 18′ 10″ W), state of Maranhão, northeastern Brazil (Figure 1). For this purpose, EDTA-blood and serum samples were collected from 139 equids (90 donkeys, 39 horses and 10 mules), sampled by convenience. Among sampled equids, 23 animals were abandoned donkeys (17 females and 6 males) caught by the technical staff from the prefecture of São Luís city and kept in Universidade Estadual do Maranhão (UEMA). Additionally, 116 equids, including 67 donkeys (37 females and 20 males), 39 horses (11 females and 28 males), and 10 mules (5 females and 5 males) were working animals belonging to coachman. All equids sampled were adult animals, whose age ranged from five to eight years old.
DNA extraction
DNA was extracted from 200µL of each whole blood sample using the QIAamp DNA Blood Mini kit (QIAGEN, Valencia, California, USA), according to the manufacturer’s instructions.
Molecular detection of equid piroplasmids
Each sample of extracted DNA was used as template in nested PCR assays, using previously described specific protocols based on ema-1 gene for T. equi (NICOLAIEWSKY et al., 2001Nicolaiewsky TB, Richter MF, Lunge VR, Cunha CW, Delagostin O, Ikuta N, et al. Detection of Babesia equi (Laveran, 1901) by nested polymerase chain reaction. Vet Parasitol 2001; 101(1): 9-21. PMid:11587829. http://dx.doi.org/10.1016/S0304-4017(01)00471-X.
http://dx.doi.org/10.1016/S0304-4017(01)...
) and rap-1 gene for B. caballi (IKADAI et al., 1999Ikadai H, Xuan X, Igarashi I, Tanaka S, Kanemaru T, Nagasawa H, et al. Cloning and expression of a 48-kilodalton Babesia caballi merozoite rhoptry protein and potential use of the recombinant antigen in an enzyme-linked immunosorbent assay. J Clin Microbiol 1999; 37(11): 3475-3480. PMid:10523537.; BATTSETSEG et al., 2001Battsetseg B, Xuan X, Ikadai H, Bautista J, Byambaa B, Boldbaatar D, et al. Detection of Babesia caballi and Babesia equi in Dermacentor nuttalli adult ticks. Int J Parasitol 2001; 31(4): 384-386. PMid:11306116. http://dx.doi.org/10.1016/S0020-7519(01)00120-5.
http://dx.doi.org/10.1016/S0020-7519(01)...
). Briefly, five microliters of DNA was used as a template in 25 µL reaction mixtures containing 10X PCR buffer, 1.0 mM MgCl2, 0.6 mM deoxynucleotide triphosphate (dNTPs) mixture, 1.5 U of Taq DNA polymerase (Life Technologies) and 0.5 µM each primer (Integrated DNA Technologies) (Table 1). PCR amplifications were performed at 94°C for 4 min followed by 40 repetitive cycles of 94°C for 1 min, 56°C for 1 min for B. caballi and 60°C for 1 min for T. equi, and 72°C for 1 min, followed by a final extension at 72°C for 5 min.
Primers and annealing temperatures used for amplification of equine piroplasmids rap-1, ema-1 and 18S rRNA genes.
Molecular characterization of equid piroplasmids
In order to perform the molecular characterization of the equid piroplasmids, eleven positive samples (seven for T. equi and four for B. caballi) were randomly selected and submitted to previously described conventional PCR protocols for T. equi and B. caballi based on 18S rRNA (~1600 bp) (BHOORA et al., 2009Bhoora R, Franssen L, Oosthuizen MC, Guthrie AJ, Zweygarth E, Penzhorn BL, et al. Sequence heterogeneity in the 18S rRNA gene within Theileria equi and Babesia caballi from horses in South Africa. Vet Parasitol 2009; 159(2): 112-120. PMid:19019541. http://dx.doi.org/10.1016/j.vetpar.2008.10.004.
http://dx.doi.org/10.1016/j.vetpar.2008....
) (Table 1). The primers namely NBabesia1F: 5'-AAGCCATGCATGTCTAAGTATAAGCTTTT-3' and 18SRev-TB: 5'- GAATAATTCACCGGATCACTCG-3' were initially used for this purpose. The reactions were performed using five microliters of DNA as a template in 25 µL reaction mixtures containing 10X PCR buffer, 1.0 mM MgCl2, 0.8 mM deoxynucleotide triphosphate (dNTPs) mixture, 1.0 U Taq DNA Polymerase (Life Technologies, Carlsbad, CA, USA) and 0.5 µM of each primer (Integrated DNA Technologies). PCR amplifications were performed by an initial cycle of 94 °C for 2 min followed by 40 cycles of 94 °C for 30 s, 60 °C for 45 s and 72 °C for 1 min, and an extension final by 72 °C for 7 min. Additionally, DNA samples were submitted to another three nested PCR assays in order to achieve the whole 18S rRNA gene fragment of T. equi and B. caballi. The amplifications conditions were the same previously described, except the primers and annealing temperature (Table 1). T. equi and B. caballi DNA samples obtained from naturally infected donkeys (MACHADO et al., 2012Machado RZ, Toledo CZ, Teixeira MC, André MR, Freschi CR, Sampaio PH. Molecular and serological detection of Theileria equi and Babesia caballi in donkeys (Equus asinus) in Brazil. Vet Parasitol 2012; 186(3-4): 461-465. PMid:22186194. http://dx.doi.org/10.1016/j.vetpar.2011.11.069.
http://dx.doi.org/10.1016/j.vetpar.2011....
) and ultra-pure sterile water were used as positive and negative controls, respectively, in all PCR assays described above. In order to avoid the PCR contamination, DNA extraction, reaction setup, PCR amplification and electrophoresis were performed in separated rooms. Ethidium-bromide stained gels were imaged under ultraviolet light using the Image Lab Software version 4.1 (Bio-Rad®).
Sequencing and phylogenetic analyzes of equine piroplasm
18S rRNA-amplicons obtained were purified using Silica Bead DNA Gel Extraction Kit (Fermentas, São Paulo, SP, Brazil). The purified PCR products were subsequently sequenced in both directions using the same PCR primers (forward and reverse). Sanger sequencing at CREBIO (Centro de Recursos Biológicos e Biologia Genômica – FCAV/UNESP) was performed using the ABI Prism 310 Genetic Analyser–Applied Byosystem/Perkin Elmer. Consensus sequences were obtained through the analysis of the sense and antisense sequences using the CAP3 program (HUANG & MADAN, 1999Huang X, Madan A. CAP3: A DNA sequence assembly program. Genome Res 1999; 9(9): 868-877. PMID: 10508846.). Comparisons with sequences deposited in GenBank were done using the basic local alignment search tool (BLASTn – using default parameters) (ALTSCHUL et al., 1990Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol 1990; 215(3): 403-410. PMid:2231712. http://dx.doi.org/10.1016/S0022-2836(05)80360-2.
http://dx.doi.org/10.1016/S0022-2836(05)...
). The 18S rRNA sequences were aligned with sequences published in GenBank using Clustal/W (THOMPSON et al., 1994Thompson JD, Higgins DG, Gibson TJ. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 1994; 22(22): 4673-4680. PMid:7984417. http://dx.doi.org/10.1093/nar/22.22.4673.
http://dx.doi.org/10.1093/nar/22.22.4673...
) and adjusted in Bioedit v.7.0.5.3 (HALL, 1999Hall TA. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 1999; 41: 95-98.). Phylogenetic inference based on Maximum Likelihood criterion (ML) was inferred with RAxML-HPC BlackBox 7.6.3 (STAMATAKIS et al., 2008Stamatakis A, Hoover P, Rougemont J, Renner S. A rapid bootstrap algorithm for the RAxML Web servers. Syst Biol 2008; 57(5): 758-771. PMid:18853362. http://dx.doi.org/10.1080/10635150802429642.
http://dx.doi.org/10.1080/10635150802429...
) through the CIPRES Science Gateway (MILLER et al., 2011Miller MA, Pfeiffer W, Schwartz T. The CIPRES science gateway: a community resource for phylogenetic analyses [online]. New Orleans: CIPRES; 2011 [cited 2017 Feb 5]. Available from: https://www.phylo.org
https://www.phylo.org...
). Akaike information criterion (AIC) was used in Mega 5.05 to identify the most appropriate model of nucleotide substitution. GTRGAMMA + I model was chosen as the most appropriate for the Maximum Likelihood analysis of the 18S rRNA alignment. The sequences amplified in the present study were deposited in GenBank database under accession numbers: KY952226-KY952236.
Enzyme-linked immunosorbent assay (ELISA)
In order to detect IgG antibodies to T. equi and B. caballi, the serum samples were submitted to ELISA assays using commercial kits (Imunodot diagnóstico, IMUNOTEST®), in according to the manufacturer’s instructions. Briefly, 100µL of crude T. equi and B. caballi recombinant antigen (concentration of 10 µg/mL and 2.5 µg/mL, respectively), were diluted in a 0.05M sodium bicarbonate-carbonated buffer (pH 9.6 and added to each well of a micro-ELISA plate (NunclonTM Surface; Nunc, Denmark). After incubation at 4°C during 18 hours, the excess of antigen was removed by washing with PBS Tween-80 at 0.05%. Blocking of the uncoated sites in the well was achieved by the addition of 200 µL of PBS Tween-80 containing 6% skim milk for 2 h at 37°C. The blocking agent was removed, the plate was washed with PBS Tween-80 at 0.05%, and individual equids sera diluted (1:100) in PBS Tween-80 with 5% skim milk (PBS-TWSM) were added to each well and then incubated for 90 min at 37°C. After washing the plates, 100 µL of alkaline phosphatase conjugated anti-horse IgG (Sigma–Aldrich, St. Louis, Missouri, USA) diluted 1:30.000 in PBS-TW-SM were added to each well and then incubated for 90 min at 37°C. The plates were washed and the appropriate substrate (p-Nitrophenyl phosphate) (Sigma–Aldrich, St. Louis, Missouri, USA) was added. Serum samples of previously positive and negative donkeys were used as controls at ELISA assays (MACHADO et al., 2012Machado RZ, Toledo CZ, Teixeira MC, André MR, Freschi CR, Sampaio PH. Molecular and serological detection of Theileria equi and Babesia caballi in donkeys (Equus asinus) in Brazil. Vet Parasitol 2012; 186(3-4): 461-465. PMid:22186194. http://dx.doi.org/10.1016/j.vetpar.2011.11.069.
http://dx.doi.org/10.1016/j.vetpar.2011....
). Absorbance at 405nm and were read after 45 minutes incubation at room temperature using an ELISA reader (Dynex Technologies). The cut-off value (0.343 and 0.346, for T. equi and B. caballi, respectively) were determined as being 2.5 times the mean absorbance value of negative controls, where readings above the cutoff value were considered positive, according to Machado et al. (1997)Machado RZ, Montassier HJ, Pinto AA, Lemos EG, Machado MR, Valadão IF, et al. An enzyme-linked immunosorbent assay (ELISA) for the detection of antibodies against Babesia bovis in cattle. Vet Parasitol 1997; 71(1): 17-26. PMid:9231985. http://dx.doi.org/10.1016/S0304-4017(97)00003-4.
http://dx.doi.org/10.1016/S0304-4017(97)...
.
Results
Molecular and serological occurrence for equids piroplasmids
Among all equids’ blood samples analyzed, 21.6% (30/139) and 55.4% (77/139) were positive for cPCR assays for T. equi and B. caballi, respectively (Table 2). Additionally, co-positivity in PCR assays for both piroplasmid species was observed in 13.6% (19/139) of animals sampled (Table 3). From 139 serum samples submitted to ELISA assay, IgG antibodies to T. equi and B. caballi were detected in 19.4% (27/139) and 25.2% (35/139), respectively (Table 2). Only two animals (1.4%) showed antibodies for both T. equi and B. cabalii (Table 3). Fourteen (10%) animals were positive for both T. equi-ELISA and cPCR assays. Lastly, 17 (12.2%) animals were positive for both B. caballi-ELISA and cPCR assays. The occurrence of DNA or antibodies anti-piroplasms among the different equids species is shown in the Table 2.
Phylogenetic analysis
Out of the 30 positive samples for T. equi, seven (23.3%) 18S rRNA amplicons were randomly selected and submitted to sequencing and ML analyses. These sequences, when analyzed by BLASTn (using default parameters), shared identity ranging from 99% to 100% with T. equi sequences previously detected in Spain (AY150062), USA (CP001669 – whole genome) and Brazil (KU240071). Among the 77 positive samples for B. caballi, four (5.2%) 18S rRNA sequences were sequenced. The BLASTn analyses reveled that all sequences shared 99% identity with B. caballi (AY534883) amplified from an equine blood sample from Spain. All sequences amplified in the present study showed query coverage ranging from 99% to 100%.
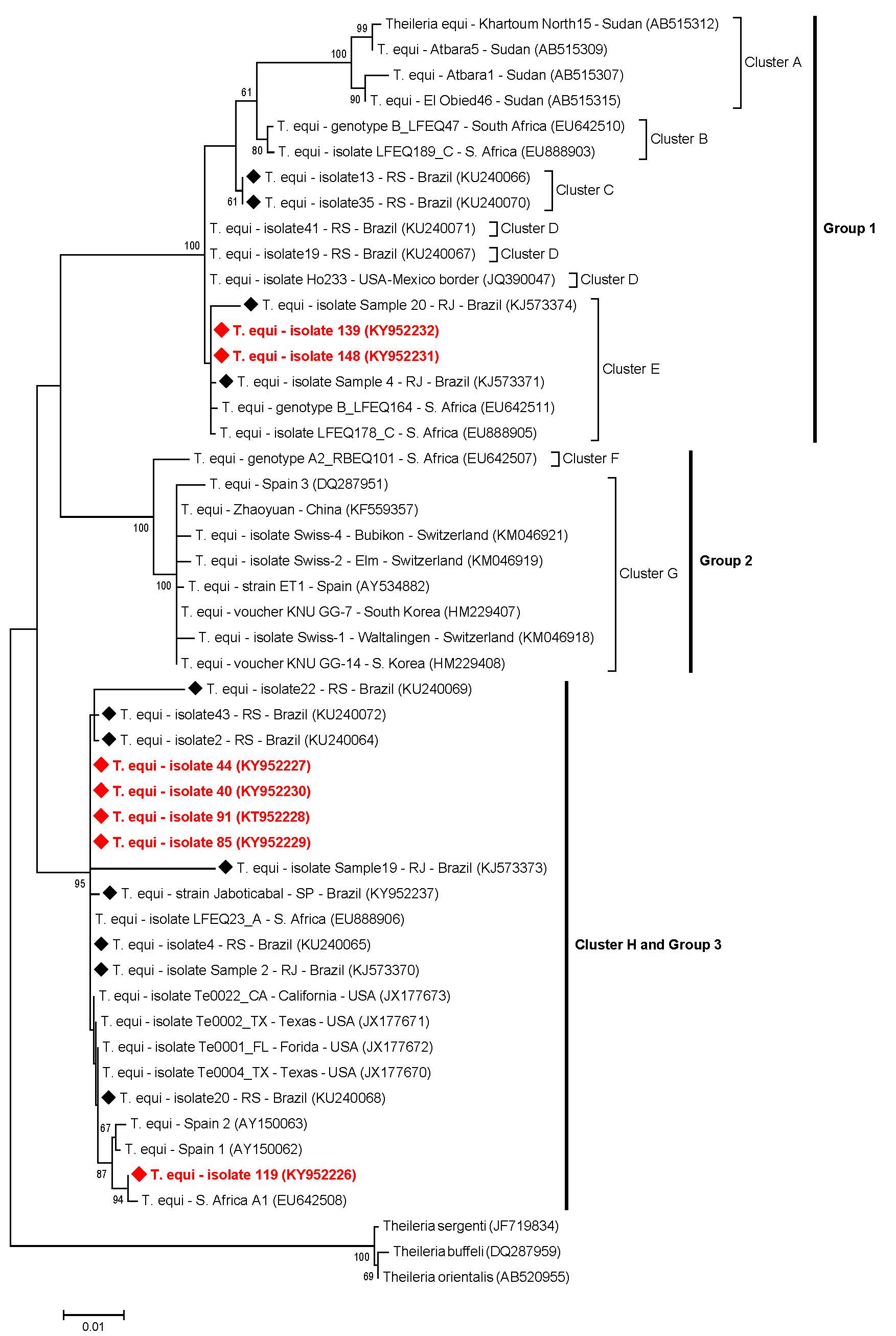
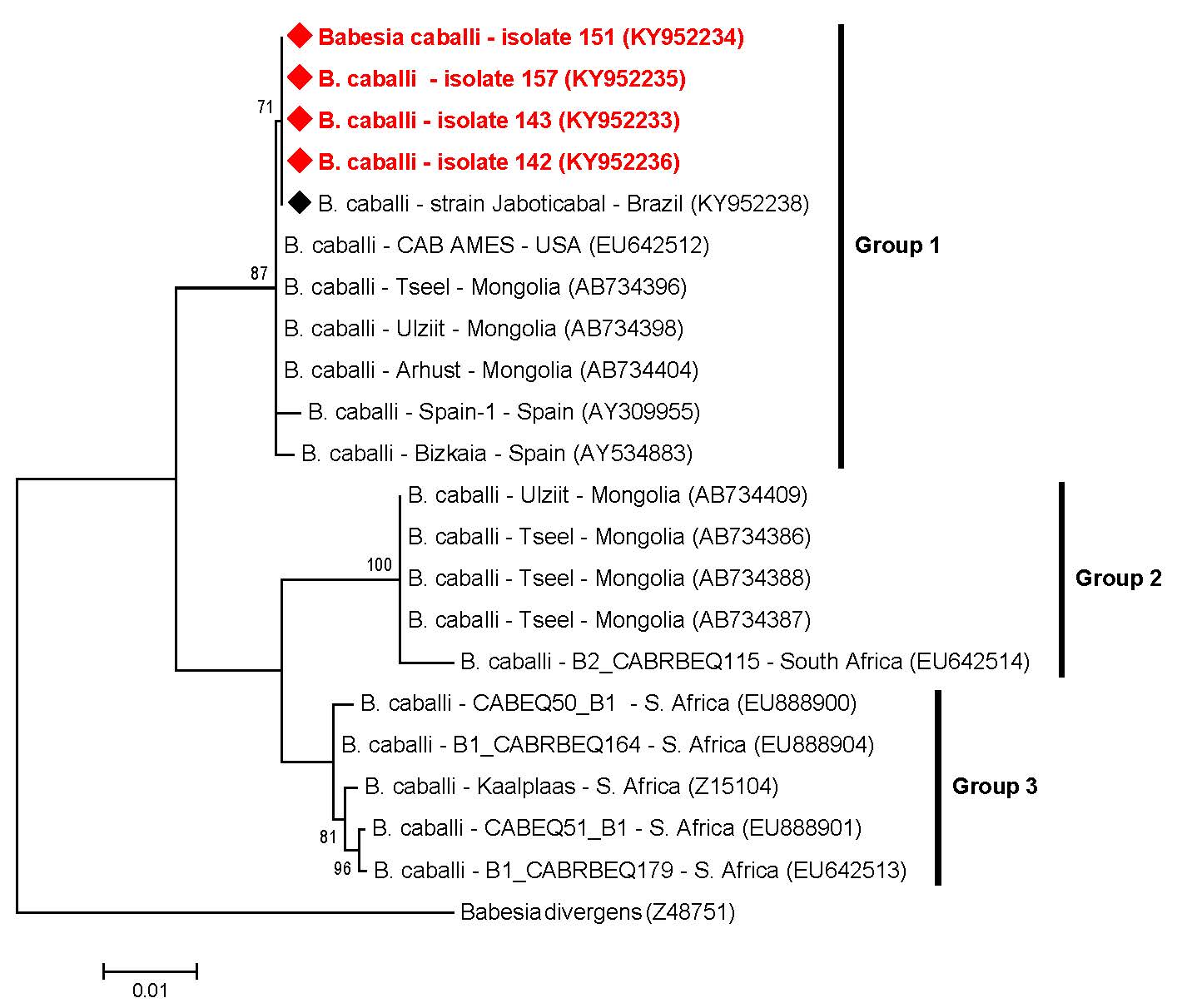
Overall, the T. equi sequences (n=7) submitted to ML analysis (based on a fragment of 1700 bp after alignment) grouped in three main groups (#1, #2 and #3), which were subdivided in eight clusters (A-H) (Figure 2). Among the seven sequences amplified, two of them detected in donkeys (#139 and #148) were positioned near to sequences previously detected in horses from Rio de Janeiro, southeastern Brazil, and to other two sequences detected in horses from South Africa, constituting the group #1 and cluster E. Five sequences (#40, #44, #85 and #91, detected in horses, and #119, detected in a mule), belonging to group #3 and cluster H, were positioned near to sequences previously identified in equids from Brazil (south and southeastern regions), USA, Spain and South Africa (Figure 2). Theileria equi genotypes belonging to group #2 were not represented by any of the sequences from our study. Additionally, four B. caballi sequences (#142, #143, #151 and #157), all detected in donkeys, submitted to ML analysis (based on a fragment of 1720 bp after alignment), were positioned in the same group (group #1) and nearest to B. caballi sequences from Brazil, USA, Mongolia and Spain (Figure 3).
Phylogenetic relationships within the T. equi sequences based on a 1700 bp fragment of the 18S rRNA gene. The phylogenetic tree was inferred by using the maximum likelihood method. The sequences detected in the present study are highlighted in red. The numbers at the nodes correspond to boostrap values higher than 60% accessed with 1.000 replicates. Theileria species was used as outgroup.
Phylogenetic relationships within the B. caballi sequences based on a 1720 bp fragment of the 18S rRNA gene. The phylogenetic tree was inferred by using the maximum likelihood method. The sequences detected in the present study are highlighted in red. The numbers at the nodes correspond to boostrap values higher than 60% accessed with 1.000 replicates. Babesia divergens was used as outgroup.
Discussion
In the present study, the frequency of piroplasm infection in donkeys, horses and mules from São Luís Island, Maranhão state, northeastern Brazil, was investigated by molecular and serological techniques. Both T. equi and B. caballi circulate in equids sampled in the studied area.
The low percentage of positive donkeys for piroplasmids found in the present study corroborates with a previous work carried out in the state of São Paulo, southeastern Brazil (MACHADO et al., 2012Machado RZ, Toledo CZ, Teixeira MC, André MR, Freschi CR, Sampaio PH. Molecular and serological detection of Theileria equi and Babesia caballi in donkeys (Equus asinus) in Brazil. Vet Parasitol 2012; 186(3-4): 461-465. PMid:22186194. http://dx.doi.org/10.1016/j.vetpar.2011.11.069.
http://dx.doi.org/10.1016/j.vetpar.2011....
), suggesting low levels of transmission in this animal species in both studied areas.
Herein, we found an overall low seroprevalence for T. equi (19.4%) and B. caballi (25.2%) among sampled equids. Horses showed a higher seroprevalence to T. equi when compared to that found among donkeys and mules. On the other hand, donkeys and mules showed a higher seroprevalence to B. caballi when compared to that found among sampled horses. These seroprevalence rates were lower than those found among donkeys sampled in São Paulo state for both T. equi and B. caballi (MACHADO et al., 2012Machado RZ, Toledo CZ, Teixeira MC, André MR, Freschi CR, Sampaio PH. Molecular and serological detection of Theileria equi and Babesia caballi in donkeys (Equus asinus) in Brazil. Vet Parasitol 2012; 186(3-4): 461-465. PMid:22186194. http://dx.doi.org/10.1016/j.vetpar.2011.11.069.
http://dx.doi.org/10.1016/j.vetpar.2011....
; PIANTEDOSI et al., 2014Piantedosi D, D’Alessio N, Di Loria A, Di Prisco F, Mariani U, Neola B, et al. Seroprevalence and risk factors associated with Babesia caballi and Theileria equi infections in donkeys from Southern Italy. Vet J 2014; 202(3): 578-582. PMid:25457263. http://dx.doi.org/10.1016/j.tvjl.2014.09.025.
http://dx.doi.org/10.1016/j.tvjl.2014.09...
; LAUS et al., 2015Laus F, Spaterna A, Faillace V, Veronesi F, Ravagnan S, Beribé F, et al. Clinical investigation on Theileria equi and Babesia caballi infections in Italian donkeys. BMC Vet Res 2015; 11(1): 100. PMid:25927984. http://dx.doi.org/10.1186/s12917-015-0411-z.
http://dx.doi.org/10.1186/s12917-015-041...
) and in horses from southeastern (BALDANI et al., 2010Baldani CD, Nakaghi ACH, Machado RZ. Occurrence of Theileria equi in horses raised in the Jaboticabal microregion, São Paulo state, Brazil. Rev Bras Parasitol Vet 2010; 19(4): 228-232. PMid:21184699. http://dx.doi.org/10.1590/S1984-29612010000400007.
http://dx.doi.org/10.1590/S1984-29612010...
; SALVAGNI et al., 2010Salvagni CA, Dagnone AS, Gomes TS, Mota JS, Andrade GM, Baldani CD, et al. Serologic evidence of equine granulocytic anaplasmosis in horses from central West Brazil. Rev Bras Parasitol Vet 2010; 19(3): 135-140. PMid:20943015. http://dx.doi.org/10.1590/S1984-29612010000300002.
http://dx.doi.org/10.1590/S1984-29612010...
; PFEIFER BARBOSA et al., 1995Pfeifer Barbosa I, Böse R, Peymann B, Friedhoff KT. Epidemiological aspects of equine babesioses in a herd of horses in Brazil. Vet Parasitol 1995; 58(1-2): 1-8. PMid:7676590. http://dx.doi.org/10.1016/0304-4017(94)00704-G.
http://dx.doi.org/10.1016/0304-4017(94)0...
; HEUCHERT et al., 1999Heuchert CM, Giulli V Jr, Athaide D, Böse R, Friedhoff KT. Seroepidemiologic studies on Babesia equi and Babesia caballi infections in Brazil. Vet Parasitol 1999; 85(1): 1-11. PMid:10447188. http://dx.doi.org/10.1016/S0304-4017(99)00108-9.
http://dx.doi.org/10.1016/S0304-4017(99)...
; HEIM et al., 2007Heim A, Passos LM, Ribeiro MF, Costa-Júnior LM, Bastos CV, Cabral DD, et al. Detection and molecular characterization of Babesia caballi and Theileria equi isolates from endemic areas of Brazil. Parasitol Res 2007; 102(1): 63-68. PMid:17828553. http://dx.doi.org/10.1007/s00436-007-0726-1.
http://dx.doi.org/10.1007/s00436-007-072...
; KERBER et al., 2009Kerber CE, Labruna MB, Ferreira F, De Waal DT, Knowles DP, Gennari SM. Prevalence of equine Piroplasmosis and its association with tick infestation in the State of São Paulo, Brazil. Rev Bras Parasitol Vet 2009; 18(4): 1-8. PMid:20040201. http://dx.doi.org/10.4322/rbpv.01804001.
http://dx.doi.org/10.4322/rbpv.01804001...
) and southern regions (VIEIRA et al., 2013Vieira TS, Vieira RF, Finger MA, Nascimento DA, Sicupira PM, Dutra LH, et al. Seroepidemiological survey of Theileria equi and Babesia caballi in horses from a rural and from urban areas of Paraná State, southern Brazil. Ticks Tick Borne Dis 2013; 4(6): 537-541. PMid:24209493. http://dx.doi.org/10.1016/j.ttbdis.2013.07.005.
http://dx.doi.org/10.1016/j.ttbdis.2013....
) of Brazil.
Previously, the occurrence of antibodies for B. caballi was more prevalent than that one found for T. equi in horses parasitized by Dermacentor nitens, Amblyomma cajennense and Rhipicephalus (Boophilus) microplus ticks (KERBER et al., 2009Kerber CE, Labruna MB, Ferreira F, De Waal DT, Knowles DP, Gennari SM. Prevalence of equine Piroplasmosis and its association with tick infestation in the State of São Paulo, Brazil. Rev Bras Parasitol Vet 2009; 18(4): 1-8. PMid:20040201. http://dx.doi.org/10.4322/rbpv.01804001.
http://dx.doi.org/10.4322/rbpv.01804001...
). Additionally, while the parasitism by D. nitens was statistically associated with B. caballi positivity, the presence of Amblyomma cajennense in horses was associated with infection and/or exposure to T. equi (KERBER et al., 2009Kerber CE, Labruna MB, Ferreira F, De Waal DT, Knowles DP, Gennari SM. Prevalence of equine Piroplasmosis and its association with tick infestation in the State of São Paulo, Brazil. Rev Bras Parasitol Vet 2009; 18(4): 1-8. PMid:20040201. http://dx.doi.org/10.4322/rbpv.01804001.
http://dx.doi.org/10.4322/rbpv.01804001...
). In fact, the intrastadial transmission of T. equi by A. cajennense male ticks has been confirmed between uninfected and infected horses in the USA (SCOLES & UETI, 2013Scoles GA, Ueti MW. Amblyomma cajennense is an intrastadial biological vector of Theileria equi. Parasit Vectors 2013; 6(1): 306. PMid:24499587. http://dx.doi.org/10.1186/1756-3305-6-306.
http://dx.doi.org/10.1186/1756-3305-6-30...
). Unfortunately, an accurate assessment of tick infestation on sampled equids was not performed, precluding any inferences about tick species found in animals and positivity for T. equi and B. caballi. Future studies aiming at assessing the vectors involved in piroplasmids transmission among equids in northeastern Brazil are much needed.
The occurrence of T. equi DNA in donkey blood samples in the present study was lower than that found among horses and mules. On the other hand, the occurrence of B. caballi DNA in donkey blood samples were higher than that found among horses and mules. While the number of seropositive animals for T. equi and B. caballi was lower than the number of PCR positive animals in the present study, the opposite was reported by Machado et al. (2012)Machado RZ, Toledo CZ, Teixeira MC, André MR, Freschi CR, Sampaio PH. Molecular and serological detection of Theileria equi and Babesia caballi in donkeys (Equus asinus) in Brazil. Vet Parasitol 2012; 186(3-4): 461-465. PMid:22186194. http://dx.doi.org/10.1016/j.vetpar.2011.11.069.
http://dx.doi.org/10.1016/j.vetpar.2011....
among equids sampled in the state of São Paulo.
The 18S rRNA gene is widely used for phylogenetic analysis of equine piroplasms (BHOORA et al., 2009Bhoora R, Franssen L, Oosthuizen MC, Guthrie AJ, Zweygarth E, Penzhorn BL, et al. Sequence heterogeneity in the 18S rRNA gene within Theileria equi and Babesia caballi from horses in South Africa. Vet Parasitol 2009; 159(2): 112-120. PMid:19019541. http://dx.doi.org/10.1016/j.vetpar.2008.10.004.
http://dx.doi.org/10.1016/j.vetpar.2008....
; HALL et al., 2013Hall CM, Busch JD, Scoles GA, Palma-Cagle KA, Ueti MW, Kappmeyer LS, et al. Genetic characterization of Theileria equi infecting horses in North America: evidence for a limited source of US introductions. Parasit Vectors 2013; 6(1): 35. PMid:23399005. http://dx.doi.org/10.1186/1756-3305-6-35.
http://dx.doi.org/10.1186/1756-3305-6-35...
; MUNKHJARGAL et al., 2013Munkhjargal T, Sivakumar T, Battsetseg B, Nyamjargal T, Aboulaila M, Purevtseren B, et al. Prevalence and genetic diversity of equine piroplasms in Tov province, Mongolia. Infect Genet Evol 2013; 16: 178-185. PMid:23416256. http://dx.doi.org/10.1016/j.meegid.2013.02.005.
http://dx.doi.org/10.1016/j.meegid.2013....
). Herein, the genetic composition and the phylogenetic positioning of T. equi and B. caballi sequences amplified among equids from São Luís, Maranhão state, Brazil was assessed.
In the present study, three distinct main groups were identified by phylogenetic analysis of T. equi and B. caballi 18S rRNA gene sequences. T. equi and B. caballi 18S rRNA sequences detected in equids from Brazil clustered within two and one groups, respectively. Although fewer B. caballi sequences were analyzed, less genetic variation within B. caballi sequences seemed to occur when compared to T. equi sequences. These findings are in agreement with previous reports (CRIADO-FORNELIO et al., 2004Criado-Fornelio A, Gónzalez-del-Río MA, Buling-Saraña A, Barba-Carretero JC. The “expanding universe” of piroplasms. Vet Parasitol 2004; 119(4): 337-345. PMid:15154598. http://dx.doi.org/10.1016/j.vetpar.2003.11.015.
http://dx.doi.org/10.1016/j.vetpar.2003....
; BHOORA et al., 2009Bhoora R, Franssen L, Oosthuizen MC, Guthrie AJ, Zweygarth E, Penzhorn BL, et al. Sequence heterogeneity in the 18S rRNA gene within Theileria equi and Babesia caballi from horses in South Africa. Vet Parasitol 2009; 159(2): 112-120. PMid:19019541. http://dx.doi.org/10.1016/j.vetpar.2008.10.004.
http://dx.doi.org/10.1016/j.vetpar.2008....
).
Although Heim et al. (2007)Heim A, Passos LM, Ribeiro MF, Costa-Júnior LM, Bastos CV, Cabral DD, et al. Detection and molecular characterization of Babesia caballi and Theileria equi isolates from endemic areas of Brazil. Parasitol Res 2007; 102(1): 63-68. PMid:17828553. http://dx.doi.org/10.1007/s00436-007-0726-1.
http://dx.doi.org/10.1007/s00436-007-072...
, when analyzing the T. equi ema-1 gene sequences (800bp) and B. caballi BC-48 gene sequences (700bp), have reported no differences within each species or among known sequences available and, therefore, no diversity was observed among the isolates, the results of the phylogenetic analysis performed in the present study showed a high genetic diversity among the T. equi-18S rRNA sequences amplified in equids from different Brazilian regions. Additionally, the findings also revealed that the genetic diversity of T. equi sequences seems to be not geographically delimited in the Brazil, as suggested by the positioning of some sequences amplified from different regions (Maranhão, Rio Grande do Sul, Rio de Janeiro and São Paulo states), which clustered all together.
The heterogeneity observed among the T. equi-18S rRNA sequences amplified in equids from Brazil is in agreement with previous studies performed in Spain (NAGORE et al., 2004Nagore D, García-Sanmartín J, Garcıa-Pérez AL, Juste RA, Hurtado A. Detection and identification of equine Theileria and Babesia species by reverse line blotting: epidemiological survey and phylogenetic analysis. Vet Parasitol 2004; 123(1-2): 41-54. PMid:15265570. http://dx.doi.org/10.1016/j.vetpar.2004.04.010.
http://dx.doi.org/10.1016/j.vetpar.2004....
), South Africa (BHOORA et al., 2009Bhoora R, Franssen L, Oosthuizen MC, Guthrie AJ, Zweygarth E, Penzhorn BL, et al. Sequence heterogeneity in the 18S rRNA gene within Theileria equi and Babesia caballi from horses in South Africa. Vet Parasitol 2009; 159(2): 112-120. PMid:19019541. http://dx.doi.org/10.1016/j.vetpar.2008.10.004.
http://dx.doi.org/10.1016/j.vetpar.2008....
), Greece (KOUAM et al., 2010Kouam MK, Kantzoura V, Gajadhar AA, Theis JH, Papadopoulos E, Theodoropoulos G. Seroprevalence of equine piroplasms and host-related factors associated with infection in Greece. Vet Parasitol 2010; 169(3-4): 273-278. PMid:20138434. http://dx.doi.org/10.1016/j.vetpar.2010.01.011.
http://dx.doi.org/10.1016/j.vetpar.2010....
), Mongolia (MUNKHJARGAL et al., 2013Munkhjargal T, Sivakumar T, Battsetseg B, Nyamjargal T, Aboulaila M, Purevtseren B, et al. Prevalence and genetic diversity of equine piroplasms in Tov province, Mongolia. Infect Genet Evol 2013; 16: 178-185. PMid:23416256. http://dx.doi.org/10.1016/j.meegid.2013.02.005.
http://dx.doi.org/10.1016/j.meegid.2013....
), Tunisia (ROS-GARCÍA et al., 2013Ros-García A, M’ghirbi Y, Hurtado A, Bouattour A. Prevalence and genetic diversity of piroplasm species in horses and ticks from Tunisia. Infect Genet Evol 2013; 17: 33-37. PMid:23542456. http://dx.doi.org/10.1016/j.meegid.2013.03.038.
http://dx.doi.org/10.1016/j.meegid.2013....
) and Israel (KETTER-RATZON et al., 2017Ketter-Ratzon D, Tirosh-Levy S, Nachum-Biala Y, Saar T, Qura’n L, Zivotofsky D, et al. Characterization of Theileria equi genotypes in horses in Israel, the Palestinian Authority and Jordan. Ticks Tick Borne Dis 2017; 8(4): 499-505. PMid:28286144. http://dx.doi.org/10.1016/j.ttbdis.2017.02.010.
http://dx.doi.org/10.1016/j.ttbdis.2017....
). On the other hand, Hall et al. (2013)Hall CM, Busch JD, Scoles GA, Palma-Cagle KA, Ueti MW, Kappmeyer LS, et al. Genetic characterization of Theileria equi infecting horses in North America: evidence for a limited source of US introductions. Parasit Vectors 2013; 6(1): 35. PMid:23399005. http://dx.doi.org/10.1186/1756-3305-6-35.
http://dx.doi.org/10.1186/1756-3305-6-35...
showed that T. equi-18S rRNA sequences from different USA regions clustered in a single cluster, exhibiting very little diversity within this group, suggesting a limited introduction of a small number of T. equi genotypes into the USA (HALL et al., 2013Hall CM, Busch JD, Scoles GA, Palma-Cagle KA, Ueti MW, Kappmeyer LS, et al. Genetic characterization of Theileria equi infecting horses in North America: evidence for a limited source of US introductions. Parasit Vectors 2013; 6(1): 35. PMid:23399005. http://dx.doi.org/10.1186/1756-3305-6-35.
http://dx.doi.org/10.1186/1756-3305-6-35...
).
Even though it is not possible to use the 18S rRNA gene sequences diversity to classify T. equi as different species (CRIADO-FORNELIO et al., 2003Criado-Fornelio A, Martinez-Marcos A, Buling-Saraña A, Barba-Carretero JC. Molecular studies on Babesia, Theileria and Hepatozoon in southern Europe Part I. Epizootiological aspects. Vet Parasitol 2003; 113(3-4): 189-201. PMid:12719133. http://dx.doi.org/10.1016/S0304-4017(03)00078-5.
http://dx.doi.org/10.1016/S0304-4017(03)...
; NAGORE et al., 2004Nagore D, García-Sanmartín J, Garcıa-Pérez AL, Juste RA, Hurtado A. Detection and identification of equine Theileria and Babesia species by reverse line blotting: epidemiological survey and phylogenetic analysis. Vet Parasitol 2004; 123(1-2): 41-54. PMid:15265570. http://dx.doi.org/10.1016/j.vetpar.2004.04.010.
http://dx.doi.org/10.1016/j.vetpar.2004....
), it is extremely necessary that future studies evaluate the pathogenic potential of these different genotypes infecting equids around the world.
Conclusion
The present work showed that different genotypes of T. equi and B. caballi circulate among equids in Brazil. Future studies aiming at elucidating the differences in pathogenic potential of different piroplasmids genotypes are much needed. Additionally, the phylogenetic positioning of these genotypes according to mitochondrial genes will contribute to the elucidation of the taxonomy of Piroplasmida.
References
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol 1990; 215(3): 403-410. PMid:2231712. http://dx.doi.org/10.1016/S0022-2836(05)80360-2
» http://dx.doi.org/10.1016/S0022-2836(05)80360-2 - Baldani CD, Nakaghi ACH, Machado RZ. Occurrence of Theileria equi in horses raised in the Jaboticabal microregion, São Paulo state, Brazil. Rev Bras Parasitol Vet 2010; 19(4): 228-232. PMid:21184699. http://dx.doi.org/10.1590/S1984-29612010000400007
» http://dx.doi.org/10.1590/S1984-29612010000400007 - Bashiruddin JB, Cammà C, Rebêlo E. Molecular detection of Babesia equi and Babesia caballi in horse blood by PCR amplification of part of the 16S RNA gene. Vet Parasitol 1999; 84(1-2): 75-83. PMid:10435792. http://dx.doi.org/10.1016/S0304-4017(99)00049-7
» http://dx.doi.org/10.1016/S0304-4017(99)00049-7 - Battsetseg B, Lucero S, Xuan X, Claveria FG, Inoue N, Alhassan A, et al. Detection of natural infection of Boophilus microplus with Babesia equi and Babesia caballi in Brazilian horses using nested polymerase chain reaction. Vet Parasitol 2002; 107(4): 351-357. PMid:12163246. http://dx.doi.org/10.1016/S0304-4017(02)00131-0
» http://dx.doi.org/10.1016/S0304-4017(02)00131-0 - Battsetseg B, Xuan X, Ikadai H, Bautista J, Byambaa B, Boldbaatar D, et al. Detection of Babesia caballi and Babesia equi in Dermacentor nuttalli adult ticks. Int J Parasitol 2001; 31(4): 384-386. PMid:11306116. http://dx.doi.org/10.1016/S0020-7519(01)00120-5
» http://dx.doi.org/10.1016/S0020-7519(01)00120-5 - Bhoora R, Franssen L, Oosthuizen MC, Guthrie AJ, Zweygarth E, Penzhorn BL, et al. Sequence heterogeneity in the 18S rRNA gene within Theileria equi and Babesia caballi from horses in South Africa. Vet Parasitol 2009; 159(2): 112-120. PMid:19019541. http://dx.doi.org/10.1016/j.vetpar.2008.10.004
» http://dx.doi.org/10.1016/j.vetpar.2008.10.004 - Bhoora R, Quan M, Franssen L, Butler CM, Van der Kolk JH, Guthrie AJ, et al. Development and evaluation of real-time PCR assays for the quantitative detection of Babesia caballi and Theileria equi infections in horses from South Africa. Vet Parasitol 2010; 168(3-4): 201-211. PMid:20031328. http://dx.doi.org/10.1016/j.vetpar.2009.11.011
» http://dx.doi.org/10.1016/j.vetpar.2009.11.011 - Böse R, Jorgensen WK, Dalgliesh RJ, Friedhoff KT, de Vos AJ. Current state and future trends in the diagnosis of babesiosis. Vet Parasitol 1995; 57(1-3): 61-74. PMid:7597794. http://dx.doi.org/10.1016/0304-4017(94)03111-9
» http://dx.doi.org/10.1016/0304-4017(94)03111-9 - Brüning A. Equine piroplasmosis an update on diagnosis, treatment and prevention. Br Vet J 1996; 152(2): 139-151. PMid:8680838. http://dx.doi.org/10.1016/S0007-1935(96)80070-4
» http://dx.doi.org/10.1016/S0007-1935(96)80070-4 - Carcy B, Précigout E, Schetters T, Gorenflot A. Genetic basis for GPI-anchor merozoite surface antigen polymorphism of Babesia and resulting antigenic diversity. Vet Parasitol 2006; 138(1-2): 33-49. PMid:16551492. http://dx.doi.org/10.1016/j.vetpar.2006.01.038
» http://dx.doi.org/10.1016/j.vetpar.2006.01.038 - Criado-Fornelio A, Gónzalez-del-Río MA, Buling-Saraña A, Barba-Carretero JC. The “expanding universe” of piroplasms. Vet Parasitol 2004; 119(4): 337-345. PMid:15154598. http://dx.doi.org/10.1016/j.vetpar.2003.11.015
» http://dx.doi.org/10.1016/j.vetpar.2003.11.015 - Criado-Fornelio A, Martinez-Marcos A, Buling-Saraña A, Barba-Carretero JC. Molecular studies on Babesia, Theileria and Hepatozoon in southern Europe Part I. Epizootiological aspects. Vet Parasitol 2003; 113(3-4): 189-201. PMid:12719133. http://dx.doi.org/10.1016/S0304-4017(03)00078-5
» http://dx.doi.org/10.1016/S0304-4017(03)00078-5 - de Waal DT. Equine piroplasmosis: a review. Br Vet J 1992; 148(1): 6-14. PMid:1551016. http://dx.doi.org/10.1016/0007-1935(92)90061-5
» http://dx.doi.org/10.1016/0007-1935(92)90061-5 - Golynski AA, Fernandes KR, Baldani CD, Golynski AL, Madeiro AS, Machado RZ, et al. Seroepidemiological studies on Babesia equi in horses from the State of Rio Grande do Sul determined by indirect immunoflourecence test and Elisa. Rev Bras Parasitol Vet 2008;17(Suppl 1): 317-321. PMid:20059868.
- Guimarães AM, Lima JD, Ribeiro MFB. Sporogony and experimental transmission of Babesia equi by Boophilus microplus. Parasitol Res 1998; 84(4): 323-327. PMid:9580425. http://dx.doi.org/10.1007/s004360050404
» http://dx.doi.org/10.1007/s004360050404 - Hall CM, Busch JD, Scoles GA, Palma-Cagle KA, Ueti MW, Kappmeyer LS, et al. Genetic characterization of Theileria equi infecting horses in North America: evidence for a limited source of US introductions. Parasit Vectors 2013; 6(1): 35. PMid:23399005. http://dx.doi.org/10.1186/1756-3305-6-35
» http://dx.doi.org/10.1186/1756-3305-6-35 - Hall TA. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 1999; 41: 95-98.
- Heim A, Passos LM, Ribeiro MF, Costa-Júnior LM, Bastos CV, Cabral DD, et al. Detection and molecular characterization of Babesia caballi and Theileria equi isolates from endemic areas of Brazil. Parasitol Res 2007; 102(1): 63-68. PMid:17828553. http://dx.doi.org/10.1007/s00436-007-0726-1
» http://dx.doi.org/10.1007/s00436-007-0726-1 - Heuchert CM, Giulli V Jr, Athaide D, Böse R, Friedhoff KT. Seroepidemiologic studies on Babesia equi and Babesia caballi infections in Brazil. Vet Parasitol 1999; 85(1): 1-11. PMid:10447188. http://dx.doi.org/10.1016/S0304-4017(99)00108-9
» http://dx.doi.org/10.1016/S0304-4017(99)00108-9 - Huang X, Madan A. CAP3: A DNA sequence assembly program. Genome Res 1999; 9(9): 868-877. PMID: 10508846.
- Ikadai H, Xuan X, Igarashi I, Tanaka S, Kanemaru T, Nagasawa H, et al. Cloning and expression of a 48-kilodalton Babesia caballi merozoite rhoptry protein and potential use of the recombinant antigen in an enzyme-linked immunosorbent assay. J Clin Microbiol 1999; 37(11): 3475-3480. PMid:10523537.
- Kerber CE, Labruna MB, Ferreira F, De Waal DT, Knowles DP, Gennari SM. Prevalence of equine Piroplasmosis and its association with tick infestation in the State of São Paulo, Brazil. Rev Bras Parasitol Vet 2009; 18(4): 1-8. PMid:20040201. http://dx.doi.org/10.4322/rbpv.01804001
» http://dx.doi.org/10.4322/rbpv.01804001 - Ketter-Ratzon D, Tirosh-Levy S, Nachum-Biala Y, Saar T, Qura’n L, Zivotofsky D, et al. Characterization of Theileria equi genotypes in horses in Israel, the Palestinian Authority and Jordan. Ticks Tick Borne Dis 2017; 8(4): 499-505. PMid:28286144. http://dx.doi.org/10.1016/j.ttbdis.2017.02.010
» http://dx.doi.org/10.1016/j.ttbdis.2017.02.010 - Kouam MK, Kantzoura V, Gajadhar AA, Theis JH, Papadopoulos E, Theodoropoulos G. Seroprevalence of equine piroplasms and host-related factors associated with infection in Greece. Vet Parasitol 2010; 169(3-4): 273-278. PMid:20138434. http://dx.doi.org/10.1016/j.vetpar.2010.01.011
» http://dx.doi.org/10.1016/j.vetpar.2010.01.011 - Kumar S, Kumar R, Sugimoto C. A perspective on Theileria equi infections in donkeys. Jpn J Vet Res 2009; 56(4): 171-180. PMid:19358444.
- Laus F, Spaterna A, Faillace V, Veronesi F, Ravagnan S, Beribé F, et al. Clinical investigation on Theileria equi and Babesia caballi infections in Italian donkeys. BMC Vet Res 2015; 11(1): 100. PMid:25927984. http://dx.doi.org/10.1186/s12917-015-0411-z
» http://dx.doi.org/10.1186/s12917-015-0411-z - Machado RZ, Montassier HJ, Pinto AA, Lemos EG, Machado MR, Valadão IF, et al. An enzyme-linked immunosorbent assay (ELISA) for the detection of antibodies against Babesia bovis in cattle. Vet Parasitol 1997; 71(1): 17-26. PMid:9231985. http://dx.doi.org/10.1016/S0304-4017(97)00003-4
» http://dx.doi.org/10.1016/S0304-4017(97)00003-4 - Machado RZ, Toledo CZ, Teixeira MC, André MR, Freschi CR, Sampaio PH. Molecular and serological detection of Theileria equi and Babesia caballi in donkeys (Equus asinus) in Brazil. Vet Parasitol 2012; 186(3-4): 461-465. PMid:22186194. http://dx.doi.org/10.1016/j.vetpar.2011.11.069
» http://dx.doi.org/10.1016/j.vetpar.2011.11.069 - Matjila PT, Leisewitz AL, Oosthuizen MC, Jongejan F, Penzhorn BL. Detection of a Theileria species in dogs in South Africa. Vet Parasitol 2008; 157(1-2): 34-40. PMid:18687528. http://dx.doi.org/10.1016/j.vetpar.2008.06.025
» http://dx.doi.org/10.1016/j.vetpar.2008.06.025 - Miller MA, Pfeiffer W, Schwartz T. The CIPRES science gateway: a community resource for phylogenetic analyses [online]. New Orleans: CIPRES; 2011 [cited 2017 Feb 5]. Available from: https://www.phylo.org
» https://www.phylo.org - Munkhjargal T, Sivakumar T, Battsetseg B, Nyamjargal T, Aboulaila M, Purevtseren B, et al. Prevalence and genetic diversity of equine piroplasms in Tov province, Mongolia. Infect Genet Evol 2013; 16: 178-185. PMid:23416256. http://dx.doi.org/10.1016/j.meegid.2013.02.005
» http://dx.doi.org/10.1016/j.meegid.2013.02.005 - Nagore D, García-Sanmartín J, Garcıa-Pérez AL, Juste RA, Hurtado A. Detection and identification of equine Theileria and Babesia species by reverse line blotting: epidemiological survey and phylogenetic analysis. Vet Parasitol 2004; 123(1-2): 41-54. PMid:15265570. http://dx.doi.org/10.1016/j.vetpar.2004.04.010
» http://dx.doi.org/10.1016/j.vetpar.2004.04.010 - Nicolaiewsky TB, Richter MF, Lunge VR, Cunha CW, Delagostin O, Ikuta N, et al. Detection of Babesia equi (Laveran, 1901) by nested polymerase chain reaction. Vet Parasitol 2001; 101(1): 9-21. PMid:11587829. http://dx.doi.org/10.1016/S0304-4017(01)00471-X
» http://dx.doi.org/10.1016/S0304-4017(01)00471-X - Oosthuizen MC, Zweygarth E, Collins NE, Troskie M, Penzhorn BL. Identification of a novel Babesia sp. from a sable antelope (Hippotragus niger Harris, 1838). J Clin Microbiol 2008; 46(7): 2247-2251. PMid:18508943. http://dx.doi.org/10.1128/JCM.00167-08
» http://dx.doi.org/10.1128/JCM.00167-08 - Pfeifer Barbosa I, Böse R, Peymann B, Friedhoff KT. Epidemiological aspects of equine babesioses in a herd of horses in Brazil. Vet Parasitol 1995; 58(1-2): 1-8. PMid:7676590. http://dx.doi.org/10.1016/0304-4017(94)00704-G
» http://dx.doi.org/10.1016/0304-4017(94)00704-G - Piantedosi D, D’Alessio N, Di Loria A, Di Prisco F, Mariani U, Neola B, et al. Seroprevalence and risk factors associated with Babesia caballi and Theileria equi infections in donkeys from Southern Italy. Vet J 2014; 202(3): 578-582. PMid:25457263. http://dx.doi.org/10.1016/j.tvjl.2014.09.025
» http://dx.doi.org/10.1016/j.tvjl.2014.09.025 - Ribeiro MFB, Costa JO, Guimarães AM. Epidemiological aspects of Babesia equi in Horses in Minas Gerais, Brazil. Vet Res Commun 1999; 23(6): 385-390. PMid:10543367. http://dx.doi.org/10.1023/A:1006389725759
» http://dx.doi.org/10.1023/A:1006389725759 - Ros-García A, M’ghirbi Y, Hurtado A, Bouattour A. Prevalence and genetic diversity of piroplasm species in horses and ticks from Tunisia. Infect Genet Evol 2013; 17: 33-37. PMid:23542456. http://dx.doi.org/10.1016/j.meegid.2013.03.038
» http://dx.doi.org/10.1016/j.meegid.2013.03.038 - Salvagni CA, Dagnone AS, Gomes TS, Mota JS, Andrade GM, Baldani CD, et al. Serologic evidence of equine granulocytic anaplasmosis in horses from central West Brazil. Rev Bras Parasitol Vet 2010; 19(3): 135-140. PMid:20943015. http://dx.doi.org/10.1590/S1984-29612010000300002
» http://dx.doi.org/10.1590/S1984-29612010000300002 - Schein E. Equine babesiosis. In: Ristic M, editors. Babesiosis of domestic animals and man. Boca Raton: CRS Press; 1988. p. 197-208.
- Scoles GA, Ueti MW. Amblyomma cajennense is an intrastadial biological vector of Theileria equi. Parasit Vectors 2013; 6(1): 306. PMid:24499587. http://dx.doi.org/10.1186/1756-3305-6-306
» http://dx.doi.org/10.1186/1756-3305-6-306 - Stamatakis A, Hoover P, Rougemont J, Renner S. A rapid bootstrap algorithm for the RAxML Web servers. Syst Biol 2008; 57(5): 758-771. PMid:18853362. http://dx.doi.org/10.1080/10635150802429642
» http://dx.doi.org/10.1080/10635150802429642 - Tenter AM, Friedhoff KT. Serodiagnosis of experimental and natural Babesia equi and B. caballi infections. Vet Parasitol 1986; 20(1-3): 49-61. PMid:3518217. http://dx.doi.org/10.1016/0304-4017(86)90092-0
» http://dx.doi.org/10.1016/0304-4017(86)90092-0 - Thompson JD, Higgins DG, Gibson TJ. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 1994; 22(22): 4673-4680. PMid:7984417. http://dx.doi.org/10.1093/nar/22.22.4673
» http://dx.doi.org/10.1093/nar/22.22.4673 - Ueti MW, Palmer GH, Kappmeyer LS, Scoles GA, Knowles DP. Expression of Equi Merozoite Antigen 2 during Development of Babesia equi in the Midgut and Salivary Gland of the Vector Tick Boophilus microplus. J Clin Microbiol 2003; 41(12): 5803-5809. PMid:14662988. http://dx.doi.org/10.1128/JCM.41.12.5803-5809.2003
» http://dx.doi.org/10.1128/JCM.41.12.5803-5809.2003 - Uilenberg G. Babesia--a historical overview. Vet Parasitol 2006; 138(1-2): 3-10. http://dx.doi.org/10.1016/j.vetpar.2006.01.035
» http://dx.doi.org/10.1016/j.vetpar.2006.01.035 - Vieira TS, Vieira RF, Finger MA, Nascimento DA, Sicupira PM, Dutra LH, et al. Seroepidemiological survey of Theileria equi and Babesia caballi in horses from a rural and from urban areas of Paraná State, southern Brazil. Ticks Tick Borne Dis 2013; 4(6): 537-541. PMid:24209493. http://dx.doi.org/10.1016/j.ttbdis.2013.07.005
» http://dx.doi.org/10.1016/j.ttbdis.2013.07.005 - World Organization for Animal Health – OIE. Manual of diagnostic tests and vaccines for terrestrial animals Paris: OIE; 2011.
Publication Dates
-
Publication in this collection
Jul-Sep 2017
History
-
Received
18 May 2017 -
Accepted
07 July 2017