Abstract
The objective of this work was to study the inheritance of soybean resistance to Soybean Steam Necrosis (SSN) and to map the resistance gene(s), using an F2 population from the cross between the soybean cultivars BRS 133 (resistant) and CD 206 (susceptible). A phenotypic evaluation was performed under controlled conditions, in a greenhouse. In a population of 114 F2 plants, 92 were resistant, and 22 were susceptible. The result is compatible with the inheritance of one dominant gene (χ2=1.98, P = 15.97%). Analysis of F1 and backcross populations confirmed the inheritance of one dominant gene. Bulk segregant analysis (BSA) was used as the mapping strategy. The resistance gene was mapped on chromosome 18 of the soybean genome, between markers Sat_308 and Satt303. This dominant gene is located on the same chromosome where the recessive SSN resistance gene rbc1 was mapped, at 8cM distance, and was named Rbc2.
Key words:
Glycine max; soybean steam necrosis; SSR; microsatellite markers; BSA; genetic mapping
INTRODUCTION
Brazil is the second largest soybean producer on the planet, representing 30% of the total world’s production. Soybean grain production accounted for 95.4 million tons in the 2015/2016 harvest season in Brazil, in a cultivated area of 33.2 million ha (CONAB 2017CONAB - Companhia Nacional de Abastecimento (2017) Séries históricas de área plantada, produtividade e produção. Available at < Available at http://www.conab.gov.br/conteudos.php?a=1252&t=2
> Accessed on March 1, 2017.
http://www.conab.gov.br/conteudos.php?a=...
), which illustrates the significance of the Brazilian soybean crops for the world’s economy.
Due to the large cultivation area, several pests constantly threaten soybean production in Brazil. The losses caused by diseases in soybean production in Brazil from 1997 to 2000 were estimated in $5.2 billion (Yorinori 2002Yorinori JT (2002) Situação atual das doenças potenciais no cone sul. In Anais do congresso brasileiro de soja, 1999. Embrapa Soja, Londrina, p. 171-186.). The use of resistant varieties is the most important alternatives to reduce losses caused by diseases, especially those caused by viruses since chemical control is non-existent or unfeasible.
The main virus disease found in soybean crops in Brazil is the soybean stem necrosis (SSN), caused by the CMMV - Cowpea mild mottle virus. This virus was identified in Brazilian soybean crops for the first time in 2000/2001. The virus has a great potential to cause production losses due to its fast widespread. Moreover, its vector, the whitefly (Bemisia tabaci), is very aggressive and difficult to control (Almeida et al. 2005Almeida AMR, Piuga FF, Marim SRR, Kitajima EW, Caspar JO, Oliveira TG and Moraes TG (2005) Detection and partial characterization of a carlavirus causing stem necrosis of soybean in Brazil. Fitopatologia Brasileira 30: 191-194.).
This virus has also damaged soybean crops in Argentina, Mexico, and Puerto Rico (Brace et al. 2012Brace RC, Fehr WR and Graham MA (2012) Inheritance and molecular mapping of an allele providing resistance to Cowpea mild mottle virus-like symptoms in soybean. Crop Science 52: 2109-2114.). Puerto Rico is not a significant soybean producer; however, the country hosts most winter nurseries of USA’s soybean breeding programs (USDA, Universities, and private companies). Thus, the disease can cause significant damage to these breeding programs. Another important fact is that seeds can also transmit the virus (Brunt and Kenten 1973Brunt AA and Kenten RH (1973) Cowpea mild mottle, a newly recognized virus infecting cowpeas (Vigna unguiculata) in Ghana. Annual Applied Biology 74: 67-74.). Thus, the seeds flow between Puerto Rico and the US can facilitate the entry of the virus into the continental USA, where almost the entire soybean area is vulnerable to CMMV.
The identification of resistance genes and the development of resistant varieties is the only way to avoid the damage caused by this virus. Brace et al. (2012Brace RC, Fehr WR and Graham MA (2012) Inheritance and molecular mapping of an allele providing resistance to Cowpea mild mottle virus-like symptoms in soybean. Crop Science 52: 2109-2114.) identified inheritance pattern of one recessive gene for resistance to SSN. The gene, named rbc1, was mapped on chromosome 18 (linkage group G) of the soybean genetic map. Suryanto et al. (2014Suryanto A, Kuswanto S and Kasno A (2014) Estimation of number and genes actions of CpMMV (Cowpea mild mottle virus) disease resistance genes on soybean crop. Journal of Agriculture and Veterinarian Sciences 7: 51-56.) identified inheritance patterns of two recessive genes or two dominant genes, depending on the source of resistance used. Arias et al. (2015Arias CAA, Almeida AMR, Mituti T and Kitakima EW (2015) Inheritance of tolerance to Cowpea Mild Mottle Virus in soybean. Crop Breeding and Applied Biotechnology 15: 132-138.) also identified inheritance of one recessive gene or two dominant genes, depending on the source of resistance.
The knowledge and the mapping of genes resistant to SSN are of fundamental for the obtainment of resistant varieties since molecular markers can be used to select resistant plants on a large scale, without the need for phenotypic evaluations. This work aimed to identify the inheritance of soybean resistance to SSN in the BRS 133 variety and map the resistance gene(s) present in this variety.
MATERIAL AND METHODS
The populations used in the study of inheritance and mapping were obtained by crossing the resistant variety BRS 133 with the susceptible variety CD 206, generating the F1, F2, BC1P1, and BC1P2 generations. BC1P1 was obtained by crossing F1 plants with the resistant parent (BRS 133), and BC1P2 was obtained by crossing F1 plants with the susceptible parent (CD206).
Populations were made of 25 plants of parent P1 (BRS 133), 27 plants of parent P2 (CD 206), 31 plants of F1 generation, 60 plants of BC1P1 generation, 81 plants of BC1P2 generation, and 114 plants of F2 generation.
SSN was phenotypically evaluated under controlled greenhouse conditions at the Coodetec research station (lat 24° 52' 54.9" S, long 53° 32' 30.4" W), in Cascavel, PR. The populations were planted in two-liter plastic pots containing a mixture of two parts of field soil, one part of sand, and one part of organic matter. Pots were daily irrigated to maintain soil moisture.
The CMMV isolate used for inoculation was obtained from infected soybean plants of the variety CD 206. The initial isolate was always kept in soybean plants of cultivar CD 206, in a greenhouse. The inoculum used for the phenotypic evaluation was prepared from symptomatic leaves macerated with a buffer solution (10 μM Sodium Phosphate pH 7.0) at the rate of 1 gram of symptomatic leaf for each 3 mL of buffer solution. The inoculum was prepared with the aid of a Wallita® mix apparatus.
For the inoculation, common charcoal was initially applied as a fine powder to the trifolium. Then, with a soft sponge, the solution of infected leaves crushed with the diluted buffer solution was slightly applied on the trifolium. The function of the milled charcoal and sponge was to cause small mechanical damage on the surface of the trifolium to allow virus infection. Two inoculations were carried out, one on the first fully expanded trifolium, at 19 days after sowing), and another at ten days after the first inoculation.
Plants were evaluated at 35 days after the second inoculation. The following scale was used to assess the SSN symptoms: 1) Absence of symptoms; 2) Chlorosis; 3) Chlorosis with light mosaic; 4) Chlorosis with strong mosaic and bullous mosaic. Plants with scores 1 and 2 were considered as resistant, and plants with scores 3 and 4 were considered as susceptible (Figure 1).
Scale for evaluation of the symptoms of soybean stem necrosis (SSN) under greenhouse conditions.
DNA was extracted as described by Doyle and Doyle (1990Doyle JJ and Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12: 13-15.). The Bulk Segregant Analysis (BSA) method was used for molecular markers identification. DNA bulks contained equimolar amounts of DNA from plants with the same phenotypes. Four bulks were formed, two consisting of DNA from resistant plants, and two consisting of DNA from susceptible plants. Each bulk contained DNA from six plants.
DNA amplification for BSA was performed with DNA samples from the parents and bulks in 20 μL solution containing 20 mM Tris-HCL (pH 8.3), 50 mM KCl, 3 mM MgCl2, 250 μM of each of the deoxynucleotides (dATP, dTTP, dGTP, and dCTP), 0.2 μM of each primer, one unit of enzyme Taq DNA polymerase, and 30 ng DNA. Amplification was performed in an MJ 96 thermal cycler programmed for an initial step of 3 minutes at 72 °C, followed by 35 cycles of 30 seconds at 94 °C, 30 seconds at 50 °C, 45 seconds at 72 °C, with a final step of 10 minutes at 72 °C.
The parents BRS 133 and CD 206 were screened with 209 microsatellite markers. Markers with different alleles for each parent were used in the BSA. After identifying a candidate marker, other markers mapped close to this candidate marker were added to the analysis to build a linkage group containing the resistance gene
The amplified SSR fragments were separated on 3% agarose gel containing Synergel and ethidium bromide, or on a 7% polyacrylamide denaturing gel. The gel type was defined as a function of the difference in size of the fragments of each allele, in each marker, obtained in the evaluation of the parents. Only polyacrylamide denaturing gel was used in the parents screening. Ethidium bromide-stained agarose gels were recorded in the Bio Capt photodocumentation system (Vilber Lourmat), and the polyacrylamide gels were scanned in a digital scanner, after being stained with silver nitrate.
The inheritance of resistance to soybean stem necrosis and the individual and joint segregation of the molecular markers were tested by the Chi-square test, using the software GENES (Cruz 2001Cruz CD (2001) Programa Genes: Versão Windows, aplicativo computacional em genética e estatística. UFV, Viçosa, 648p.). The linkage analysis between molecular markers and resistance gene was performed in the software GQMOL (Cruz and Schuster 2004Cruz CD and Schuster I (2004) GQMOL: aplicativo computacional para análise de dados moleculares e de suas associações com caracteres quantitativos. Version 2.1. UFV, Viçosa . Available at <Available at www.ufv.br/dbg/gqmol/gqmol.htm
> Accessed in March, 2017
www.ufv.br/dbg/gqmol/gqmol.htm...
), using the Kosambi mapping functions. Two loci were considered as linked when the recombination frequency between them was smaller than or equal to 35%, and the LOD score was greater than or equal to 3.0.
RESULTS AND DISCUSSION
The method used to evaluate the reaction of soybean plants to SSN under controlled conditions was efficient in discriminating resistant and susceptible plants. All plants of the susceptible cultivar (CD 206) obtained score 4, and presented symptoms of strong and bullous mosaic on the leaves. All the plants of the resistant cultivar (BRS 133) obtained score 1, without visible symptoms provoked by the infection with the virus.
In a population of 142 F2 plants, 92 were resistant, and 22 were susceptible (Table 1). This observed ratio agrees with the hypothesis of resistance conditioned by one dominant gene (P = 15.97%). In the BC1P2 population, the ratio of resistant and susceptible plants also fitted the hypothesis of one dominant gene (P = 43.67%).
Under the hypothesis of resistance conditioned by one dominant gene, it would be expected that 100% of the F1 plants and 100% of the plants of the BC1P1 population are resistant. One susceptible F1 plant and one susceptible BC1P1 plant were observed. The analysis of these plants using molecular markers confirmed they were F1 and BC1P1. The susceptibility of these plants may have been due to the stringency of the phenotypic evaluation protocol (possibility of confusing scores 2 and 3).
In Brazil, Arias et al. (2015Arias CAA, Almeida AMR, Mituti T and Kitakima EW (2015) Inheritance of tolerance to Cowpea Mild Mottle Virus in soybean. Crop Breeding and Applied Biotechnology 15: 132-138.) evaluated a population derived from the cross between the susceptible parent CD 206 and the resistant variety BRSMT Pintado and observed the inheritance of one recessive gene in BRSMT Pintado. When the authors evaluated the population derived from the cross between BRS 133 and BRSMT Pintado (two resistant varieties), they observed the segregation of two dominant genes, one coming from each parent, and the presence of only one gene resulted in resistant plants. The authors did not clarify the controversy between the expression of a recessive gene in BRSMT Pintado in the population of the cross with CD 206 and the inheritance of a dominant gene in the cross with BRS 133. In the present work, a dominant gene was observed in BRS 133 when crossed with the susceptible variety CD 206.
In Puerto Rico, Brace et al. (2012Brace RC, Fehr WR and Graham MA (2012) Inheritance and molecular mapping of an allele providing resistance to Cowpea mild mottle virus-like symptoms in soybean. Crop Science 52: 2109-2114.) identified one recessive gene for resistance of soybean to SSN in the resistant cultivar IA3023 when crossed with the susceptible cultivar IA3024. In Indonesia, Suryanto et al. (2014Suryanto A, Kuswanto S and Kasno A (2014) Estimation of number and genes actions of CpMMV (Cowpea mild mottle virus) disease resistance genes on soybean crop. Journal of Agriculture and Veterinarian Sciences 7: 51-56.) identified inheritance pattern of two recessive genes in the resistant variety MLG 0120, when crossed with the susceptible varieties Gumitir and Argopuro, and of two dominant genes in the resistant variety MLG 0278, when crossed with the same susceptible varieties. In both cases, the genes were epistatic, that is, the plants needed to have both genes to be resistant.
Resistance to some other viruses in soybean also presents a simple inheritance pattern, with alternation between dominant and recessive alleles expressing resistance. Goodrick et al. (1991Goodrick BJ, Kuhn CW and Boerma HR (1991) Inheritance of nonnecrotic resistance to cowpea chlorotic mottle virus in soybean. The Journal of Heredity 82: 512-514.) reported two recessive genes responsible for soybean resistance to Cowpea chlorotic mottle virus - CpCMV. Silva et al. (2004Silva MF, Kiihl RAS and Almeida AMR (2004) Inheritance of resistance to soybean mosaic virus in FT-10 soybean. Euphytica 135: 339-343.) assessed the inheritance to a new strain of Soybean mosaic virus (SMV) that broke the resistance of the soybean cultivar FT-10 in Brazil and identified an alternative allele for the Rsv1 locus, named as Rsv1 d .
Resistance to viruses in plants varied from simple to complex. The resistance of wheat to Barley yellow dwarf (BYDV) is polygenic (Ayala et al. 2002Ayala L, Henry M, Van Ginkel M, Singh R, Keller B and Khairallah M (2002) Identification of QTLs for BYDV tolerance in bread wheat. Euphytica 128: 249-259.). In cotton, one dominant gene provides resistance to blue disease, caused by the Cotton leaf roll dwarf virus (CLRDV) (Pupim Junior et al. 2008Pupin Junior O, Schuster I, Pinto RB, Pires E, Belot J-L, Silvie P, Chitarra Lg, Hoffmann Lv and Barroso P (2008) Inheritance of resistance to cotton blue disease. Pesquisa Agropecuaria Brasileira 43: 661-665.). In bean, resistance to Watermelon mosaic virus 2 (WMV 2) is controlled by two dominant genes (Kyle and Provvidenti 1987Kyle MM and Provvidenti R (1987) Inheritance of resistance to potato Y viruses in Phaseolus vulgaris L. Theoretical and Applied Genetics 74: 595-600.), and resistance to Soybean mosaic virus (SMV) is controlled by one gene (Smv) with incomplete dominance in the cultivar Great Northem 1140 (Provvidenti et al. 1982Provvidenti R, Gonsalves D and Ranall P (1982) Inheritance of resistance to soybean mosaic virus in Phaseolus vulgaris. The Journal of Heredity 73: 302-303.). In watercress, the inheritance of resistance to Watermelon mosaic virus 1 (WMV 1) is controlled by three genes with partial dominance in the cultivar ABL-010, and by two genes with additive effects in the cultivar Redlands Trailblazer (Maluf et al. 1997Maluf WR, Pereira JJ and Figueira AR (1997) Inheritance of resistance to the papaya ringspot virus-watermelon strain from two different accessions of winter squash Cucurbita maxima Duch. Euphytica 94: 163-168. ). Conversely, in watermelon, resistance to Watermelon mosaic virus (WMV) is polygenic (Beserra Júnior et al. 2006Beserra Junior JEA, Maluf WR, Figueira AR and Barguil BM (2006) Herança da resistência ao Watermelon mosaic virus em melancia (Citrullus lanatus L.). Fitopatologia Brasileira 31: 302-305., Azevedo et al. 2012Azevedo SM, Maluf WR, Faria MV, Resende JTV, Menezes CB and Nascimento IR (2012) Inheritance of resistance to the Papaya ringspot virus-watermelon strain (PRSV-W) from watermelon accession ‘PI 595201’. Crop Breeding and Applied Biotechnology 12: 67-75.). In ‘Whitaker’ summer squash line, one dominant gene summed to polygenic effect are associated to papaya ringspot virus-watermelon strain (PRSV-W) resistance (Menezes at al. 2015Menezes CB, Maluf WR, Faria MV, Azevedo SM, Resende JTV, Figueira AR and Gomes LAA (2015) Inheritance of resistance to papaya ringspot virus-watermelon strain (PRSV-W) in ‘Whitaker’ summer squash line. Crop Breeding and Applied Biotechnology 15: 203-209.).
In a set of 209 microsatellite markers used in parent screening, 56 (26.8%) were polymorphic among the varieties BRS 133 and CD 206. As the study of inheritance identified one dominant gene segregating in the F2 population, the bulk of resistant plants is expected to present homozygous dominant and heterozygous individuals, and the group of susceptible plants is expected to have only homozygous recessive individuals. Therefore, for the candidate marker, the resistant bulk should have only the allele of the resistant parent or both resistant and susceptible alleles. The susceptible bulk must present only the susceptible allele. This pattern was observed in the evaluation of the bulks with the marker Sat_308 (Figure 2).
A) Bulk Segregant Analysis (BSA) with the marker Sat_308. RP: resistant parent (BRS 133); SP: susceptible parent (CD 206); RB1 and RB2: resistant bulks; SB1 and SB2: susceptible bulks. Each bulk contains the mixture of DNA of six plants. B) Amplification of individual plants constituent for each bulk, with the marker Sat_308. RBC: Resistant bulk components; SBC: Susceptible bulk components.
Evaluation of individual plants from the bulks confirmed the possibility of linkage between the marker Sat_308 and the resistance gene since all susceptible plants were homozygous with the same genotype as CD 206 (Figure 2B). The DNA evaluation of the rest of the population confirmed this linkage (Table 2). The significance of the χ2 test indicates that the marker and the resistance gene do not segregate independently.
Sat_308 is located on chromosome 18 (Linkage group G) of the soybean genetic map (Song et al. 2004Song QJ, Marek LF, Shoemaker RC, Lark KG, Concibido VC, Delannay X, Specht JE and Cregan PB (2004) A new integrated genetic linkage map of the soybean. Theoretical and Applied Genetics 109: 122-128.). Other markers of chromosome 18 were added in the study to assist the localization of the gene in the soybean genetic map. The polymorphic markers of chromosome 18 segregated, as expected for an F2 population (Table 3), and, together with the resistance gene, four SSR markers formed a linkage group containing 89.14 cM of chromosome 18 (Figure 3).
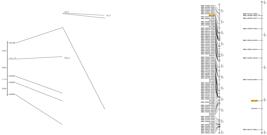
Mapping of the Rbc2 gene for resistance to soybean stem necrosis (SSN) on soybean chromosome 18, and comparison with the position of the rbc1 gene on the map. To the left, the linkage group obtained in the present work. To the center, part of chromosome 18 of the Soybean GmConsensus4.0 map: to the right, detail of this chromosome, highlighting the region between the Sat_308 marker and the BARC-901121-00988 marker. The alignment of the maps allows identifying the regions where the Rbc2 and rbc1 genes are mapped.
The order of the molecular markers in the linkage group agreed with the order of the same markers in the soybean consensus map (Song et al. 2004Song QJ, Marek LF, Shoemaker RC, Lark KG, Concibido VC, Delannay X, Specht JE and Cregan PB (2004) A new integrated genetic linkage map of the soybean. Theoretical and Applied Genetics 109: 122-128.). The SSN resistance gene was mapped between the SSR markers Sat_308 and Satt303, with a distance of 27.87 cM from Sat_308 and 29.64 cM from Satt303. In the Soybean-GmConsensus4.0 map, the markers Sat_308 and Satt303 are located to the positions 43 and 55 cM, respectively (Figure 3). In the middle of this interval, to the position expected for the resistance gene, the marker Satt340 is mapped at 48.5. However, this marker was monomorphic in the evaluated population.
Brace et al. (2012Brace RC, Fehr WR and Graham MA (2012) Inheritance and molecular mapping of an allele providing resistance to Cowpea mild mottle virus-like symptoms in soybean. Crop Science 52: 2109-2114.) mapped a recessive allele for resistance to SSN on chromosome 18, in cultivar IA3023. The authors mapped this gene at the same position as that of marker BARCSOYSSR_18_0443 and named it Rbc1 gene, and the recessive allele rbc1 was the resistance allele. The marker BARCSOYSSR_18_0443 is not mapped in the Soybean-GmConsensus4.0 map nor in the Soybean-GmComposite2003 map (www.soybease.org). In the Williams82 sequence map (versions Wm82.a1.v1 and Wm82.a2.v1), this marker is located between the markers BARC-901121-00988 and BARC-063589-18917, which are mapped to the positions 40 and 41 cM, on chromosome 18 of the Soybean-GmConsensus4.0 map (Figure 3).
The dominant resistance gene for SSN, identified in the present work, was mapped between the markers Sat_308 and Satt303, or between the positions 43 and 55 cM on the consensus map, approximately in the middle of this interval. The distance between the recessive gene previously mapped by Brace et al. (2012Brace RC, Fehr WR and Graham MA (2012) Inheritance and molecular mapping of an allele providing resistance to Cowpea mild mottle virus-like symptoms in soybean. Crop Science 52: 2109-2114.) and the dominant gene mapped in this study is 8 cM, or approximately 7Gbp, based on Wm82 map.a1.v1 (www.soybase.org). The new locus containing the dominant gene from variety BRS 133, mapped in this study, was named as Rps2, and the resistance allele was named Rps2, with the susceptible allele rps2.
In this work, another gene for resistance to soybean diseases was identified on chromosome 18 of the soybean genome. In this same chromosome several other genes was already identified for resistance to other soybean diseases, such as Heterodera glycines (Concibido et al. 1994Concibido VC, Denny RL, Boutin SR, Hautea R, Orf JH and Young ND (1994) DNA marker analysis of loci underlying resistance to soybean cyst-nematode (Heterodera glycines ichinohe). Crop Science 34: 240-246., Creagan et al. 1999Creagan PB, Mudge J, Fickus EW, Danesh D, Denny R and Young ND (1999) Two simple sequence repeat markers to select for soybean cyst nematode resistance conditioned by the rhg1 locus. Theoretical and Applied Genetics 99: 811-818., Yue et al. 2001Yue P, Arelli PR and Sleper DA (2001) Molecular characterization of resistance to Heterodera glycines in soybean PI 438489b. Theoretical and Applied Genetics 102: 921-928, Cervigni et al 2004Cervigni GDL, Schuster I, Barros EG and Moreira MA (2004) Two microsatellite markers flanking a dominant gene for resistance to soybean cyst nematode race 3. Euphytica 135: 99-105.); Meloydogine incognita (Li et al. 2001Li Z, Jakkula L, Hussey RS, Tamulonis JP and Boerma HR (2001) SSR mapping and confirmation of the QTL from PI 96354 conditioning soybean resistance to southern root-knot nematode. Theoretical and Applied Genetics 103:1167-1173.); Sclerotinia sclerotiorum (Arahana et al. 2001Arahana VS, Graef GL, Specht JE, Steadman JR and Eskridge KM (2001) Identification of QTLs for resistance to Sclerotinia sclerotiorum in soybean. Crop Science 41: 180-188.); Phytophthora sojae (Demirbas et al. 2001Demirbas A, Rector BG, Lohnes DG, Fioritto RJ, Graef GL, Cregan PB, Shoemaker RC and Specht JE (2001) Simple sequence repeat markers linked to the soybean rps genes for phytophthora resistance. Crop Science 41: 1220-1227.); Fusarium tucumaniae (Chang et al. 1997Chang SIC, Doubler TW, Kilo VY, Abuthredeih J, Prabhu R, Freire V, Suttner R, Klein J, Schmidt ME, Gibson PT and Lightfoot DA (1997) Association of loci underlying field resistance to soybean sudden death syndrome (SDS) and cyst nematode (SCN) Race 3. Crop Science 37: 965-971.); and Phakopsora pachyrhizi (Hyten et al. 2007Hyten DL, Hartman GL, Nelson RL, Frederick RD, Concibido VC, Narvel JM and Cregan PB (2007) Map location of the Rpp1 locus that confers resistance to soybean rust in soybean. Crop Science 47: 837-838., Silva et al. 2008Silva DCG, Yamanaka N, Brogin RL, Arias CAA, Nepomuceno AL, Di Mauro AL, Pereira SS, Nogueira LM, Passianoto ALL and Abdelnoor RV (2008) Molecular mapping of two loci that confer resistance to Asian rust in soybean. Theoretical and Applied Genetics 117: 57-63.). Linked genes were also identified on the chromosome 18 for most of these diseases.
Since each one of these genes comes from different sources of resistance, resistance alleles should be linked in the repulsion phase. For this reason, genetic breeding strategies should be used to break these linkages and generate progenies with linkage in the coupling phase between resistance alleles for different diseases. By obtaining such kind of germplasm with coupling linkage with many genes for resistance to diseases, breeding programs will easily obtain cultivars containing multiple resistance genes. The use of molecular markers identified in the various genetic mapping studies, including the markers identified in this work, will facilitate the obtainment of these recombined germplasm.
REFERENCES
- Almeida AMR, Piuga FF, Marim SRR, Kitajima EW, Caspar JO, Oliveira TG and Moraes TG (2005) Detection and partial characterization of a carlavirus causing stem necrosis of soybean in Brazil. Fitopatologia Brasileira 30: 191-194.
- Arahana VS, Graef GL, Specht JE, Steadman JR and Eskridge KM (2001) Identification of QTLs for resistance to Sclerotinia sclerotiorum in soybean. Crop Science 41: 180-188.
- Arias CAA, Almeida AMR, Mituti T and Kitakima EW (2015) Inheritance of tolerance to Cowpea Mild Mottle Virus in soybean. Crop Breeding and Applied Biotechnology 15: 132-138.
- Ayala L, Henry M, Van Ginkel M, Singh R, Keller B and Khairallah M (2002) Identification of QTLs for BYDV tolerance in bread wheat. Euphytica 128: 249-259.
- Azevedo SM, Maluf WR, Faria MV, Resende JTV, Menezes CB and Nascimento IR (2012) Inheritance of resistance to the Papaya ringspot virus-watermelon strain (PRSV-W) from watermelon accession ‘PI 595201’. Crop Breeding and Applied Biotechnology 12: 67-75.
- Beserra Junior JEA, Maluf WR, Figueira AR and Barguil BM (2006) Herança da resistência ao Watermelon mosaic virus em melancia (Citrullus lanatus L.). Fitopatologia Brasileira 31: 302-305.
- Brace RC, Fehr WR and Graham MA (2012) Inheritance and molecular mapping of an allele providing resistance to Cowpea mild mottle virus-like symptoms in soybean. Crop Science 52: 2109-2114.
- Brunt AA and Kenten RH (1973) Cowpea mild mottle, a newly recognized virus infecting cowpeas (Vigna unguiculata) in Ghana. Annual Applied Biology 74: 67-74.
- Cervigni GDL, Schuster I, Barros EG and Moreira MA (2004) Two microsatellite markers flanking a dominant gene for resistance to soybean cyst nematode race 3. Euphytica 135: 99-105.
- Chang SIC, Doubler TW, Kilo VY, Abuthredeih J, Prabhu R, Freire V, Suttner R, Klein J, Schmidt ME, Gibson PT and Lightfoot DA (1997) Association of loci underlying field resistance to soybean sudden death syndrome (SDS) and cyst nematode (SCN) Race 3. Crop Science 37: 965-971.
- CONAB - Companhia Nacional de Abastecimento (2017) Séries históricas de área plantada, produtividade e produção. Available at < Available at http://www.conab.gov.br/conteudos.php?a=1252&t=2 > Accessed on March 1, 2017.
» http://www.conab.gov.br/conteudos.php?a=1252&t=2 - Concibido VC, Denny RL, Boutin SR, Hautea R, Orf JH and Young ND (1994) DNA marker analysis of loci underlying resistance to soybean cyst-nematode (Heterodera glycines ichinohe). Crop Science 34: 240-246.
- Creagan PB, Mudge J, Fickus EW, Danesh D, Denny R and Young ND (1999) Two simple sequence repeat markers to select for soybean cyst nematode resistance conditioned by the rhg1 locus. Theoretical and Applied Genetics 99: 811-818.
- Cruz CD (2001) Programa Genes: Versão Windows, aplicativo computacional em genética e estatística. UFV, Viçosa, 648p.
- Cruz CD and Schuster I (2004) GQMOL: aplicativo computacional para análise de dados moleculares e de suas associações com caracteres quantitativos. Version 2.1. UFV, Viçosa . Available at <Available at www.ufv.br/dbg/gqmol/gqmol.htm > Accessed in March, 2017
» www.ufv.br/dbg/gqmol/gqmol.htm - Demirbas A, Rector BG, Lohnes DG, Fioritto RJ, Graef GL, Cregan PB, Shoemaker RC and Specht JE (2001) Simple sequence repeat markers linked to the soybean rps genes for phytophthora resistance. Crop Science 41: 1220-1227.
- Doyle JJ and Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12: 13-15.
- Goodrick BJ, Kuhn CW and Boerma HR (1991) Inheritance of nonnecrotic resistance to cowpea chlorotic mottle virus in soybean. The Journal of Heredity 82: 512-514.
- Hyten DL, Hartman GL, Nelson RL, Frederick RD, Concibido VC, Narvel JM and Cregan PB (2007) Map location of the Rpp1 locus that confers resistance to soybean rust in soybean. Crop Science 47: 837-838.
- Kyle MM and Provvidenti R (1987) Inheritance of resistance to potato Y viruses in Phaseolus vulgaris L. Theoretical and Applied Genetics 74: 595-600.
- Li Z, Jakkula L, Hussey RS, Tamulonis JP and Boerma HR (2001) SSR mapping and confirmation of the QTL from PI 96354 conditioning soybean resistance to southern root-knot nematode. Theoretical and Applied Genetics 103:1167-1173.
- Maluf WR, Pereira JJ and Figueira AR (1997) Inheritance of resistance to the papaya ringspot virus-watermelon strain from two different accessions of winter squash Cucurbita maxima Duch. Euphytica 94: 163-168.
- Menezes CB, Maluf WR, Faria MV, Azevedo SM, Resende JTV, Figueira AR and Gomes LAA (2015) Inheritance of resistance to papaya ringspot virus-watermelon strain (PRSV-W) in ‘Whitaker’ summer squash line. Crop Breeding and Applied Biotechnology 15: 203-209.
- Provvidenti R, Gonsalves D and Ranall P (1982) Inheritance of resistance to soybean mosaic virus in Phaseolus vulgaris The Journal of Heredity 73: 302-303.
- Pupin Junior O, Schuster I, Pinto RB, Pires E, Belot J-L, Silvie P, Chitarra Lg, Hoffmann Lv and Barroso P (2008) Inheritance of resistance to cotton blue disease. Pesquisa Agropecuaria Brasileira 43: 661-665.
- Silva DCG, Yamanaka N, Brogin RL, Arias CAA, Nepomuceno AL, Di Mauro AL, Pereira SS, Nogueira LM, Passianoto ALL and Abdelnoor RV (2008) Molecular mapping of two loci that confer resistance to Asian rust in soybean. Theoretical and Applied Genetics 117: 57-63.
- Silva MF, Kiihl RAS and Almeida AMR (2004) Inheritance of resistance to soybean mosaic virus in FT-10 soybean. Euphytica 135: 339-343.
- Song QJ, Marek LF, Shoemaker RC, Lark KG, Concibido VC, Delannay X, Specht JE and Cregan PB (2004) A new integrated genetic linkage map of the soybean. Theoretical and Applied Genetics 109: 122-128.
- Suryanto A, Kuswanto S and Kasno A (2014) Estimation of number and genes actions of CpMMV (Cowpea mild mottle virus) disease resistance genes on soybean crop. Journal of Agriculture and Veterinarian Sciences 7: 51-56.
- Yorinori JT (2002) Situação atual das doenças potenciais no cone sul. In Anais do congresso brasileiro de soja, 1999. Embrapa Soja, Londrina, p. 171-186.
- Yue P, Arelli PR and Sleper DA (2001) Molecular characterization of resistance to Heterodera glycines in soybean PI 438489b. Theoretical and Applied Genetics 102: 921-928
Publication Dates
-
Publication in this collection
Apr-Jun 2018 -
Date of issue
Apr 2018
History
-
Received
05 Apr 2017 -
Accepted
27 Aug 2017