Abstract
The clustered regularly interspaced short palindromic repeat (CRISPR)/CRISPR-associated (Cas) technology has recently emerged as a powerful genomic editing tool with great potential for crop breeding. However, commonly used protocols for screening of CRISPR/Cas9-induced mutations are laborious, time-consuming, and costly. In the present study we examined the applicability of high resolution melting (HRM) analysis fast screening CRISPR/Cas9-induced mutations in T0 plants and subsequent genotyping of T1 populations. Comparative analysis demonstrated that HRM analysis could identify mutant T0 plants carrying various types of mutation, including single nucleotide substitutions and short insertion/deletions, with false positive/negative rates of 0-2.78%. Furthermore, T1 plants derived from single T0 plants could be correctly genotyped by HRM analysis using WT parents and T0 plants as controls. We hence recommend the adoption of HRM analysis in CRISPR/Cas9 mediated genetic studies and breeding in rice and other crop species.
Key words:
Genomic editing; mutation screening; OsLCT1; bell
INTRODUCTION
The clustered regularly interspaced short palindromic repeat (CRISPR)/CRISPR-associated (Cas) system has become the most efficient genome editing tool for biological research in a wide spectrum of organisms (Mao et al. 2013Mao Y, Zhang H, Xu N, Zhang B and Gao F (2013) Application of the CRISPR/Cas system for efficient genome engineering in plants. Molecular Plant 6: 691-693.). In plants, this technology was first successfully applied to Arabidopsis, tobacco, sorghum and rice (Jiang et al. 2013Jiang W, Zhou H, Bi H, Fromm M, Yang B and Weeks DP (2013) Demonstration of CRISPR/Cas/sgRNA-mediated targeted gene modification in Arabidopsis, tobacco, sorghum and rice. Nucleic Acids Research 41: e188. doi:10.1093/nar/gkt780.
https://doi.org/e188. doi:10.1093/nar/gk...
, Li et al. 2013Li JF, Norville JE, Aach J, McCormack M, Zhang DD, Bush J, Church GM and Sheen J (2013) Multiplex and homologous recombination-mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas. Nature Biotechnology 31: 688-691., Miao et al. 2013Miao J, Guo DS, Zhang JZ, Huang QP, Qin GJ, Zhang X, Wan JM, Gu HY, Qu LJ (2013) Targeted mutagenesis in rice using CRISPR-Cas system. Cell Research 23: 1233-1236., Nekrasov et al. 2013Nekrasov V, Staskawicz B, Weigel D, Jones JD and Kamoun S (2013) Targeted mutagenesis in the model plant Nicotiana benthamiana using Cas RNA-guided endonuclease. Nature Biotechnology 31: 691-693.), and has since been proven to be an effective and simple method for targeted mutagenesis in many other plant species. This technology has now gained consensus among experts as being a major breakthrough in plant studies and genetic improvement (see reviews by Luo et al. 2016Luo M, Gilbert B and Ayliffe M (2016) Applications of CRISPR/Cas technology for targeted mutagenesis, gene replacement and stacking of genes in higher plants. Plant Cell Reports 7: 1439-1450., Ma et al. 2016Ma X, Zhu Q, Chen Y and Liu YG (2016) CRISPR/Cas platforms for genome editing in plants: developments and applications. Molecular Plant 9: 961-974.).
Identification of mutations in the targeted region is an important step in CRISPR/Cas9 mutagenesis. Although there are already a few methods available for this purpose, detection using endonucleases is often the method of choice (reviewed by Ma et al. 2016Ma X, Zhu Q, Chen Y and Liu YG (2016) CRISPR/Cas platforms for genome editing in plants: developments and applications. Molecular Plant 9: 961-974.). For instance, endonucleases such as CELI, appropriate restriction enzymes Pst I, Aat II, and Msc I were used in mutation identification (Wang et al. 2015Wang C, Shen L, Fu YP, Yan CJ and Wang KJ (2015) A simple CRISPR/Cas system for multiplex genome editing in rice. Journal of Genetics and Genomics 42: 703-706., Li et al. 2016Li MR, Li XX, Zhou ZJ, Wu PZ, Fang MC, Pan XP, Lin QP, Luo WB, Wu GJ and Li HQ (2016) Reassessment of the four yield-related genes Gn1a, DEP1, GS3, and IPA1 in rice using a CRISPR/Cas system. Frontiers in Plant Science 7: 377. doi: 10.3389/fpls.2016.00377.
https://doi.org/10.3389/fpls.2016.00377....
), and T7 endonuclease I (T7EI, Hadden et al. 2007Hadden JM, Déclais AC, Carr SB, Lilley DMJ and Phillips SEV (2007) The structural basis of Holliday junction resolution by T7 endonuclease I. Nature 449: 621-624. ) was recommended for the standard CRISPR/Cas9-based genomic editing studies (Cho et al. 2013Cho SW, Kim S, Kim JM and Kim JS (2013) Targeted genome engineering in human cells with the Cas RNA-guided endonuclease. Nature Biotechnology 31: 230-232.). Recently, Zheng et al. (2016Zheng XL, Yang SX, Zhang DW, Zhong ZH, Xu T, Deng KJ, Zhou JP, Qi YP and Zhang Y (2016) Effective screen of CRISPR/Cas-induced mutants in rice by single-strand conformation polymorphism. Plant Cell Reports 35: 1545-1554.) reported that single-strand conformation polymorphism (SSCP) analysis can also be used for the effective screening of CRISPR/Cas9-induced mutants in rice. While both T7EI (restriction) and SSCP analysis can identify different types of mutations generated by CRISPR/Cas9 mutagenesis, they are time-consuming and may generate false positive results when T7EI is used (Hadden et al. 2007Hadden JM, Déclais AC, Carr SB, Lilley DMJ and Phillips SEV (2007) The structural basis of Holliday junction resolution by T7 endonuclease I. Nature 449: 621-624. ).
High resolution melting (HRM) curve analysis, which is based on fluorescence changes during the melting of the DNA duplex, is a simple, cost-effective and high throughput method. It has been widely used for mutation scanning and genotyping in general (Koeyer et al. 2010Koeyer DD, Douglass K, Murphy A, Whitney S, Nolan L, Song Y and Jong WD (2010) Application of high-resolution DNA melting for genotyping and variant scanning of diploid and autotetraploid potato. Molecular Breeding 25: 67-90., Lochlainn et al. 2011Lochlainn SO, Amoah S, Graham NS, Alamer K, Rios JJ, Kurup S, Stoute A, Hammond JP, Østergaard L, King KJ, White PJ and Broadley MR (2011) High resolution melt (HRM) analysis is an efficient tool to genotype EMS mutants in complex crop genomes. Plant Methods 7: 43. doi:10.1186/1746-4811-7-43.
https://doi.org/10.1186/1746-4811-7-43...
), and for testing food products and seeds in particular (reviewed by Simko et al. 2016Simko I (2016) High-resolution DNA melting analysis in plant research. Trends in Plant Science 21: 528-537.). However, the use of HRM for identification of mutations generated by CRISPR/Cas9-based research was mentioned by Fauser et al. (2014Fauser F, Schiml S and Puchta H (2014) Both CRISPR/Cas-based nucleases and nickases can be used efficiently for genome engineering in Arabidospsis thaliana. Plant Journal 79: 348-359. ) and Shen et al. (2017Shen H, Strunks GD, Klemann BJ, Hooykaas PJ and Pater DS (2017) CRISPR/Cas9-induced double-strand break repair in Arabidopsis nonhomologous end -joining mutants.G3-Genes Genomes Genetics 7: 193-202.) in Arabidopsis. Contrary to the general notion that HRM is a sensitive mutation detection method (Dahlem et al. 2012Dahlem TJ, Hoshijima K, Jurynec MJ, Gunther D, Starker CG, Locke AS, Weis AM, Voytas DF and Grunwald DJ (2012) Simple methods for generating and detecting locus-specific mutations induced with TALENs in the zebrafish genome. PLoS Genetics 8: e1002861. [doi:10.1371/journal.pgen.1002861]
https://doi.org/e1002861. [doi:10.1371/j...
), Ma et al. (2016Ma X, Zhu Q, Chen Y and Liu YG (2016) CRISPR/Cas platforms for genome editing in plants: developments and applications. Molecular Plant 9: 961-974.) suggest that the detection sensitivity of HRM is relatively low; however they did not provide additional material to substantiate this claim. Therefore, more studies are needed to assess the suitability of HRM analysis for detection and genotyping of CRISPR/Cas9-induced mutations.
In the present study we mutagenized two genes in rice via CRISPR/Cas9-based genome editing, and assessed the applicability of HRM for mutation screening and genotyping. Our results demonstrate that HRM analysis provides an efficient method for screening and genotyping genetic variations induced by CRISPR/Cas9-based genome editing.
MATERIAL AND METHODS
Production of CRISPR/Cas9 transgenic plants
For CRISPR/Cas9 mutagenesis, we selected two rice genes, namely rice low-affinity cation transporter1 (OsLCT1, LOC_Os06g38120), which encodes a cadmium transporter (Uraguchi et al. 2011Uraguchi S, Kamiya T, Sakamoto T, Kasai K, Sato Y, Nagamura Y, Yoshida A, Kyozuka J, Ishikawa S and Fujiwara T (2011) Low-affinity cation transporter (oslct1) regulates cadmium transport into rice grains. Proceeding of the National Academy of Sciences of the United States of America 108: 20959-20964.), and rice Bentazon Lethal 1 (BEL1, LOC_Os03g55240), which encodes a P450 protein CYP81A6 that confers resistance to bentazon and sulfonylurea herbicides (Pan et al. 2006Pan G, Zhang X, Liu K, Zhang J, Wu X, Zhu J and Tu J (2006) Map-based cloning of a novel rice cytochrome p450 gene cyp81a6, that confers resistance to two different classes of herbicides. Plant Molecular Biology 61: 933-943.). Oligo primers for sgRNAs were designed using the following web resource at http://e-crisp-test.dkfz.de/E-CRISP/designcrispr.html (Table 1). These primers were subsequently annealed and inserted into the sgRNA scaffold vector PHUN4C12 (Xu et al. 2014Xu RF, Li H, Qin RY, Wang L, Li L, Wei PC and Yang JB (2014) Gene targeting using the Agrobacterium tumefaciens-mediated CRISPR-Cas system in rice. Rice 7: 5. doi:10.1186/s12284-014-0005-6.
https://doi.org/10.1186/s12284-014-0005-...
). These loaded vectors were named phun4c12-lct1 and phun4c12-bel, respectively, for OsLCT1 and BEL1. They were then transformed into the Agrobacterium tumefaciens pSOUP-EHA105 to produce transgenic rice plants via Agrobacterium-mediated transformation according to Xu et al. (2014Xu RF, Li H, Qin RY, Wang L, Li L, Wei PC and Yang JB (2014) Gene targeting using the Agrobacterium tumefaciens-mediated CRISPR-Cas system in rice. Rice 7: 5. doi:10.1186/s12284-014-0005-6.
https://doi.org/10.1186/s12284-014-0005-...
). Rice cultivars Xidao #3 and Nipponbare were used for transformation of phun4c12-lct1 and phun4c12-bel, respectively.
DNA Oligos used for CRISPR/Cas 9 vector construction and PCR primer sequences for HRM analysis and T-DNA detection of two genes
The plantlets regenerated from a common hygromycin-resistant callus were considered as a single independent T0 plant. Seeds harvested from individual panicles of these T0 plants were grown into T1 populations in panicle rows. For identification of transgene positive T0 plants, genomic DNA was extracted from leaf tissues using a modified CTAB method (Zhang et al. 2014Zhang HL, Huang JZ, Chen XY, Tan YY and Shu QY (2014) Competitive amplification of differentially melting amplicons facilitates efficient genotyping of photoperiod- and temperature-sensitive genic male sterility in rice. Molecular Breeding 34: 1765-1776.), and the concentration diluted to approximately 50 ng/µl after quantification using a Nanodrop 2000 (Thermo Scientific, USA). The presence of the transgene was assessed by amplification of the U6 promoter driving the Cas9 gene in the vectors using Ubi primers (Table 1) based on the Ubi promoter (Xu et al. 2014Xu RF, Li H, Qin RY, Wang L, Li L, Wei PC and Yang JB (2014) Gene targeting using the Agrobacterium tumefaciens-mediated CRISPR-Cas system in rice. Rice 7: 5. doi:10.1186/s12284-014-0005-6.
https://doi.org/10.1186/s12284-014-0005-...
). PCRs were performed in aliquots of 20 µL with 40 ng of genomic DNA, 10 µL of 2× master mix and 0.4 µL each of 10 µM primers. The following PCR conditions were used: 5 min at 94 °C, followed by 35 cycles of 30 s at 94 °C, 30 s at 55 °C and 30 s at 72 °C, with a final extension at 72 °C for 8 min and hold at 4 °C. The PCR products were separated on 1.0 % agarose gels and detected with ethidium bromide.
HRM identification of genetic variations
For detection of mutations in the target region in T0 and T1 plants, site-specific primers were used to amplify fragments of 100-500 bp encompassing potential mutations induced by CRISPR/Cas9 (Table 1).
PCRs were performed in a 10 µL volume with 25 ng of genomic DNA, 5 µL of 2×master mix (containing 2× PCR buffer, 4 mM MgCl2, 0.4 mM dNTPs, 50 units mL-1 Taq DNA polymerase, TOYOBO Co., Ltd.), 0.2 µL each of 10 µM primers and 1 µL of 10 × EvaGreen (Biotium, USA), covered with a drop of mineral oil to prevent solution evaporation. The following PCR conditions were used: 5 min at 94 °C, followed by 35-40 cycles of 30 s at 94 °C, 30 s at 55-62 °C and 30 s at 72 °C, with a final extension at 72 °C for 8 min and hold at 16 °C. The annealing temperatures were optimized for amplification of the respective fragments.
Following PCR, plates were transferred to a Lightscanner (Idaho Technology Inc., USA) and subjected to HRM analysis according to Tan et al. (2016Tan YY, Yu XM, Shu QY, Zhang HL, Wang SG, Yuan FJ and Shi CH (2016) Development of an HRM-based, safe and high-throughput genotyping system for two low phytic acid, mutations in soybean. Molecular Breeding 36: 1-9.). In brief, the temperature was ramped up from 55 to 95 °C at 0.1 °C per second and data were analyzed using its proprietary software, Call ITTM 2.0 (Idaho Technology Inc., USA) after normalization and temperature shifting of the melting curves according to the Lightscanner Operator’s Manual (Idaho Technology Inc.). Samples with relative fluorescence differences (|ΔF|) > 0.05 were considered to be significantly different from the reference (Hofinger et al. 2009Hofinger BJ, Jing HC, Kosack KEH and Kanyuka K (2009) High-resolution melting analysis of cDNA-derived PCR amplicons for rapid and cost-effective identification of novel alleles in barley. Theoretical and Applied Genetics 119: 851-865.).
Validation by TA cloning and sequencing analysis
During the process of CRISPR/Cas9 vector construction, the plasmids were sequenced to assure no mismatch had occurred. The target fragments for T0 and T1 plants were also sequenced to validate the effectiveness of HRM analysis.
For DNA sequencing, PCRs were performed in 50 µL volumes and the amplicons were separated on 1.0 % agarose gels. The target DNAs were recovered using DNA Gel Extraction Kit (Axygen Biotechnology Co. Ltd., Hangzhou, China) for subsequent sequencing by Shanghai RuiDi Biological Technology Co., Ltd. (Shanghai, China). The sequencing results were analyzed by DSDecode software (http://dsdecode.scgene.com/) (Liu et al. 2015Liu WZ, Xie XR, Ma XL, Li J, Chen JH and Liu YG (2015) DSDecode: a web-based tool for decoding of sequencing chromatograms for genotyping of targeted mutations. Molecular Plant 8: 1431-1433.). All the T0 plants of phun4c12-bel were sent for TA cloning. For T0 plants of phun4c12-lct1, when sequencing data were not readily recognized by DSDecode, samples were further analyzed by TA cloning: target DNAs were purified and cloned into pGEM®-T Easy Vector (Promega Biotechnology Co. Ltd., Beijing, China), and then transformed into competent E. coli DH5α cells. Six white colonies (recombinant colonies) were selected and sequenced for each fragment.
Primers were designed using Primer premier 5 software and were synthesized by Shanghai Sangong Biological Engineering Technology & Services Co., Ltd. (Shanghai, China).
RESULTS AND DISCUSSION
A total of 128 and 72 independent T0 plants were obtained for the phun4c12-lct1 and phun4c12-bel construct, respectively. PCR analysis confirmed that they all carried the U6 promoter, hence all were transgenic. These independent T0 plants were subjected to subsequent mutation analysis.
Mutant T0 plants revealed by HRM analysis
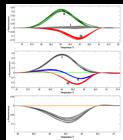
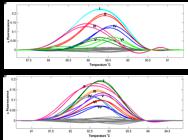
Among the 128 transgenic phun4c12-lct1 T0 plants, the |ΔF| of 70 plants (54.69%) was greater than 0.05 when their HRM curves were compared with that of Xidao #3 (their wild type parent), while |ΔF| of the remaining 58 was < 0.05 (Figure 1a). Similarly, 47 plants (65.28%) out of the 72 transgenic phun4c12-bel T0 plants were shown to have HRM curves significantly different from their wild type parent Nipponbare (Figure 1b).
Representative HRM analysis of T0 plants with their OsLCT1 (a) or Bel1 (b) genes target-mutagenized by CRISPR/Cas9. The wild type parents, which are shown as the horizontal lines, are chosen as reference for the development of fluorescence difference curves of T0 plants. Fluorescence difference curves are automatically grouped by the HRM system as shown in different colors. These groups are also labeled by different Roman numerals for subsequent analysis.
HRM curves that have small differences from each other were automatically grouped together and shown in the same color (Figure 1). HRM curves that are significantly different from their wild type parent were further divided into different groups marked with different colors. A total of six and seven major groups were found in T0 plants of phun4c12-lct1 and phun4c12-bel, respectively (Figure 1).
Mutations revealed by sequencing
To evaluate the sensitivity of mutation detection by HRM analysis, all transgenic T0 plants were subjected to Sanger sequencing. All T0 plants that had HRM curves significantly different from their respective wild type parents carried at least one mutant allele at the target region (Tables 2 and 3), suggesting they were true mutants. Hence the false positive rate of HRM detection of mutants was zero in both T0 populations.
Sequencing also revealed that each group of T0 plants based on HRM analysis could be further divided into more than two subgroups that have different genotypes (Tables 2 and 3). For instance, there were four types of oslct1 mutant T0 plant in group I (Table 2) and five types of bel1 mutant T0 plant in group II (Table 3). Nevertheless, common mutations were always identified in different T0 plants within the same group, e.g., 15 T0 plants including #33-37, #69, #80-81, #83-84, #88-#90, and #101-102, in group III of phun4c12-lct1 had the same biallelic mutation of 1-bp insertion and 1-bp deletion (Table 2). In total, 24 genotypes of OsLCT1 and 31 genotypes of BEL1 were detected. Similar mutational profiles in T0 plants were reported in previous studies (Feng et al. 2014Feng Z, Mao Y, Xu N, Zhang B, Wei P, Yang DL, Wang Z, Zhang Z, Zheng R, Yang L, Zeng L, Liu X and Zhu JK (2014) Multigeneration analysis reveals the inheritance, specificity, and patterns of CRISPR/Cas-induced gene modifications in Arabidopsis. Proceeding of the National Academy of Sciences of the United States of America 111: 4632-4637. , Ma et al. 2015Ma XL, Zhang QY, Zhu QL, Liu W, Chen Y, Qiu R, Wang B, Yang ZF, Li HY, Lin Y, Xie YY, Shen RX, Chen SF, Wang Z, Chen YL, Guo JX, Chen LT, Zhao XC, Dong ZC and Liu YG (2015) A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Molecular Plant 8: 1274-1284.). These results demonstrated that HRM analysis is suitable for identification of various kinds of mutation generated by CRISPR/Cas9 mutagenesis.
Because false negatives may exist in HRM analysis, samples having |ΔF| < 0.05 from their wild type parent were also sequenced. Two transgenic phun4c12-lct1 plants and two phun4c12-bel plants were detected to carry mutations (data not shown), amounting to false negative rates of 1.56% and 2.78%, respectively, for OsLCT1 and BEL1. HRM analysis may produce false positive or negative results. Chen et al (2012Chen ST, Su YN, Ni YH, Hu WL, Lee NC, Chien YH, Chang CC, Chen HL and Chang MH (2012) Diagnosis of neonatal intrahepatic cholestasis caused by citrin deficiency using high-resolution melting analysis and a clinical scoring system. Journal of Pediatrics 161: 626-631. ) found that the false positive rate and false negative rate of HRM analysis were 1.25% and null, respectively. Furthermore, since we were also unable to distinguish a mutant (with a 6-bp deletion) from its wild type parent by direct HRM analysis (Tan et al. 2013Tan YY, Fu HW, Zhao HJ, Lu S, Fu JJ, Li YF, Cui HR and Shu QY (2013) Functional molecular markers and high-resolution melting curve analysis of low phytic acid mutations for marker-assisted selection in rice. Molecular Breeding 31: 517-528.), there is thus a chance that certain types of mutation cannot be detected by this technique. In the present study, the rate of false positives is zero and the false negative rate is less than 2.78%. Our study thus suggests that HRM analysis is a sensitive method for mutation detection. This finding concurs with the observation of Dahlem et al. (2012Dahlem TJ, Hoshijima K, Jurynec MJ, Gunther D, Starker CG, Locke AS, Weis AM, Voytas DF and Grunwald DJ (2012) Simple methods for generating and detecting locus-specific mutations induced with TALENs in the zebrafish genome. PLoS Genetics 8: e1002861. [doi:10.1371/journal.pgen.1002861]
https://doi.org/e1002861. [doi:10.1371/j...
) in TALEN studies, but is contrary to reports by Ma et al. (2016Ma X, Zhu Q, Chen Y and Liu YG (2016) CRISPR/Cas platforms for genome editing in plants: developments and applications. Molecular Plant 9: 961-974.).
Genotyping of T1 plants
To assess the usefulness of HRM analysis for genotyping mutations generated through CRISPR/ Cas9 mutagenesis, T1 plants derived from three independent T0 phun4c12-lct1 plants, i.e., #73 (heterozygous), #6 (homozygous) and #77 (biallelic) (Table 1), were selected for HRM analysis. Their wild type parent Xidao #3 was used as reference in fluorescence difference curve development and their T0 plants used as a control.
In the T1 progeny of T0 plant #73, three groups were established based on their |ΔF| values as compared with that of the wild type: Group I plants were indistinguishable from Xidao #3, suggesting that they were homozygous wild type lines; Group II plants were similar to T0 plants, suggesting they were heterozygous at the target region; by contrast, group III plants differed from group I and II, hence they should be homozygous mutant lines (Figure 2a).
HRM analysis of T1 plants derived from three representative T0 plants (A: #73; B: #77; C: #6) of phun4c12-lct1. The wild type parents Xidao #3 was used as reference for fluorescence curve development and is shown as the horizontal lines. Fluorescence difference curves are automatically grouped by the HRM system as shown in different colors. These groups are also labeled by different Roman numerals for subsequent analysis. The curve of respective T0 plant was included in each analysis and is indicated in black.
In the T1 progeny of T0 plant #77, test samples were all significantly different from Xidao #3, suggesting there were no wild type plants. Samples of group I had |ΔF| > 0.3 from that of the wild type, with HRM curves similar to that of the T0 plant #77 (Figure 2b), suggesting they also carried biallelic mutations of OsLCT1. Samples of the other two groups had |ΔF| greater than 0.1 (Figure 2b), implicating they should be homozygous mutants for the 1-bp insertion or the 8-bp deletion. Sequencing of representative samples of group II and group III indicated that the former had the 1-bp insertion and the latter had the 8-bp deletion.
Genotyping of the T1 plants derived from T0 plant #6 revealed only a single type of HRM curve, with |ΔF| > 0.05 from the wild type and indistinguishable from the T0 plant (Figure 2c), suggesting these plants were all homozygous mutants as was the T0 plant #6.
Advantages of HRM analysis for mutation screening and genotyping in CRISPR/Cas9 mediated plant genetics and breeding
Although a number of methods are used in mutation detection in CRISPR/Cas9 studies, the HRM method appears to have several advantages: 1) It is easier and simpler to handle since there is no need for post-PCR sample processing or separation; the post-PCR operation is a closed-tube process (which minimizes operation errors) and takes only 15 min. Compared with HRM, other methods take much longer. For example, the T7EI method takes at least two hours after PCR; PAGE-based assay and SSCP all need PAGE electrophoresis and staining, which can take more than six hours. 2) The HRM method enables 96 samples to be tested at one time, making it more cost-effective and more suitable for high throughput screening.
Montgomery et al. (2007Montgomery J, Wittwer CT, Palais R and Zhou L (2007) Simultaneous mutation scanning and genotyping by high-resolution DNA melting analysis. Nature Protocols 2: 59-66.) reported that the fragment length has an impact on the screening efficiency of mutations randomly positioned and that fragments of 100 - 400 bp were best for HRM analysis. The mutations induced by CRISPR/Cas9 are known to be concentrated around the target (Shan et al. 2013Shan Q, Wang Y, Li J, Zhang Y, Chen K, Liang Z, Zhang K, Liu J, Xi JJ, Qiu JL and Gao C (2013) Targeted genome modification of crop plants using a CRISPR/Cas system. Nature Biotechnology 31: 686-688.), hence a fragment of 400 bp is sufficient to encompass expected mutations and hence, the length limitation of HRM has little impact on detecting mutations induced by CRISPR/Cas9. Indeed, the use of HRM as a method for screening and genotyping of CRISPR/Cas9 induced mutations was already adopted in a few recent studies, e.g. Chang et al. (2016Chang ZY, Chen ZF, Wang N, Xie G, Lu JW, Yan W, Zhou JL, Tang XY and Deng XW (2016) Construction of a male sterility system for hybrid rice breeding and seed production using a nuclear male sterility gene. Proceeding of the National Academy of Sciences of the United States of America 113: 14145-14150.) used HRM for screening OsNP1 mutants in rice and Shen et al. (2017Shen H, Strunks GD, Klemann BJ, Hooykaas PJ and Pater DS (2017) CRISPR/Cas9-induced double-strand break repair in Arabidopsis nonhomologous end -joining mutants.G3-Genes Genomes Genetics 7: 193-202.) for identifying mutants that escaped restriction enzyme analysis in Arabidopsis. Therefore, our study may bring more attention to this method for its use in future CRISPR/Cas9 projects.
In summary, the present study demonstrates that HRM analysis is a suitable method for detection and genotyping of mutations generated by CRISPR/Cas9, and recommends its use as a tool for CRISPR/Cas9-based studies and breeding.
ACKNOWLEDGMENTS
This research was supported by grants from the Ministry of Science and Technology, China (No.: SQ2015IM3600010) and the Fundamental Research Funds for the Central Universities (2016XZZX001-09).
REFERENCES
- Chang ZY, Chen ZF, Wang N, Xie G, Lu JW, Yan W, Zhou JL, Tang XY and Deng XW (2016) Construction of a male sterility system for hybrid rice breeding and seed production using a nuclear male sterility gene. Proceeding of the National Academy of Sciences of the United States of America 113: 14145-14150.
- Chen ST, Su YN, Ni YH, Hu WL, Lee NC, Chien YH, Chang CC, Chen HL and Chang MH (2012) Diagnosis of neonatal intrahepatic cholestasis caused by citrin deficiency using high-resolution melting analysis and a clinical scoring system. Journal of Pediatrics 161: 626-631.
- Cho SW, Kim S, Kim JM and Kim JS (2013) Targeted genome engineering in human cells with the Cas RNA-guided endonuclease. Nature Biotechnology 31: 230-232.
- Dahlem TJ, Hoshijima K, Jurynec MJ, Gunther D, Starker CG, Locke AS, Weis AM, Voytas DF and Grunwald DJ (2012) Simple methods for generating and detecting locus-specific mutations induced with TALENs in the zebrafish genome. PLoS Genetics 8: e1002861. [doi:10.1371/journal.pgen.1002861]
» https://doi.org/e1002861. [doi:10.1371/journal.pgen.1002861] - Fauser F, Schiml S and Puchta H (2014) Both CRISPR/Cas-based nucleases and nickases can be used efficiently for genome engineering in Arabidospsis thaliana. Plant Journal 79: 348-359.
- Feng Z, Mao Y, Xu N, Zhang B, Wei P, Yang DL, Wang Z, Zhang Z, Zheng R, Yang L, Zeng L, Liu X and Zhu JK (2014) Multigeneration analysis reveals the inheritance, specificity, and patterns of CRISPR/Cas-induced gene modifications in Arabidopsis. Proceeding of the National Academy of Sciences of the United States of America 111: 4632-4637.
- Hadden JM, Déclais AC, Carr SB, Lilley DMJ and Phillips SEV (2007) The structural basis of Holliday junction resolution by T7 endonuclease I. Nature 449: 621-624.
- Hofinger BJ, Jing HC, Kosack KEH and Kanyuka K (2009) High-resolution melting analysis of cDNA-derived PCR amplicons for rapid and cost-effective identification of novel alleles in barley. Theoretical and Applied Genetics 119: 851-865.
- Jiang W, Zhou H, Bi H, Fromm M, Yang B and Weeks DP (2013) Demonstration of CRISPR/Cas/sgRNA-mediated targeted gene modification in Arabidopsis, tobacco, sorghum and rice. Nucleic Acids Research 41: e188. doi:10.1093/nar/gkt780.
» https://doi.org/e188. doi:10.1093/nar/gkt780 - Koeyer DD, Douglass K, Murphy A, Whitney S, Nolan L, Song Y and Jong WD (2010) Application of high-resolution DNA melting for genotyping and variant scanning of diploid and autotetraploid potato. Molecular Breeding 25: 67-90.
- Li JF, Norville JE, Aach J, McCormack M, Zhang DD, Bush J, Church GM and Sheen J (2013) Multiplex and homologous recombination-mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas. Nature Biotechnology 31: 688-691.
- Li MR, Li XX, Zhou ZJ, Wu PZ, Fang MC, Pan XP, Lin QP, Luo WB, Wu GJ and Li HQ (2016) Reassessment of the four yield-related genes Gn1a, DEP1, GS3, and IPA1 in rice using a CRISPR/Cas system. Frontiers in Plant Science 7: 377. doi: 10.3389/fpls.2016.00377.
» https://doi.org/10.3389/fpls.2016.00377. - Liu WZ, Xie XR, Ma XL, Li J, Chen JH and Liu YG (2015) DSDecode: a web-based tool for decoding of sequencing chromatograms for genotyping of targeted mutations. Molecular Plant 8: 1431-1433.
- Lochlainn SO, Amoah S, Graham NS, Alamer K, Rios JJ, Kurup S, Stoute A, Hammond JP, Østergaard L, King KJ, White PJ and Broadley MR (2011) High resolution melt (HRM) analysis is an efficient tool to genotype EMS mutants in complex crop genomes. Plant Methods 7: 43. doi:10.1186/1746-4811-7-43.
» https://doi.org/10.1186/1746-4811-7-43 - Luo M, Gilbert B and Ayliffe M (2016) Applications of CRISPR/Cas technology for targeted mutagenesis, gene replacement and stacking of genes in higher plants. Plant Cell Reports 7: 1439-1450.
- Ma X, Zhu Q, Chen Y and Liu YG (2016) CRISPR/Cas platforms for genome editing in plants: developments and applications. Molecular Plant 9: 961-974.
- Ma XL, Zhang QY, Zhu QL, Liu W, Chen Y, Qiu R, Wang B, Yang ZF, Li HY, Lin Y, Xie YY, Shen RX, Chen SF, Wang Z, Chen YL, Guo JX, Chen LT, Zhao XC, Dong ZC and Liu YG (2015) A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Molecular Plant 8: 1274-1284.
- Mao Y, Zhang H, Xu N, Zhang B and Gao F (2013) Application of the CRISPR/Cas system for efficient genome engineering in plants. Molecular Plant 6: 691-693.
- Miao J, Guo DS, Zhang JZ, Huang QP, Qin GJ, Zhang X, Wan JM, Gu HY, Qu LJ (2013) Targeted mutagenesis in rice using CRISPR-Cas system. Cell Research 23: 1233-1236.
- Montgomery J, Wittwer CT, Palais R and Zhou L (2007) Simultaneous mutation scanning and genotyping by high-resolution DNA melting analysis. Nature Protocols 2: 59-66.
- Nekrasov V, Staskawicz B, Weigel D, Jones JD and Kamoun S (2013) Targeted mutagenesis in the model plant Nicotiana benthamiana using Cas RNA-guided endonuclease. Nature Biotechnology 31: 691-693.
- Pan G, Zhang X, Liu K, Zhang J, Wu X, Zhu J and Tu J (2006) Map-based cloning of a novel rice cytochrome p450 gene cyp81a6, that confers resistance to two different classes of herbicides. Plant Molecular Biology 61: 933-943.
- Shan Q, Wang Y, Li J, Zhang Y, Chen K, Liang Z, Zhang K, Liu J, Xi JJ, Qiu JL and Gao C (2013) Targeted genome modification of crop plants using a CRISPR/Cas system. Nature Biotechnology 31: 686-688.
- Shen H, Strunks GD, Klemann BJ, Hooykaas PJ and Pater DS (2017) CRISPR/Cas9-induced double-strand break repair in Arabidopsis nonhomologous end -joining mutants.G3-Genes Genomes Genetics 7: 193-202.
- Simko I (2016) High-resolution DNA melting analysis in plant research. Trends in Plant Science 21: 528-537.
- Tan YY, Yu XM, Shu QY, Zhang HL, Wang SG, Yuan FJ and Shi CH (2016) Development of an HRM-based, safe and high-throughput genotyping system for two low phytic acid, mutations in soybean. Molecular Breeding 36: 1-9.
- Tan YY, Fu HW, Zhao HJ, Lu S, Fu JJ, Li YF, Cui HR and Shu QY (2013) Functional molecular markers and high-resolution melting curve analysis of low phytic acid mutations for marker-assisted selection in rice. Molecular Breeding 31: 517-528.
- Uraguchi S, Kamiya T, Sakamoto T, Kasai K, Sato Y, Nagamura Y, Yoshida A, Kyozuka J, Ishikawa S and Fujiwara T (2011) Low-affinity cation transporter (oslct1) regulates cadmium transport into rice grains. Proceeding of the National Academy of Sciences of the United States of America 108: 20959-20964.
- Wang C, Shen L, Fu YP, Yan CJ and Wang KJ (2015) A simple CRISPR/Cas system for multiplex genome editing in rice. Journal of Genetics and Genomics 42: 703-706.
- Xu RF, Li H, Qin RY, Wang L, Li L, Wei PC and Yang JB (2014) Gene targeting using the Agrobacterium tumefaciens-mediated CRISPR-Cas system in rice. Rice 7: 5. doi:10.1186/s12284-014-0005-6.
» https://doi.org/10.1186/s12284-014-0005-6 - Zheng XL, Yang SX, Zhang DW, Zhong ZH, Xu T, Deng KJ, Zhou JP, Qi YP and Zhang Y (2016) Effective screen of CRISPR/Cas-induced mutants in rice by single-strand conformation polymorphism. Plant Cell Reports 35: 1545-1554.
- Zhang HL, Huang JZ, Chen XY, Tan YY and Shu QY (2014) Competitive amplification of differentially melting amplicons facilitates efficient genotyping of photoperiod- and temperature-sensitive genic male sterility in rice. Molecular Breeding 34: 1765-1776.
Publication Dates
-
Publication in this collection
Apr-Jun 2018 -
Date of issue
Apr 2018
History
-
Received
27 Sept 2017 -
Accepted
28 Nov 2017