Abstracts
Duplicates are common in germplasm banks and their identification is needed to facilitate germplasm bank management and to reduce maintenance costs. The aim of this work was to identify duplicates of cassava from a germplasm bank in Eastern Amazon, which had been previously characterized both morphological and agronomically. In order to be genotyped with 15 microsatellite loci, 36 accessions were selected. These accessions were classified into 13 groups of similar morpho-agronomical characteristics. All loci were polymorphic, and 75 alleles were identified, with an average of five alleles per loci and H E = 0.66. There were determined 34 pairs of genotypes with identical multiloci profiles and the probability of genetic identity was 1.1x10-12 with probability of exclusion of 99.9999%. Among these duplicates, there are accessions sampled on different years and places, but with different names and accessions with the same name sampled in different places and years. The study identified genotypes that are grown in different places and that have been maintained over the years by local farmers.
Manihot esculenta; germplasm; molecular markers
Duplicatas costumam ocorrer em bancos de germoplasma e a sua identificação é necessária para facilitar o manejo dos bancos ativos de germoplasma (BAGs) e diminuir custos de manutenção. O objetivo deste trabalho foi identificar duplicatas de mandioca determinadas previamente pela caracterização morfo-agronômica, em um BAG da Amazônia Oriental. Foram selecionados 36 acessos que se agrupavam em 13 grupos de similaridade morfo-agronômica para serem genotipados com 15 locos microssatélites. Todos os locos foram polimórficos, sendo obtidos 75 alelos, com média de cinco alelos por loco e HE = 0,66. Foram encontrados 34 pares de genótipos que apresentaram perfis multilocos idênticos e a probabilidade de identidade genética foi de 1,1x10-12 com probabilidade de exclusão de 99,9999%. Entre essas duplicatas, estão materiais coletados em épocas e locais diferentes, e com diferentes denominações e acessos com o mesmo nome coletados em diferentes locais e anos. O estudo identificou genótipos que vem sendo cultivados em diferentes locais e que vêm sendo mantidos pelos agricultores ao longo dos anos.
Manihot esculenta; germoplasma; marcadores moleculares
GENÉTICA
Elisa Ferreira MouraI; João Tomé de Farias NetoII; José Edson SampaioIII; Diehgo Tuloza da SilvaIV; Girena Fernandes RamalhoV
IEmbrapa Amazônia Oriental, Trav. Dr. Enéas Pinheiro s/n caixa postal 48, Belém, Pará CEP: 66095-100. E-mail: elisa.moura@embrapa.br
IIEmbrapa Amazônia Oriental, Trav. Dr. Enéas Pinheiro s/n caixa postal 48, Belém, Pará CEP: 66095-100. E-mail: joao.farias@embrapa.br
IIIEmbrapa Amazônia Oriental, Trav. Dr. Enéas Pinheiro s/n caixa postal 48, Belém, Pará CEP: 66095-100. E-mail: edson.sampaio@embrapa.br
IVUniversidade Federal do Pará, Campus Universitário do Guamá, Rua Augusto Corrêa,no. 1, Instituto de Ciências Biológicas - Campus Básico, CEP: 66075-110. E-mail: dbiotuloza@hotmail.com
VUniversidade Federal Rural da Amazônia, Universidade Federal Rural da Amazônia, Avenida Presidente Tancredo Neves, nº 2501, Bairro: Terra Firme Cep: 66.077-530 Caixa Postal: 917, Belém, Pará, Brasil. E-mail: girenaufpa@yahoo.com.br
ABSTRACT
Duplicates are common in germplasm banks and their identification is needed to facilitate germplasm bank management and to reduce maintenance costs. The aim of this work was to identify duplicates of cassava from a germplasm bank in Eastern Amazon, which had been previously characterized both morphological and agronomically. In order to be genotyped with 15 microsatellite loci, 36 accessions were selected. These accessions were classified into 13 groups of similar morpho-agronomical characteristics. All loci were polymorphic, and 75 alleles were identified, with an average of five alleles per loci and HE = 0.66. There were determined 34 pairs of genotypes with identical multiloci profiles and the probability of genetic identity was 1.1x10-12 with probability of exclusion of 99.9999%. Among these duplicates, there are accessions sampled on different years and places, but with different names and accessions with the same name sampled in different places and years. The study identified genotypes that are grown in different places and that have been maintained over the years by local farmers.
Keywords:Manihot esculenta; germplasm; molecular markers
RESUMO
Duplicatas costumam ocorrer em bancos de germoplasma e a sua identificação é necessária para facilitar o manejo dos bancos ativos de germoplasma (BAGs) e diminuir custos de manutenção. O objetivo deste trabalho foi identificar duplicatas de mandioca determinadas previamente pela caracterização morfo-agronômica, em um BAG da Amazônia Oriental. Foram selecionados 36 acessos que se agrupavam em 13 grupos de similaridade morfo-agronômica para serem genotipados com 15 locos microssatélites. Todos os locos foram polimórficos, sendo obtidos 75 alelos, com média de cinco alelos por loco e HE = 0,66. Foram encontrados 34 pares de genótipos que apresentaram perfis multilocos idênticos e a probabilidade de identidade genética foi de 1,1x10-12 com probabilidade de exclusão de 99,9999%. Entre essas duplicatas, estão materiais coletados em épocas e locais diferentes, e com diferentes denominações e acessos com o mesmo nome coletados em diferentes locais e anos. O estudo identificou genótipos que vem sendo cultivados em diferentes locais e que vêm sendo mantidos pelos agricultores ao longo dos anos.
Palavras-chave:Manihot esculenta; germoplasma; marcadores moleculares
INTRODUCTION
Cassava (Manihot esculenta) is an important source of carbohydrates for more than 800 million people around the world, especially in underdeveloped countries. There are reasons to believe that the species domestication has occurred towards the Southern border of the Amazon Basin (Olsen and Schaal 2001; Leotárd et al. 2009). Possibly the North region keeps a great portion of cassava genetic variation, specially due to the diversity of products that local farmers generate from this root.
In Brazil, cassava is grown almost all across its territorial extension, which means the country has genetic variability for adaptation to different environments and climatic conditions. This variability has been maintained on germplasm bank and collections established in different sites of the country. One of them is established on Pará State, the main producer of cassava in Brazil and also known for the diversity of products its population generates from cassava root and even from its leaves. The bases of this germplasm bank are genotypes sampled in properties of familiar farmers from Pará State. At the present, the germplasm bank maintains 470 accessions, including genotypes of 'sweet', 'bitter' and 'sugary'cassava. The accessions have been characterized according to 39 morpho-agronomical characters, established by Fukuda and Guevara (1998), which is followed by the majority of Brazilian germplasm collections and banks. Based on this characterization, some morpho-agronomical similarities have been observed among accessions, including ones with different names and sampled in different places. The identification of duplicates on germplasm banks is interesting to reduce its size, which facilitates its management. Also, it helps the genetic breeding program, since identical genotypes will not be chosen for field trial experiments or for controlled crosses.
The main procedure to identify duplicates is using molecular genotyping, since molecular markers represent a portion of the genome that does not suffer environmental influence. Microsatellite markers, due to high information generated per locus, have been used to identify duplicates on germplasm banks and populations of different species (Robichaud et al. 2006; Irish et al. 2010; van Treuren et al. 2010). Methods of identification that use multiloci profiles of microsatellites are more accurate than the ones based on dominant markers, since genotypes can present a complete coincidence and the probability of casual coincidence can be measured based on allelic frequencies (Paetkau and Strobeck 1994).
Thus, the aim of this work was to genotype cassava accessions previously identified as morpho-agronomically similar with microsatellite markers, in order to verify if they represent duplicates.
MATERIALS AND METHODS
It was selected 36 accessions from the Germplasm Bank of Embrapa Eastern Amazon, located in Belém, Pará, Brazil. The accessions were morphologically characterized according to Fukuda and Guevara (1998) and they showed similarity. These accessions were divided according to the morphological characterization in 13 groups (Table 1). When samples were being collected, it was decided to maintain the same name given by farmers. The selected accessions were composed of 16 samples of 'sweet' cassava, differentiated by the letter 'M' (as for 'macaxeira') and 20 accessions of 'bitter' cassava. On similarity groups where accessions came from the same locality, they were sampled on different farms. Figure 1 shows the geographical sample locations on the map. Genomic DNA was extracted according to the method of Doyle and Doyle (1990) with modifications. Leaves were macerated with liquid nitrogen and polivinilpirrolidone and 3 mL CTAB extraction buffer (2% CTAB, 5 M NaCl, 0.5 M EDTA, PVP, 1 M Tris-HCl and sterilized water) was added to the macerate. Then, it was homogenized and incubated in water bath at 65 oC for one hour. After that period, it was added cloroformium: isoamilic alcohol (24:1), the extract was homogenized and samples were centrifuged for 10 minutes at 10,000 rpm. It was added 3 mL 95% ethylic alcohol to the supernatant to precipitate the DNA and samples were centrifuged for 10 minutes at 10,000 rpm. After that phase, the precipitate was washed with 70% ethylic alcohol for 10 minutes and at 5,000 rpm. DNA samples were ressuspended with 300 mL TE buffer (10 mM Tris-HCl, 1 mM EDTA, pH 8.0) and RNAse. DNA was quantified in 1% agarose gel using samples of phage lambda DNA on different concentrations (50, 100 e 200 ng mL-1) as pattern. DNA was diluted to 10 ng mL-1. It was used 15 microsatellite primers (Table 2), developed by Chavarriaga-Aguire et al. (1998) and Mba et al. (2001). Polymerase chain reactions (PCR) were prepared for a final volume of 20 mL, containing 30 ng of genomic DNA, 50 mM of each triphosphate deoxiribonucleotides (dATP, dCTP, dGTP e dTTP), 0.1 mM of each pair of primer (forward and reverse), 10 mg mL-1 of BSA (bovine serum albumin), 0.6 units of Taq DNA polymerase (Invitrogen, Brazil) and 1X reaction buffer containing MgCl2 (1 mM) supplied by the manufacturer. PCR were performed on a thermocycler (Applied Byosystems, GeneAmp®, PCR Instrument System 9700, USA). The temperature cycling profile was: an initial denaturation step for 5 min at 94 oC, followed by 30 cycles of denaturation at 94 oC for 1 min, annealing at 55 oC to 59 oC (depending on the primer) for 2 min and primer extension at 72 oC for 2 min and a final extension cycle of 5 min at 72 oC. For primer SSRY82, the conditions of amplification were: an initial denaturation step for 5 min at 94 oC, followed by 35 cycles of DNA denaturation at 95 ºC for 30s, annealing at 55 ºC for 30 s and elongation at 72 ºC for 45s; after 35 cycles, there was a primer extension at 72 ºC for 10 min.
Amplification products were separated on vertical electrophoresis (Omniphor, HMEDI15, England), using 6% polyacrilamide gel. Gels were revealed with silver nitrate and scanned for image analyses. Gels were visually interpreted and each primer represented a locus and each band with different migration pattern was considered an allele. For the construction of the dendrogram, bands were analyzed as presence (1) and absence (0) for the 36 accessions. These data were used to construct a dendrogram based on the unweighted pair group method with arithmetic average (UPGMA) using the Jaccard similarity index. A bootstrap resampling method was performed to determine the robustness of the dendrogram, and 1,000 bootstrap replicates were obtained from the original data of 36 accessions. All calculations were performed using FreeTree 0.9.1.50 (Pavlicek et al. 1999) and the dendrogram was drawn using TreeView 1.6.6 (Page 2001). Genetic diversity parameters, such as the number of alleles per locus, allelic frequency, percent of polymorphic loci, observed average heterozygosity (HO), gene diversity (expected heterozygosity - HE) obtained per locus, and allelic frequencies per locus were estimated using GenAlEx 6.4.1 (Peakall and Smouse 2006). The probability of genetic identity and the probability of exclusion were obtained according to the method suggested by Paetkau and Strobeck (1994) and calculated with GenAlEx.
RESULTS
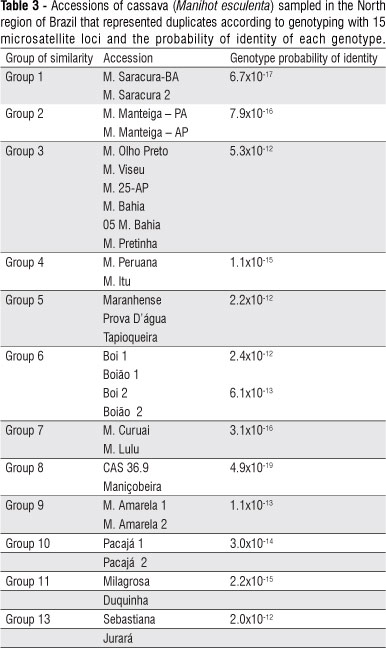
All microsatellite loci used were 100% polymorphic and generated 75 alleles, with an average of five alleles per locus (Table 2). The number of alleles varied from two (GAGG05 and SSRY102) to eight (SSRY04). The Jaccard coefficient of similarities was estimated among cassava accessions. It was identified 34 pairs of duplicates, which showed identical multiloci profiles and, consequently, genetic similarity = 1.0. The probability that two individuals share the same genetic profile by chance with the 15 loci used and the allelic frequencies obtained was 1.1x10-12. It means that the probability that they represent duplicates is 99.9999%. The accessions that correspond to duplicates are listed on Table 3, with the probability of occurrence of each genotype considering the allelic frequencies estimated for the 36 accessions. All probabilities of occurrence of genotypes were very low. For genotype of group 4 (Table 3), for example, an exact match between unrelated genotypes in a randomly mating population is one in hundreds of quintillions. Considering the 13 groups of morphological similarity, only three groups had individuals that were not identical. In group 4, the genotype 'Hamburguesa' was different from 'M. Itu' and 'M. Peruana'. In group 10, 'Pretinha' and 'Manivão' were not duplicates and were different from two accessions named 'Pacajá'. In group 12, the two accessions named 'Tumase' were different.
After the detection of duplicates, sample was reduced to 18 genotypes. Genetic diversity parameters were estimated after the removal of duplicates, to avoid overestimatives (Table 2). Average HE was 0.66 and HE per locus varied from 0.80 (SSRY04) to 0.44 (GAGG05 and SSRY102). Average HO was 0.61 and HO per locus varied from 1.00 (GA21) to 0.28 (SSRY164). Total coefficient of endogamy was 0.08.
The dendrogram with bootstrap calculations clearly showed the grouping of the duplicates (Figure 2). The dendrogram showed that accessions did not cluster according to the sample location or according to 'sweet' or 'bitter' type.
The analysis showed three different situations: confirmation of materials with the same name that share the same genotype; materials with the same name that are different and genotypes with different names spread through different places. Confirmation of materials with the same name and the same genotype occurred even for accessions sampled in different places. Among the genotypes identified as duplicates, there are accessions sampled in very distant periods, such as genotypes 'Sebastiana', sampled in 2005, and 'Jurará', sampled in 1947, both in Pará State (Table 1). This situation occurs for the duplicates of groups 1, 2, 3, 4, 5, 6 and 13 (Table 1). Also, the same genotype was sampled in different places in groups 2, 3, 4, 5, 6, 8, 10, 11 and 13.
DISCUSSION
We have shown in this study that some cassava genotypes are spread through a large area in the North region, more than 99.9999% confident. The sample size could be considered small to estimate the allelic frequencies, which could lead to an underestimation. However, the HE (Table 2) obtained were similar to the ones obtained in other analyses done with a higher number of samples, when primers GA21, GA126, GA131 and GA136 were compared (Elias et al. 2001; Siqueira et al. 2009; Siqueira et al. 2010).
HO values, although high, were comparable to the values obtained with cassava sampled in several parts of the world (Elias et al. 2001; Fregene et al. 2003). High levels of heterozygosis are characteristic of a plant with allogamous reproduction and asexual propagation.
The occurrence of the same genotypes in different places confirms the existence of exchanges of propagative material of cassava genotypes among farmers in the North region. This may have contributed to the lack of correlation between genetic grouping in the dendrogram and geographical places, as observed for cassava in other geographic regions (Elias et al. 2004; Siqueira et al. 2009). The different denominations that farmers attribute to cassava landraces seem to be common. Surveys of cassava names in different countries have identified that the nomination of the landraces is often given separately by farmers, even from the same place or community (Salick et al. 1997; Mkumbira et al. 2003). However, there were varieties that kept their names over the years and in different places, such as 'Pacajá', 'M. Manteiga' and 'M. Saracura'. This may be related to the importance and value of the landrace among farmers of a more extensive area. Otherwise, the accessions named 'Tumase' were very genetically different, evidencing another common situation among cassava landraces: the coincidence of names between genetically distinct genotypes.
The identification of duplicates on germplasm banks of vegetatively propagated plants seems to be common (Irish et al. 2010; van Treuren et al. 2010). This may be due to usual exchanges of propagules among farmers of different regions, especially when the species has economical importance. In the new place, that genotype may receive a new name, causing certain confusion on samples and maintenance of accessions on germplasm banks. However, as the species are vegetatively propagated, there may be spontaneous mutations in a single gene that generates an interesting variation. Thus, although genotyping with molecular markers can identify duplicates, the accessions genetically identical may not be discarded immediately, not before characterization is complete. Chemical characterization and analysis of genotypes resistance/tolerance to diseases may reveal differences among individuals with the same multiloci profile that may be due to a single mutation. However, duplicates identification is important to cluster accessions and to avoid crossings between them.
This study showed the occurrence of duplicates in a cassava germplasm bank composed mainly of landraces sampled on the North region of Brazil and evidenced specific genotypes that are spread through different sites of the region. The identification of these duplicates will be useful to the germplasm bank management and also in the selection of accessions for field trial experiments.
The resampling of the same genotype over many years in different places of the North region of Brazil may reflect the importance that some genotypes have for the farmers. These genotypes may be the most productive or have an important characteristic for food. Also, it emphasizes the importance that local farmers have as keepers of the genetic variation of cassava.
CONCLUSIONS
The study showed that the availability of polymorphic microsatellite loci for cassava allowed identification of duplicates in a germplasm bank composed of accessions sampled mainly in the North region of Brazil. Identification of duplicates was 99.9999% confident. It showed that molecular characterization with highly informative markers is important to determine duplications in germplasm banks of vegetatively propagated species, such as cassava, and to reduce numbers of accessions in the bank, helping in its management. It also showed the great dispersion that some genotypes of cassava have in the North region, confirming that local farmers are keepers of genetic variability of cassava.
ACKNOWLEDGMENTS
The authors thank FAPESPA, CNPq and Embrapa for financial support and scholarship grants.
Recebido em: 25/06/2012
Aceito em: 04/02/2013
- Chavarriaga-Aguirre, P.P.; Maya, M.M.; Bonierbale, M.W.; Kresovich, S.; Fregene, M.A.; Tohme. J.; Kochert, G. 1998. Microsatellites in cassava (Manihot esculenta Crantz): discovery, inheritance and variability. Theoretical and Applied Genetics, 97: 493-501.
- Doyle, J.J.; Doyle, J.L. 1990. Isolation of plant DNA from fresh tissue. Focus, 12: 13-15.
- Elias, M.; Muhlen, G.S.; McKey, D.; Roa, A.C.; Tohme, J. 2004. Genetic diversity of traditional South American landraces of cassava (Manihot esculenta Crantz): an analysis using microsatellites. Economic Botany, 52: 242-25.
- Elias, M.; Penet, L.; Vindry, P.; McKey, D.; Panaud, O.; Robert, T. 2001. Unmanaged sexual reproduction and the dynamics of genetic diversity of a vegetatively propagated crop plant, cassava (Manihot esculenta Crantz) in a traditional farming system. Molecular Ecology, 10: 1895-1907.
- Fregene, M.; Suarez, M.; Mkumbira, J.; Kulembeka, H.; Ndedya, E.; Kulaya, A.; Mitchel, S.; Gullberg, U.; Rosling, H.; Dixon, A.G.O.; Dean, R.; Kresovich, S. 2003. Simple sequence repeat marker diversity in cassava landraces: genetic diversity and differentiation in an asexually propagated crop. Theoretical and Applied Genetics, 107:1083-1093.
- Fukuda, W.M.G.; Guevara, C.L. 1998. Descritores morfológicos e agronômicos para a caracterização de mandioca (Manihot esculenta Crantz) Embrapa, Cruz das Almas, BA, Brasil. 38p.
- Irish, B.M.; Goenaga, R.; Zhang, D.; Schnell, R.; Brown, J.S.; Motamayor, J.C. 2010. Microsatellite fingerprinting of the USDA-ARS tropical agriculture research station cacao (Theobroma cacao L) germplasm collection. Crop Science, 50: 656-667.
- Léotard, G.; Duputié, A.; Kjellberg, F.; Douzery, E.J.P.; Debain, C.; Granville, J.J.; McKey, D. 2009. Phylogeography and the origin of cassava: new insights from the Northern rim of the Amazonian basin. Molecular Phylogenetics and Evolution, 53: 329-334.
- Mba, R.E.C.; Stephenson, P.; Edwards, K.; Melzer, S.; Nkumbira, J.; Gullberg; U.; Ape, K.; Gale, M.; Tohme, J.; Fregene, M. 2001. Simple sequence repeats (SSR) markers survey of the cassava (Manihot esculenta Crantz) genome: towards an SSR-based molecular genetic map. Theoretical and Applied Genetics, 102: 21-31.
- Mkumbira, J.; Chiwona-Karltun, L.; Lagercrantz, U.; Mahungu, N.M.; Saka, J.; Mhone, A.; Bokanga, M.; Brimer, L.; Gullberg, U.; Roling, H. 2003. Classification of cassava into 'bitter' and 'cool' in Malawi: from farmers perception to characterization by molecular markers. Euphytica, 132: 7 - 22.
- Olsen, K.M.; Schaal, B.A. 2001. Microsatellite variation in cassava (Manihot esculenta, Euphorbiaceae) and its wild relatives: further evidence for a Southern Amazonian origin of domestication. American Journal of Botany, 88:131-142.
- Page, R.D.M. TreeView. Version 1.6.6. Glasgow: University of Glasgow, 2001. (http://taxonomy.zoology.gla.ac.uk/rod/treeview/treeview_manual.htmll) Accessed on: 11/01/2013.
- Pavlícek, A.; Hrdá, S.; Flegr, J. 1999. FreeTree - freeware program for construction of phylogenetic trees on the basis of distance data and bootstrap/jackknife analysis of the tree robustness. Application in the RAPD analysis of genus Frenkelia Folia Biologica 45: 97-99.
- Paetkau, D.; Strobeck, C. 1994. Microsatellite analysis of genetic variation in black bear populations. Molecular Ecology 3: 489 - 495.
- Peakall, R.; Smouse, P.E. 2006. GENALEX 6: genetic analysis in excel. Population genetic software for teaching and research. Molecular Ecology Notes 6:288 - 295.
- Robichaud, R.; Glaubitz, J.C.; Rhodes Jr., O.E.; Woeste, K. 2006. A robust set of black walnut microsatellites for parentage and clonal identification. New Forests, 32: 179-196.
- Salick, J.; Cellinese, N.; Knapp, S. 1997. Indigenous diversity of cassava: generation, maintenance, use and loss among the Amuesha, Peruvian upper Amazon. Economic Botany, 51: 7 - 17.
- Siqueira, M.V.B.M.; Queiroz-Silva, J.R.; Bressan, E.A.; Borges, A.; Pereira, K.J.C.; Pinto, J.G.; Veasey, E.A. 2009. Genetic characterization of cassava (Manihot esculenta) landraces in Brazil assessed with simple sequence repeats. Genetics and Molecular Biology, 32: 104-110.
- Siqueira, M.V.B.M.; Pinheiro, T.T.; Borges, A.; Valle, T.L.; Zatarim, M.; Veasey, E.A. 2010. Microsatellite polymorphisms in cassava landraces from the Cerrado Biome, Mato Grosso do Sul, Brazil. Biochemical Genetics, 48: 879-895.
- Van Treuren, R.; Kemp, H.; Ernsting, G.; Jongejans, B.; Houtman, H.; Visser, L. 2010. Microsatellite genotyping of apple (Malus x domestica Borkh) genetic resources in the Netherlands: application in collection management and variety identification. Genetic Resources and Crop Evolution, 57: 853-865.
Identification of duplicates of cassava accessions sampled on the North Region of Brazil using microsatellite markers
Publication Dates
-
Publication in this collection
09 Aug 2013 -
Date of issue
Dec 2013
History
-
Received
25 June 2012 -
Accepted
04 Feb 2013