Abstracts
This study evaluated the expression of CD14, toll-like receptor (TLR) 2 and TLR4 on the surface of milk neutrophils in bovine mammary glands infected with Corynebacterium bovis. Here, we used 23 culture-negative control quarters with no abnormal secretion on the strip cup test and milk somatic cell count lower than 1x105 cells/mL, and 14 C. bovis infected quarters. The identification of neutrophils, as well as, the percentage of neutrophils that expressed CD14, TLR2 and TLR4 were analyzed by flow cytometry using monoclonal antibodies. The present study encountered no significant difference in the percentages of milk neutrophils that expressed TLR2 and TLR4 or in the expression of TLR4 by milk neutrophils. Conversely, a lower median fluorescence intensity of TLR2 in milk neutrophils was observed in C. bovis-infected quarters. The percentage of neutrophils that expressed CD14 and the median fluorescence intensity of CD14 in milk neutrophils was also lower in C. bovis-infected quarters.
Immune response; pattern recognition receptor; mastitis; milk; polymorphonuclear neutrophil leukocyte
O presente estudo objetivou avaliar alterações na expressão de CD14, e dos receptores do tipo toll (TLR) 2 e 4 na superfície de neutrófilos lácteos provenientes de glândulas mamárias infectadas por Corynebacterium bovis. O presente estudo utilizou 23 quartos negativos no exame bacteriológico, sem alterações na prova de fundo escuro e com contagem automática de células somáticas menor que 1 x105 células/mL, e 14 quartos mamários infectados por C. bovis A identificação de neutrófilos, assim como a porcentagem de neutrófilos lácteos que expressaram CD14, TLR2 e 4 foram avaliadas por citometria de fluxo utilizando anticorpos monoclonais. A porcentagem de neutrófilos que expressaram TLR2 e TLR4 nos quartos mamários infectados por C. bovis não diferiu dos quartos mamários sadios, assim como na expressão de TLR4. No entanto, a intensidade de fluorescência do TLR2 na superfície dos neutrófilos foi menor nos quartos mamários infectados por C. bovis. A porcentagem de neutrófilos que expressaram CD14 e a intensidade de fluorescência da molécula de CD14 foi menor na superfície dos neutrófilos lácteos dos quartos infectados por C. bovis.
Resposta imune; receptores de reconhecimento padrão; mastite; leite; leucócitos polimorfonucleares.
Introduction
Corynebacterium bovis is one of the most common bacteria isolated from milk samples that are submitted for the identification of intramammary infection (IMI)-causing pathogens around the world (Haltia et al. 2006Haltia L., Honkanen-buzalski T., Spiridonova I., Olkonen A. & Myllys V. 2006. A study of bovine mastitis, milking procedures and management practices on 25 Estonian dairy herds. Acta Vet. Scand. 48:22., Tenhagen et al. 2006Tenhagen B.A., Köster G., Wallmann J. & Heuwieser W. 2006. Prevalence of mastitis pathogens and their resistance against antimicrobial agents in dairy cows in Brandenburg, Germany. J. Dairy Sci.89:2542-2551., Souza et al. 2009Souza G.N., Brito J.R.F., Moreira E.C., Brito M.A.V.P. & Silva M.V.G.B. 2009. Variação da contagem de células somáticas em vacas leiteiras de acordo com patógenos da mastite. Arq. Bras. Med. Vet. Zoot. 61:1015-1020.). Despite its high prevalence, C. bovis is regarded as a minor pathogen with limited clinical significance (Huxley et al. 2004Huxley J.N., Helps C.R. & Bradley A.J. 2004. Identification of Corynebacterium bovis by endonuclease restriction analysis of the 16S rRNA gene sequence. J. Dairy Sci.87:38-45.) and has previously been regarded as a commensal organism (Brooks & Barnum 1984Brooks B.W. & Barnum D.A. 1984. The susceptibility of bovine udder quarters colonized with Corynebacterium bovis to experimental infection with Staphylococcus aureus or Streptococcus agalactiae. Can. J. Comp. Med. 48:146-150.). This bacterium colonizes the teat canal region but can also be isolated from the teat cistern, gland cistern and mammary parenchyma (Benites et al. 2003Benites N.R., Melville P.A. & Costa E.O. 2003. Evaluation of the microbiological status of milk and various structures in mammary glands of from naturally infected dairy cows. Trop. Anim. Health Prod. 35:301-307.). Furthermore, although the mean milk somatic cell count (MSCC) recorded for milk samples from which C. bovis is isolated is relatively low, the value is higher than the mean MSCC recorded for samples yielding no growth (Djabri et al. 2002Djabri B., Bareille N., Beaudeau F. & Seegers H. 2002. Quarter milk somatic cell count in infected dairy cows: a meta-analysis. Vet. Res. 33:334-357.).
C. bovis is of interest to mastitis researchers because of the observation that quarters infected with this bacterium are significantly less likely to become infected with other, more pathogenic bacteria (Brooks & Barnum 1984Brooks B.W. & Barnum D.A. 1984. The susceptibility of bovine udder quarters colonized with Corynebacterium bovis to experimental infection with Staphylococcus aureus or Streptococcus agalactiae. Can. J. Comp. Med. 48:146-150., Sordillo et al. 1988Sordillo L.M., Oliver S.P., Guidry A.J. & Dermody J.T. 1988. Humoral immune response of bovine mammary glands colonized with Corynebacterium bovis: enumeration of plasma cell populations in tissue and immunoglobulin concentrations in milk. J. Vet. Med. B 35:617-627., Lam et al. 1997Lam T.J., Schukken Y.H., Van Vliet J.H., Grommer F.J., Tielen M.J. & Brand A. 1997. Effect of natural infection with minor pathogens on susceptibility to natural infection with major pathogens in the bovine mammary gland. Am. J. Vet. Res.58:17-22., Blagitz et al. 2013Blagitz M.G., Souza F.N., Batista C.F., Santos B.P., Parra A.C., Azevedo L.F.F., Melville P.A., Benites N.R.& Della Libera A.M.M.P.2013. Function of milk polymorphonuclear neutrophil leukocytes in bovine mammary glands infected with Corynebacterium bovis. J. Dairy Sci. 96:3750-3757.), although no consensus has been reached (Zadoks et al. 2001Zadoks R.N., Allore H.G., Barkema H.W., Sampimon O.C., Wellenberg G.J., Gröhn Y.T. & Schukken Y.H.2001. Cow- and quarter-level risk factors for Streptococcus uberis and Staphylococcus aureus mastitis. J. Dairy Sci.84:2649-2663., Reyher et al. 2012Reyher K.K., Dohoo I.R., Scholl D.T. & Keefe G.P. 2012. Evaluation of minor pathogens intramammary infection, susceptibility parameters, and somatic cell counts on the development of new intramammary infections with major mastitis pathogens. J. Dairy Sci.95:3766-3780.). With this concept in mind, it should be noted that polymorphonuclear neutrophil leukocytes (PMNLs) provide the first cellular line of defense against invading mastitis pathogens and are the main source of resistance during the early stages of bacterial invasion (Schukken et al. 2011Schukken Y.H., Güntler J., Fitzpatrick J., Fontaine M.C., Goetza L., Holst O., Leigh J., Petzl W., Schuberth H.-J., Sipka A., Smith D.G.E., Quesnell R., Watts J., Yancey R., Zerbe H., Gurjar A., Zadoks R.N., Seyfert H.-M. & Members of Pfizer mastitis research consortium. 2011. Host-response patterns of intramammary infections in dairy cows. Vet. Immunol. Immunopathol. 144:270-289., Souza et al. 2012Souza F.N., Ramos Sanchez E.M., Heinemann M.B., Gidlund M.A., Reis L.C., Blagitz M.G., Della Libera A.M.M.P.& Cerqueira M.M.O.P. 2012. The innate immunity in bovine mastitis: the role of pattern-recognition receptors.. Am J. Immunol.8:166-178.).
The recognition of microbial pathogens is an essential element of the initiation of innate immune responses, such as inflammation, and is mediated by germline-encoded pattern recognition receptors (PRRs) that recognize molecular structures that are broadly shared by pathogens, known as pathogen-associated molecular patterns (PAMPs). Upon PAMP recognition, PRRs initiate a series of signaling programs that execute the first line of host defensive responses that are necessary for killing infectious microbes. Toll-like receptors (TLRs) are the best-characterized PRRs and recognize a wide range of PAMPs. These receptors are either expressed on the cell surface or associated with intracellular vesicles (Schukken et al. 2011Schukken Y.H., Güntler J., Fitzpatrick J., Fontaine M.C., Goetza L., Holst O., Leigh J., Petzl W., Schuberth H.-J., Sipka A., Smith D.G.E., Quesnell R., Watts J., Yancey R., Zerbe H., Gurjar A., Zadoks R.N., Seyfert H.-M. & Members of Pfizer mastitis research consortium. 2011. Host-response patterns of intramammary infections in dairy cows. Vet. Immunol. Immunopathol. 144:270-289., Souza et al. 2012Souza F.N., Ramos Sanchez E.M., Heinemann M.B., Gidlund M.A., Reis L.C., Blagitz M.G., Della Libera A.M.M.P.& Cerqueira M.M.O.P. 2012. The innate immunity in bovine mastitis: the role of pattern-recognition receptors.. Am J. Immunol.8:166-178.). In most cases, fully efficient microbial recognition by TLR2 and TLR4 requires the critical activity of a co-receptor, CD14. In fact, in the absence of CD14, cellular responses to most microbial components, activated via TLR2 or TLR4, are particularly low (LeBouder et al. 2003LeBouder E., Rey-Nores J.E., Rushmere N.K., Grigorov M., Lawn S.D., Affolter M., Griffin G.E., Ferrara P., Schiffrin E.J., Morgan B.P. & Labéta M.O. 2003. Soluble forms of toll-like receptors (TLR)2 capable of modulating TLR2 signaling are present in human plasma and breast milk. J. Immunol.171:6680-6689.). CD14 is also expressed in milk neutrophils (Paape et al. 1996Paape M.J., Liluis E.-M., Wiitanen P.A., Kontio M.P. & Miller R.H. 1996. Intramammary defense against infections induced by Escherichia coli in cows. Am J Vet Res. 57:477-482.). Nonetheless, to the best of our knowledge, this is the first study that compared the expression of CD14, TLR2 and TLR4 in milk PMNLs in C. bovis-infected and uninfected quarters. Thus, the aim of this study was to investigate the expression of CD14, TLR2 and TLR4 on the surface of milk neutrophils in C. bovis-infected quarters.
Materials and Methods
The present study used 37 mammary quarters from 13 clinically healthy Holstein dairy cows that were collected at different lactation stages from a commercial herd. From this sample, we selected 23 culture-negative control quarters from 10 dairy cows with no abnormal secretions in the strip cup test and MSCC lower than 1 × 105 cells/mL, which is the threshold for the MSCC described by Bansal et al. (2005)Bansal B.K., Hamann J., Grabowskit N.T. & Singh B. 2005. Variation in the composition of selected milk fraction samples from healthy and mastitic quarters, and its significance for mastitis diagnosis. J. Dairy Res. 72:144-152. for uninfected quarters. Additionally, 14 C. bovis-infected quarters from eight dairy cows were included.
First, the strip cup test was performed to identify the presence of clots, flakes or otherwise obviously abnormal secretions. Predipping was then performed, and the first three milk streams were discarded. Teat ends were scrubbed with cotton moistened with 70% ethanol, and single milk samples from individual mammary quarters were aseptically collected into sterile vials for bacteriological analysis. Lastly, milk samples were collected for a somatic cell count (SCC) and an evaluation of the expression of CD14, TLR2 and TLR4 by milk neutrophils. We emphasize that this research complied with the Ethical Principles in Animal Research and was approved by the Bioethics Commission.
A bacterial analysis was performed by culturing 0.01 mL of each milk-quarter sample on 5% ovine blood agar plates. The plates were incubated for 72 hours at 37°C, followed by Gram staining, observation of colony morphology and biochemical testing (Oliver et al. 2004Oliver S.P., Lewis M.J., Gillespie B.E., Dowlen H.H., Jaenicke E.C. & Roberts R.K. 2004. Microbiological Procedures for the Diagnosis of Bovine Udder Infection and Determination of Milk Quality. 4th ed. National Mastitis Council, Verona. 47p.). A milk sample was considered culture positive when the growth of ≥4 pure C. bovis colonies was detected. Samples yielding no bacterial growth were regarded as culture negative.
The SCC measurements of milk samples were performed using an automated fluorescent microscope-based somatic cell counter (Somacount 300, Bentley Instruments(r), Chaska, USA).
The separation of milk cells was performed as described by Koess & Hamann (2008)Koess C. & Hamann J.2008. Detection of mastitis in the bovine mammary gland by flow cytometry at early stages. J. Dairy Res.75:225-232.. Briefly, 1 L of milk was diluted with 1 L of phosphate-buffered saline (PBS; pH 7.4; 1.06mM Na2HPO4, 155.17mM NaCl, 2.97mM Na2HPO4.7H2O). After centrifugation at 1,000xg for 15 min, the cream layer and the supernatant were discarded. The cell pellet at the bottom of the container was then washed once with 30mL of PBS and centrifuged at 400xg for 10 min. The cells were placed in 1 mL of RPMI-1640 nutrition media (R7638; Sigma-Aldrich, USA) supplemented with 10% fetal bovine serum (Cultilab, Brazil) and counted in a Neubauer chamber. The cell viability was first evaluated by trypan blue exclusion. The milk cells were then diluted with the nutrition media and 10% fetal bovine serum to a concentration of 2×106 viable cells/mL.
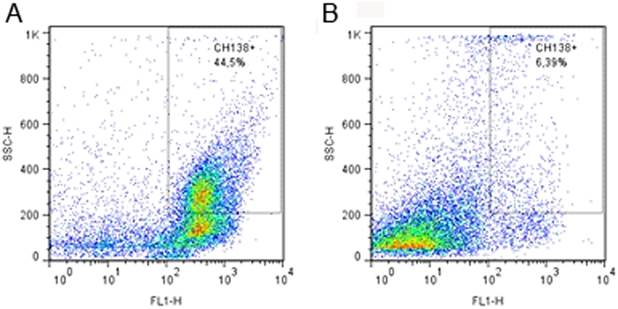
Milk neutrophils were differentiated from other cells by indirect fluorescent labeling using 1μL of the primary unlabeled monoclonal antibody (mAb) anti-bovine granulocyte (CH138A; VMRD Pullman Inc. Corp(r), Pullman, USA) that was incubated for 30 min at room temperature. Next, 1mL of PBS was added to the cell suspension, which was centrifuged at 400xg for 8 min. Lastly, a labeled secondary antibody was added and incubated for 30 min at room temperature in the dark to visualize CH138A binding. Neutrophils were identified using flow cytometry based on their cytoplasmic granularity and CH138A positivity (Fig.1), as previously described by Piepers et al. (2009)Piepers S., De Vliegher S., Demeyere K., Lamrecht B.N., Kruif A., Meyer E. & Opsomer G. 2009. Technical note: Flow cytometric identification of bovine milk neutrophils and simultaneous quantification of their viability. J. Dairy Sci.92:626-631. and Blagitz et al. (2013)Blagitz M.G., Souza F.N., Batista C.F., Santos B.P., Parra A.C., Azevedo L.F.F., Melville P.A., Benites N.R.& Della Libera A.M.M.P.2013. Function of milk polymorphonuclear neutrophil leukocytes in bovine mammary glands infected with Corynebacterium bovis. J. Dairy Sci. 96:3750-3757.. The secondary labeled mAb consisted of 1μL of fluorescein isothiocyanate (FITC) (M31501; Invitrogen, Carlsbad, USA).
(A)Flow cytometric identification of milk polymorphonuclear leukocytes isolated from quarters infected with Corynebacterium bovis, and (B) uninfected quarters. The recordings of scatter (SCC-H) and fluorescent (FL1-H; CH138A- fluorescein isothiocyanate) properties were performed with 20,000 events gated on a standard population excluding most cell debris.
The identification of PMNLs expressing TLR2 and TLR4 was performed using flow cytometry using the following monoclonal antibodies: 1μL of Alexa Fluor 647 mouse anti-human CD282 (51-902182; eBioscience, San Diego, USA) for TLR2 expression and 10μL of phycoerythrin (PE) mouse anti-human CD284 (MCA2061PE; AbD Serotec, Oxford, England) for TLR4 expression. For the identification of neutrophils expressing CD14 using flow cytometry, 1μL of the primary mouse IgG1 anti-bovine CD14 (MM61A; VMRD Pullman Inc. Corp(r), Pullman, USA) and 1μL of a Allophycocyanin-conjugated goat anti-mouse IgG1 antibody (A10541; Invitrogen, Carlsbad, USA) were used. The anti-human CD282 (Ibeagha-Awemu et al. 2008Ibeagha-Awemu E.M., Lee J-W., Ibeagha A.E., Bannerman D.D., Paape M.J. & Zhao X. 2008. Bacterial lipopolysaccharide induces increased expression of toll-loke receptors (TLR) 4 and downstream TLR signaling molecules in bovine mammary epithelial cells. Vet. Res.39:11.) and CD284 mAb (Ibeagha-Awemu et al. 2008Ibeagha-Awemu E.M., Lee J-W., Ibeagha A.E., Bannerman D.D., Paape M.J. & Zhao X. 2008. Bacterial lipopolysaccharide induces increased expression of toll-loke receptors (TLR) 4 and downstream TLR signaling molecules in bovine mammary epithelial cells. Vet. Res.39:11., Catalani et al. 2010Catalani E., Amadori M., Vitali A., Bernabucci U., Nardone A. & Lacetera N. 2010. The Hsp72 response in peri-parturient dairy cows: relationship with metabolic and immunological parameters. Cell Stress and Chaperones 15:781-790.) under use have been previously shown to stain bovine cells. First, dot plots were gated for PMNLs (CH138A), as previously described. The PMNLs were first identified as aforementioned. Unlabeled primary mAb for the identification of CD14 or labeled mAb for TLR2 and TLR4 were then added to the cell suspension and incubated for 30 min at room temperature in the dark. Next, the isolated milk cell suspension was centrifuged at 400xg for 8 min, and in the tube in which CD14 expression would be assessed, the secondary labeled mAb for CD14 was added. Lastly, the isolated milk cell suspension was centrifuged at 400xg for 8 min, and 300μL of PBS was added to the cell suspension.
The expression of CD14, TLR2 and TLR4 on the cell surface of neutrophils was analyzed in gated CH138A-positive cells. The percentage of PMNLs expressing CD14, TLR2 or TLR4 was calculated as the number of fluorescent PMNLs divided by the total PMNL count and multiplied by 100. The median fluorescence intensities of CD14, TLR2 and TLR4 expression were estimated by the geometric median of the fluorescence divided by the number of PMNLs that expressed CD14, TLR2 or TLR4. In this assay, 20,000 milk cells were examined in each sample. FlowJo TreeStar Software (TreeStar Inc., Ashland, OR, USA) was used to analyze the data.
Statistical analyses were performed using GraphPad Prism 5.0 software(r) (GraphPad Software, Inc., San Diego, CA, USA). First, the homoscedasticity and the distribution of all variables were examined using normal probability plots and the Shapiro-Wilk test. All SCCs were log10 transformed, as the distribution was not normal. To compare the differences between the groups, Student's t-test for unpaired data was used for the data with normal distributions (SCC, percentage of neutrophils that CD14, and expression of TLR4 by milk neutrophils), and the Mann-Whitney rank test was used for the nonparametric data (percentage of milk neutrophils, percentage neutrophils that expressed CD14, TLR2 and TLR4, and expression of CD14 and TLR2 by milk neutrophils). The results are reported as the mean ±SD. The association between TLR2 expression by milk neutrophils, CD14 expression by milk neutrophils and the percentage of neutrophils (CH138+) was assessed by linear regression analysis. The significance was set at P≤0.05 (Sampaio 2010Sampaio I.B.M.2010. Estatística Aplicada à Experimentação Animal. Fundação de Ensino e Pesquisa em Medicina Veterinária e Zootecnia, Belo Horizonte. 264p.).
Results
The mammary glands infected with Corynebactyerium bovis showed a higher logarithmic SCC (5.38±0.66, P≤0.0001) and percentage of neutrophils (30.66±14.82%, P =0.018) compared with the healthy quarters (4.02±0.65; 18.88 ±14.64%). Here, no significant difference in the median fluorescence intensity of TLR4 in milk neutrophils (75.64±27.61, P=0.45) and the percentage of milk neutrophils that expressed TLR4 (20.06±28.96%; P=0.18) was found between C. bovis-infected quarters and healthy quarters (85.01±40.45; 20.78±19.18%). Furthermore, the percentage of neutrophils that expressed TLR2 was not different between C. bovis-infected quarters 15.19±19.73%, P=0.34) and uninfected quarters (17.25±13.26%).
Conversely, a lower median fluorescence intensity of TLR2 in milk neutrophils was observed in C. bovis-infected quarters (46.96±47.71; P=0.03) compared with uninfected quarters (70.02±53.23) (Fig.2). The percentage of neutrophils that expressed CD14 was lower in C. bovis infected quarters (57.32+22.56) compared to the healthy ones (72.53+15.32; P=0.008) (Fig.3). The median fluorescence intensity of CD14 by milk neutrophils was also lower in C. bovis infected quarters (183.8+156.3) compared to uninfected quarters (385.9+348.7; P=0.01). A negative association between TLR2 expression by milk neutrophils and the percentage of neutrophils (CH138A+) in C. bovis infection (r= -0.64; P=0.01) was observed, which was not found in healthy quarters (r= -0.03; P=0.86) by linear regression analysis. Furthermore, a positive association between TLR2 and CD14 expression by milk neutrophils was observed in uninfected quarters (r=0.80; P<0.0001) and a tendency toward this association was observed in C. bovis infected quarters (r=0.50; P=0.07), although CD14 expression was not associated with percentage of milk neutrophils in uninfected (P=0.44) and C. bovis-infected quarters (P=0.76). Otherwise, the percentage of milk neutrophils that expressed CD14 was negatively associated with the percentage of neutrophils in uninfected quarters (r= -0.41; P=0.05), although no association was found in C. bovis-infected quarters (P=0.11).
(A)Flow cytometric analysis of toll-like receptor 2 (TLR2) on cell surface molecule expression in CH138A+ neutrophils in Corynebacterium bovis, and (B) uninfected quarters. For analysis of TLR2 expression, neutrophils were identified by gating on CH138A+ granulocytes.
(A) Percentage of neutrophils (CH138A+) that expressed CD14 (FL3-H) molecule in Corynebacterium bovis infected, and (B) healthy quarters assessed by flow cytometry. The recordings of scatter (SSC-H) and fluorescent (FL4-H) (CD14-Allophycocyanin) properties were performed with 20,000 events gated on milk neutrophils (CH138+).
Discussion
The SCC and the percentage of neutrophils were higher in Corynebacterium bovis infected quarters, as previously described by others (Djabri et al. 2002Djabri B., Bareille N., Beaudeau F. & Seegers H. 2002. Quarter milk somatic cell count in infected dairy cows: a meta-analysis. Vet. Res. 33:334-357.). Although, in the present study we did not know when the intramammary infections occurred, as we worked with naturally C. bovis-infected quarters, as well as we did not assess the nucleotide polymorphism within the PRRs in dairy cows, the present study found significant alteration in TLR2 and CD14 expression by milk neutrophils. It is well known that many TLR2 ligands were found on C. bovis, a Gram-positive bacterium. TLR2 recognizes a broad range of different bacterial structures known as TLR2 ligands (i.e. peptidoglycan). In contrast, TLR4 ligands (i.e. lypopolysaccharides) are not present in C. bovis which can explain the no significant alteration in the expression of TLR4 by milk neutrophils (Schukken et al. 2011Schukken Y.H., Güntler J., Fitzpatrick J., Fontaine M.C., Goetza L., Holst O., Leigh J., Petzl W., Schuberth H.-J., Sipka A., Smith D.G.E., Quesnell R., Watts J., Yancey R., Zerbe H., Gurjar A., Zadoks R.N., Seyfert H.-M. & Members of Pfizer mastitis research consortium. 2011. Host-response patterns of intramammary infections in dairy cows. Vet. Immunol. Immunopathol. 144:270-289., Souza et al. 2012Souza F.N., Ramos Sanchez E.M., Heinemann M.B., Gidlund M.A., Reis L.C., Blagitz M.G., Della Libera A.M.M.P.& Cerqueira M.M.O.P. 2012. The innate immunity in bovine mastitis: the role of pattern-recognition receptors.. Am J. Immunol.8:166-178.). Although, CD14 presumably acts with TLR4 in the recognition of lipopolysaccharide (Souza et al. 2012Souza F.N., Ramos Sanchez E.M., Heinemann M.B., Gidlund M.A., Reis L.C., Blagitz M.G., Della Libera A.M.M.P.& Cerqueira M.M.O.P. 2012. The innate immunity in bovine mastitis: the role of pattern-recognition receptors.. Am J. Immunol.8:166-178.), in the last few years some researchers have pointed out to an interaction between CD14 molecule and TLR2 (Raby et al. 2013Raby A.-C., Holst B., Le Bouder E., Diaz C., Ferran E., Conraux L., Guillemot J.-C., Coles B., Kift-Morgan A., Colmont C.S., Szakmamy T., Ferrara P., Hall J.E., Topley N. & Labéta M.O.2013. Targeting the TLR co-receptor CD14 with TLR2-derived peptides modulates immune response to pathogens. Sci. Transl. Med. 5:185ra64.). This idea was strengthen by the positive association between TLR2 and CD14 expression by milk neutrophils found here.
This fact presumably leads to alteration of the recognition of a broad range of bacterial ligands by neutrophils. Besides this TLRs cannot only respond appropriately but also self-regulate host responses to invading pathogens. For instance, a TLR itself may be degraded, thereby making the receptor unavailable for subsequent ligand activation, or a TLR's expression may be inhibited by anti-inflammatory cytokines (Miggin & O´Neil 2006Miggin S.M. & O´Neil L.A.J. 2006. New insights into the regulation of TLR signaling. J. Leuk. Biol. 80:220-226.). Furthermore, it was demonstrated that TLR2 release is modulated by cell activation, and soluble TLR2 (sTLR2) and the TLR co-receptor soluble CD14 (sCD14) may interact in the natural milieu. This fact resulted in the rapid down-modulation of cell-surface TLR2, thus avoiding an excessive proinflammatory response (LeBouder et al. 2003LeBouder E., Rey-Nores J.E., Rushmere N.K., Grigorov M., Lawn S.D., Affolter M., Griffin G.E., Ferrara P., Schiffrin E.J., Morgan B.P. & Labéta M.O. 2003. Soluble forms of toll-like receptors (TLR)2 capable of modulating TLR2 signaling are present in human plasma and breast milk. J. Immunol.171:6680-6689.). The lack of an association of the percentage of milk neutrophils and the percentage of neutrophils that expressed CD14 in C. bovis-infected quarters reinforced the idea of a release of CD14 co-receptor during infection, as this association was encountered in uninfected quarters.
In addition, TLR activation has been shown to be linked to conditions of oxidative stress, especially considering the mechanisms by which inflammatory cascades are activated (Kim et al. 2006Kim O.S., Lee C.S., Joe E.-H. & Joi I. 2006. Oxidized low density lipoprotein supresses lipopolysaccharide-induced inflammatory response in microglia: Oxidative stress acts through control of inflammation. Biochem. Biophys. Res. Commun. 342:9-18., Gill, Tsung & Billiar 2010Gill R., Tsung A. & Billiar T.R. 2010. Linking oxidative stress to inflammation: toll-like receptors. Free Radic. Biol. Med. 48:1121-1132.). In particular, C. bovis IMIs lead to the discrete augmentation of milk SCCs (Djabri et al. 2002Djabri B., Bareille N., Beaudeau F. & Seegers H. 2002. Quarter milk somatic cell count in infected dairy cows: a meta-analysis. Vet. Res. 33:334-357.), and SCCs appear to be negatively associated with the release of hydrogen peroxide by leukocytes (Bastos et al. 2012Bastos C.R., Blagitz M.G., Souza F.N., Batista C.F., Strincagnolo C.R., Azedo M.R. & Della Libera A.M.M.P. 2012. Viabilidade celular, fagocitose e espraiamento de fagócitos mononucleares, e liberação de peróxido de hidrogênio por leucócitos de glândulas mamárias sadias e infectadas. Pesq. Vet. Bras. 32:850-854.). It can also be speculated that C. bovis modulates TLR2 expression in an attempt to control inflammation and tissue injury, especially given that hydrogen peroxide plays a crucial role in the recruitment of leukocytes to the site of infection (Niethammer et al. 2009Niethammer P., Grabher C., Look T. & Mitchison T.J. 2009. A tissue-scale gradient of hydrogen peroxide mediates rapid wound detection in zebrafish. Nature 459:996-1000.). This idea is reinforced by the negative association between TLR2 expression by milk neutrophils and the percentage of neutrophils (CH138A+) in C. bovis infection, which was not found in healthy quarters.
We hypothesize that the lower expression of CD14 by milk neutrophils may be a result of cell activation, which releases CD14 to the cell surface. If it is true, this fact also strength the idea that IMIs by C. bovis may offer protection against IMIs by other bacteria as sCD14 inhibit the growth of bacteria especially in the early stages of infection (Ohnishi, Muroi & Tanamoto 2010Ohnishi T., Muroi M. & Tanamoto K.-I. 2010. Inhibitory effects of soluble MD-2 and soluble CD14 on bacterial growth. Microbiol. Immunol. 54:74-80.). Nevertheless, here, we did not measure sTLR2 and sCD14.
Conclusion
Overall, the present study indicated that Corynebacterium bovis IMIs have implications on mammary gland immunity, showed here by the alteration in the expression of CD14 and TLR2, which may interfere in the probability of new infections.
Acknowledgements
The authors are grateful to FAPESP for financial support. Blagitz and Souza contributed equally to this manuscript and are the first authors.
- Bansal B.K., Hamann J., Grabowskit N.T. & Singh B. 2005. Variation in the composition of selected milk fraction samples from healthy and mastitic quarters, and its significance for mastitis diagnosis. J. Dairy Res. 72:144-152.
- Bastos C.R., Blagitz M.G., Souza F.N., Batista C.F., Strincagnolo C.R., Azedo M.R. & Della Libera A.M.M.P. 2012. Viabilidade celular, fagocitose e espraiamento de fagócitos mononucleares, e liberação de peróxido de hidrogênio por leucócitos de glândulas mamárias sadias e infectadas. Pesq. Vet. Bras. 32:850-854.
- Benites N.R., Melville P.A. & Costa E.O. 2003. Evaluation of the microbiological status of milk and various structures in mammary glands of from naturally infected dairy cows. Trop. Anim. Health Prod. 35:301-307.
- Blagitz M.G., Souza F.N., Batista C.F., Santos B.P., Parra A.C., Azevedo L.F.F., Melville P.A., Benites N.R.& Della Libera A.M.M.P.2013. Function of milk polymorphonuclear neutrophil leukocytes in bovine mammary glands infected with Corynebacterium bovis. J. Dairy Sci. 96:3750-3757.
- Brooks B.W. & Barnum D.A. 1984. The susceptibility of bovine udder quarters colonized with Corynebacterium bovis to experimental infection with Staphylococcus aureus or Streptococcus agalactiae. Can. J. Comp. Med. 48:146-150.
- Catalani E., Amadori M., Vitali A., Bernabucci U., Nardone A. & Lacetera N. 2010. The Hsp72 response in peri-parturient dairy cows: relationship with metabolic and immunological parameters. Cell Stress and Chaperones 15:781-790.
- Djabri B., Bareille N., Beaudeau F. & Seegers H. 2002. Quarter milk somatic cell count in infected dairy cows: a meta-analysis. Vet. Res. 33:334-357.
- Gill R., Tsung A. & Billiar T.R. 2010. Linking oxidative stress to inflammation: toll-like receptors. Free Radic. Biol. Med. 48:1121-1132.
- Haltia L., Honkanen-buzalski T., Spiridonova I., Olkonen A. & Myllys V. 2006. A study of bovine mastitis, milking procedures and management practices on 25 Estonian dairy herds. Acta Vet. Scand. 48:22.
- Huxley J.N., Helps C.R. & Bradley A.J. 2004. Identification of Corynebacterium bovis by endonuclease restriction analysis of the 16S rRNA gene sequence. J. Dairy Sci.87:38-45.
- Ibeagha-Awemu E.M., Lee J-W., Ibeagha A.E., Bannerman D.D., Paape M.J. & Zhao X. 2008. Bacterial lipopolysaccharide induces increased expression of toll-loke receptors (TLR) 4 and downstream TLR signaling molecules in bovine mammary epithelial cells. Vet. Res.39:11.
- Kim O.S., Lee C.S., Joe E.-H. & Joi I. 2006. Oxidized low density lipoprotein supresses lipopolysaccharide-induced inflammatory response in microglia: Oxidative stress acts through control of inflammation. Biochem. Biophys. Res. Commun. 342:9-18.
- Koess C. & Hamann J.2008. Detection of mastitis in the bovine mammary gland by flow cytometry at early stages. J. Dairy Res.75:225-232.
- Lam T.J., Schukken Y.H., Van Vliet J.H., Grommer F.J., Tielen M.J. & Brand A. 1997. Effect of natural infection with minor pathogens on susceptibility to natural infection with major pathogens in the bovine mammary gland. Am. J. Vet. Res.58:17-22.
- LeBouder E., Rey-Nores J.E., Rushmere N.K., Grigorov M., Lawn S.D., Affolter M., Griffin G.E., Ferrara P., Schiffrin E.J., Morgan B.P. & Labéta M.O. 2003. Soluble forms of toll-like receptors (TLR)2 capable of modulating TLR2 signaling are present in human plasma and breast milk. J. Immunol.171:6680-6689.
- Miggin S.M. & O´Neil L.A.J. 2006. New insights into the regulation of TLR signaling. J. Leuk. Biol. 80:220-226.
- Niethammer P., Grabher C., Look T. & Mitchison T.J. 2009. A tissue-scale gradient of hydrogen peroxide mediates rapid wound detection in zebrafish. Nature 459:996-1000.
- Ohnishi T., Muroi M. & Tanamoto K.-I. 2010. Inhibitory effects of soluble MD-2 and soluble CD14 on bacterial growth. Microbiol. Immunol. 54:74-80.
- Oliver S.P., Lewis M.J., Gillespie B.E., Dowlen H.H., Jaenicke E.C. & Roberts R.K. 2004. Microbiological Procedures for the Diagnosis of Bovine Udder Infection and Determination of Milk Quality. 4th ed. National Mastitis Council, Verona. 47p.
- Paape M.J., Liluis E.-M., Wiitanen P.A., Kontio M.P. & Miller R.H. 1996. Intramammary defense against infections induced by Escherichia coli in cows. Am J Vet Res. 57:477-482.
- Piepers S., De Vliegher S., Demeyere K., Lamrecht B.N., Kruif A., Meyer E. & Opsomer G. 2009. Technical note: Flow cytometric identification of bovine milk neutrophils and simultaneous quantification of their viability. J. Dairy Sci.92:626-631.
- Raby A.-C., Holst B., Le Bouder E., Diaz C., Ferran E., Conraux L., Guillemot J.-C., Coles B., Kift-Morgan A., Colmont C.S., Szakmamy T., Ferrara P., Hall J.E., Topley N. & Labéta M.O.2013. Targeting the TLR co-receptor CD14 with TLR2-derived peptides modulates immune response to pathogens. Sci. Transl. Med. 5:185ra64.
- Reyher K.K., Dohoo I.R., Scholl D.T. & Keefe G.P. 2012. Evaluation of minor pathogens intramammary infection, susceptibility parameters, and somatic cell counts on the development of new intramammary infections with major mastitis pathogens. J. Dairy Sci.95:3766-3780.
- Sampaio I.B.M.2010. Estatística Aplicada à Experimentação Animal. Fundação de Ensino e Pesquisa em Medicina Veterinária e Zootecnia, Belo Horizonte. 264p.
- Schukken Y.H., Güntler J., Fitzpatrick J., Fontaine M.C., Goetza L., Holst O., Leigh J., Petzl W., Schuberth H.-J., Sipka A., Smith D.G.E., Quesnell R., Watts J., Yancey R., Zerbe H., Gurjar A., Zadoks R.N., Seyfert H.-M. & Members of Pfizer mastitis research consortium. 2011. Host-response patterns of intramammary infections in dairy cows. Vet. Immunol. Immunopathol. 144:270-289.
- Sordillo L.M., Oliver S.P., Guidry A.J. & Dermody J.T. 1988. Humoral immune response of bovine mammary glands colonized with Corynebacterium bovis: enumeration of plasma cell populations in tissue and immunoglobulin concentrations in milk. J. Vet. Med. B 35:617-627.
- Souza F.N., Ramos Sanchez E.M., Heinemann M.B., Gidlund M.A., Reis L.C., Blagitz M.G., Della Libera A.M.M.P.& Cerqueira M.M.O.P. 2012. The innate immunity in bovine mastitis: the role of pattern-recognition receptors.. Am J. Immunol.8:166-178.
- Souza G.N., Brito J.R.F., Moreira E.C., Brito M.A.V.P. & Silva M.V.G.B. 2009. Variação da contagem de células somáticas em vacas leiteiras de acordo com patógenos da mastite. Arq. Bras. Med. Vet. Zoot. 61:1015-1020.
- Tenhagen B.A., Köster G., Wallmann J. & Heuwieser W. 2006. Prevalence of mastitis pathogens and their resistance against antimicrobial agents in dairy cows in Brandenburg, Germany. J. Dairy Sci.89:2542-2551.
- Zadoks R.N., Allore H.G., Barkema H.W., Sampimon O.C., Wellenberg G.J., Gröhn Y.T. & Schukken Y.H.2001. Cow- and quarter-level risk factors for Streptococcus uberis and Staphylococcus aureus mastitis. J. Dairy Sci.84:2649-2663.
Publication Dates
-
Publication in this collection
Jan 2015
History
-
Received
19 Feb 2014 -
Accepted
12 Mar 2014