Abstract
Chemical modifications were used to identify some of the functionally important amino acid residues of the potato plant uncoupling protein (StUCP). The proton-dependent swelling of potato mitochondria in K+-acetate in the presence of linoleic acid and valinomycin was inhibited by mersalyl (Ki = 5 µM) and other hydrophilic SH reagents such as Thiolyte MB, iodoacetate and 5,5'-dithio-bis-(2-nitrobenzoate), but not by hydrophobic N-ethylmaleimide. This pattern of inhibition by SH reagents was similar to that of brown adipose tissue uncoupling protein (UCP1). As with UCP1, the arginine reagent 2,3-butadione, but not N-ethylmaleimide or other hydrophobic SH reagents, prevented the inhibition of StUCP-mediated transport by ATP in isolated potato mitochondria or with reconstituted StUCP. The results indicate that the most reactive amino acid residues in UCP1 and StUCP are similar, with the exception of N-ethylmaleimide-reactive cysteines in the purine nucleotide-binding site.
plant mitochondria; uncoupling protein; chemical modification; mitochondrial swelling; reconstitution
Braz J Med Biol Res, December 2000, Volume 33(12) 1413-1420
Important amino acid residues of potato plant uncoupling protein (StUCP)
P. Jezek1, A.D.T. Costa2 and A.E. Vercesi2
1Department of Membrane Transport Biophysics, Institute of Physiology, Academy of Science, Prague, Czech Republic
2Departamento de Patologia Clínica (NMCE), Faculdade de Ciências Médicas, Universidade Estadual de Campinas, Campinas, SP, Brasil
References
Correspondence and Footnotes Correspondence and Footnotes Correspondence and Footnotes
Abstract
Chemical modifications were used to identify some of the functionally important amino acid residues of the potato plant uncoupling protein (StUCP). The proton-dependent swelling of potato mitochondria in K+-acetate in the presence of linoleic acid and valinomycin was inhibited by mersalyl (Ki = 5 µM) and other hydrophilic SH reagents such as Thiolyte MB, iodoacetate and 5,5'-dithio-bis-(2-nitrobenzoate), but not by hydrophobic N-ethylmaleimide. This pattern of inhibition by SH reagents was similar to that of brown adipose tissue uncoupling protein (UCP1). As with UCP1, the arginine reagent 2,3-butadione, but not N-ethylmaleimide or other hydrophobic SH reagents, prevented the inhibition of StUCP-mediated transport by ATP in isolated potato mitochondria or with reconstituted StUCP. The results indicate that the most reactive amino acid residues in UCP1 and StUCP are similar, with the exception of N-ethylmaleimide-reactive cysteines in the purine nucleotide-binding site.
Key words: plant mitochondria, uncoupling protein, chemical modification, mitochondrial swelling, reconstitution
Introduction
The functionally well-characterized plant uncoupling mitochondrial protein (PUMP) (1-13) has been cloned from potato (StUCP) (14) and Arabidopsis thaliana (AtPUMP) (15) gene libraries. We provided evidence that potato PUMP is a product of the StUCP gene (16). Consequently, PUMP has been recognized as a member of the uncoupling protein (UCP) subfamily, homologous with mammalian UCPs such as UCP1 of brown adipose tissue mitochondria (2,17-19), the ubiquitous UCP2 (20), UCP3 of striated muscle (21), and two brain-specific uncoupling proteins, UCP4 (22) and BMCP1 (23). The physiological roles of UCPs in mammals include nonshivering thermogenesis in neonates (UCP1), regulation of weight balance and inflammatory responses such as fever (UCP2), nonshivering thermogenesis in skeletal muscle (UCP3), and possibly the prevention or regulation of apoptosis in the brain (UCP4, BMCP1). We have hypothesized (2,4-6) that, in plants, StUCP may be responsible for a respiratory burst in climacteric fruits and for all physiological events when a sudden cessation of ATP synthesis is required such as during seed formation and senescence. Several climacteric fruits such as tomato (7), banana, mango, apple and others (2) contain StUCP. A mild thermogenesis mediated by StUCP can accelerate respiration and, hence, metabolic rates (2) and mild StUCP-mediated uncoupling leads to a decreased formation of reactive oxygen species (8). Both functions are beneficial for plant growth and development. Thus, fully activated thermogenesis could facilitate plant growth at low temperature, e.g., in roots, tubers (9), or during seed germination. StUCP may also play specific roles during plant senescence and could contribute to processes that maintain the seed dormancy.
All UCPs presumably enable the passage of fatty acid (FA) anions and thus promote FA cycling, leading to H+ uniport mediated by neutral FA molecules, which results in uncoupling (2,19,24,25). The FA cycling mechanism has been confirmed for UCP1 (26), UCP2 and UCP3 (24) and StUCP (6). UCP1 also translocates a wide variety of monovalent, unipolar anions, including short-, medium-, and long-chain alkylsulfonates (27). Hexane and undecanesulfonate transport has also been demonstrated for StUCP (5,6). Nevertheless, StUCP is not able to translocate small hydrophilic anions such as Cl- and pyruvate (5,6) which are good transport substrates for UCP1 (2,19,27,28).
The protein chemistry of StUCP has not been studied to the same extent as has UCP1. A 32-kDa StUCP has been characterized as a hydrophobic protein which is not retained on hydroxylapatite in the detergent micellar solution (1,6,7). Chemical modifications of reactive amino acid residues, the cleavage pattern produced by proteases, and ligand binding (except for studies with 8-azido-ATP (13)) have not been studied in StUCP. In the present study, we examined the effects of several chemical modifiers on StUCP-mediated transport as well as StUCP inhibition by purine nucleotides. Our results clearly show that the pattern of reactive amino acid residues in StUCP is similar to that of UCP1, with the exception that no N-ethylmaleimide (NEM)-reactive cysteines were found in the purine nucleotide-binding site of StUCP.
Material and Methods
Biological material and chemicals
Potatoes (Solanum tuberosum, L. cv. Bintje) were purchased locally. Nucleotides, bovine serum albumin (BSA), valinomycin, nigericin, iodoacetate, linoleic acid, N-tris [hydroxymethyl]methyl-2-aminoethanesulfonic acid (TES), N-[2-hydroxyethyl]piperazine-N'-[2-ethane] sulfonic acid (HEPES), tetraethylammonium hydroxide (TEA-OH), carbonyl cyanide trifluoromethoxyphenylhydrazone (FCCP), ethylene glycol-bis(ß-aminoethyl ether) N,N,N',N'-tetraacetic acid (EGTA), ethylenediaminetetraacetic acid (EDTA), 5,5'-dithio-bis-(2-nitrobenzoate) (DTNB), 4,4'-diisothiocyanatostilbene-2,2'-disulfonic acid (DIDS), 2,4,6-trinitrobenzenesulfonic acid (TNBS), pyridoxalphosphate, phenylglyoxal, propranolol, phenylarsineoxide, eosinmaleimid, 2,3-butadione and phospholipids were purchased from Sigma Chemical Co. (St. Louis, MO, USA). The fluorescent probe SPQ (6-methoxy-N-(3-sulfopropyl) quinolinium) and thiol reagent Thiolyte® MB were from Calbiochem (La Jolla, CA, USA). The potassium probe PBFI (potassium-binding benzofuraneisophthalate) was from Molecular Probes (Eugene, OR, USA). All other reagents were commercial products of the purest grade available.
Isolation of mitochondria and protein determination
Potato mitochondria were isolated as described previously (5,8,9) in medium containing 250 mM sucrose, 10 mM HEPES, pH 7.2, and 0.3 mM EGTA. The protein concentration was 30-40 mg/ml, as determined by the biuret method. A crude fraction was used for swelling studies and for most of the isolations. For some isolations, a Percoll gradient centrifugation was used to remove contamination by plastid proteins, starch and other substances. Qualitatively, transport measurements using the crude fraction gave identical results as those performed with Percoll-purified mitochondria.
Swelling assay of StUCP transport function
Proton-dependent swelling of potato mitochondria (0.2 mg protein/ml) in K+-acetate (55 mM K+-acetate, 5 mM K+-HEPES, 0.2 mM Tris-EDTA, 0.1 mM Tris-EGTA, pH 6.9) initiated by valinomycin in the presence of linoleic acid (16 µM) has been used as a standard assay for StUCP-mediated transport (5). Since valinomycin allows the uniport uptake of K+ and neutral acetic acid is able to penetrate the lipid bilayer, an efflux of H+ is necessary to induce swelling. In our assay, this H+ efflux was concomitant with linoleic acid cycling which allowed swelling since StUCP mediated the uptake of linoleic acid anion, while protonated linoleic acid passed spontaneously through the lipid bilayer by a flip-flop mechanism and released H+ externally. Hexanesulfonate uniport was assayed as valinomycin-induced swelling in medium containing 51.1 mM Na+-hexanesulfonate, 30.8 mM K+-HEPES, pH 7.2, 190 µM Tris-EDTA and 95 µM Tris-EGTA. The side effects caused by the chemical modifiers used, including the induction of mitochondrial swelling without the addition of ionophore and membrane stiffening, were controlled by performing a swelling assay in K+-acetate containing nigericin, which does not depend on protein carriers. When a decrease in this rate (vNig[c]) was observed at a given concentration [c] of modifier, the rates of valinomycin-induced StUCP-mediated swelling were corrected by multiplying this decrease by the factor vNig[c = 0]/vNig[c].
Chemical modifications of potato mitochondria
For carrying reactions, mitochondria were resuspended in the sucrose isolation medium (5 mg protein/ml) and aliquots of stock solutions (aqueous or in dimethylsulfoxide) of various reagents were added and incubated for 1 h (unless otherwise indicated) at 0oC. For NEM, DTNB and phenylglyoxal, pH was raised to 8.2 by adding 20 mM Tris-HEPES, pH 8.4, to the stock solution and 2 µM propranolol was added.
StUCP isolation and reconstitution
StUCP was isolated from potato mitochondria on hydroxylapatite as described previously (6). The same procedure was used for the isolation of potato mitochondria pretreated with Thiolyte MB or 2,3-butadione. Thirty micrograms of isolated StUCP was incorporated into liposomes by detergent removal using Bio-Beads SM2 (BioRad, Hercules, CA, USA) and the vesicles were depleted of the external probe by passage through Sephadex G25-300 spin columns (6). The FA-induced H+ fluxes initiated by valinomycin were monitored either as the counterflux of K+, using PBFI (6,7), or by TES anion quenching of SPQ, as described previously (6,7), on an F-4010 Hitachi fluorescence spectrophotometer (Hitachi Ltd., Tokyo, Japan). For PBFI monitoring, the vesicles (25 µl per assay) contained 75 mM TEA sulfate, 75 mM TEA-TES, pH 7.2, 0.05 mM K2SO4 and 300 µM PBFI. The external medium contained 75 mM K2SO4 and 75 mM TEA-TES, pH 7.2. For SPQ monitoring, the vesicles (25 µl per assay) contained 84.4 mM TEA sulfate, 28.85 mM TEA-TES, pH 7.2 ([TEA] was 9.2 mM) and 0.6 mM Tris-EGTA. In the external medium, 84.4 mM K2SO4 replaced TEA sulfate.

Effect of hydrophilic SH reagents on StUCP-mediated transport in mitochondria
Proton-dependent swelling of potato mitochondria initiated by valinomycin in K+-acetate containing linoleic acid was reversibly inhibited by the organomercurial SH reagent mersalyl with an apparent Ki of 5 µM (Figure 1A, only 10-s preincubations). This type of swelling reflected the ability of StUCP to translocate linoleic acid anions (5). The effect of mersalyl can be considered as a specific inhibition, since swelling independent of a protein carrier, i.e., the nigericin-induced swelling in K+-acetate, was not affected up to 100 µM mersalyl (Figure 1A). Above 100 µM, and above 40 µM in the presence of linoleic acid, mersalyl induced nonspecific permeability changes which were observed as mitochondrial swelling without the ionophore. Some mitochondrial preparations were more sensitive to mersalyl and this made measurements with them more difficult.
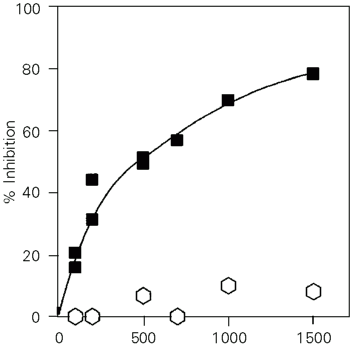
To avoid the interference of nonspecific permeability changes, we used Thiolyte MB, a covalently interacting SH modifier. Mitochondria were preincubated for 1 h with increasing Thiolyte MB doses (Figure 2). The IC50 for Thiolyte MB was around 500 nmol/mg protein. Carrier-independent swelling was not significantly affected by Thiolyte MB, indicating that the modification of the SH groups in StUCP inhibits the transport activity of this protein. Carboxymethylation by iodoacetate (which also affects SH groups) also inhibited StUCP transport activity at higher doses (IC50 of 100 µmol/mg protein), but only with 10-s preincubations (Figure 1B). Ellman's reagent (DTNB) inhibited the activity by 18 and 31% at 1000 and 3000 nmol/mg protein, respectively, after a 2-h incubation, as calculated from the rates corrected for the nonspecific effect (incubations at pH >8 lead to preswelling after a few hours). In contrast, NEM and other hydrophobic SH reagents (eosinmaleimide, phenylarsineoxide) were not inhibitory up to 10 µmol/mg protein. Hexanesulfonate uniport via StUCP was partially inhibited by hydrophilic SH reagents, e.g., by 1000 nmol Thiolyte MB/mg protein.
Effect of arginine reagents on ATP inhibition of StUCP-mediated transport
Reagents specific for other amino acid residues did not inhibit transport or prevent the inhibition by ATP at doses up to 10 µmol/mg protein. The reagents tested included DIDS, TNBS and lysine-specific pyridoxalphosphate. Only an arginine-specific reagent, 2,3-butadione, completely prevented the inhibition of linoleic acid transport by 4 mM ATP (Figure 3) at doses above 100 nmol/mg protein(see inset in Figure 3). Thus, a 1-h incubation with 4000 nmol/mg protein 2,3-butadione shifted the ATP dose-response curve so that the extrapolated apparent Ki was much greater than 10 mM (Figure 3). Surprisingly, phenylglyoxal, a more bulky arginine reagent, had no effect at doses up to 10 µmol/mg protein. NEM, which prevented nucleotide inhibition of UCP, also had no effect on ATP inhibition of StUCP (data not shown).
Confirmation of the effects of 2,3-butadione and Thiolyte MB using reconstituted StUCP
The effect of 2,3-butadione on StUCP reconstituted into proteoliposomes after premodification by 2,3-butadione in mitochondria was identical to that found in potato mitochondria. 2,3-Butadione prevented purine nucleotide inhibition of H+ efflux, including inhibition by GTP, when the H+ efflux was monitored with the fluorescent probe PBFI concomitant with K+ influx (Figure 4). The inhibitory effect of Thiolyte MB was also confirmed for isolated StUCP reconstituted into liposomes for which linoleic acid uniport or concomitant H+ efflux was detected by TES quenching of the fluorescent probe SPQ (Figure 5). Reconstituted Thiolyte MB-modified StUCP showed no transport activity (Figure 5).
Results

The pattern of reactive amino acid residues in StUCP was surprisingly similar to that of mammalian UCP1 (29-33; for reviews, see 17-19). This similarity suggests that the structures of StUCP and UCP1 are very likely to be closely related, despite only about 40% identity in their sequences (14,15).
Discussion
The chemical modification of reactive amino acid residues in proteins has been widely used to study protein structure/function relationships. Site-directed mutagenesis has shown that the identification of a residue as essential for a given function is not a straightforward task. In many cases, the effects of modifiers differ from the phenotypes of the corresponding substitution mutants. Interference by the reagent and/or the mutation with the protein function may indicate that i) the residue is essential for that function, i.e., is involved in the required functional interactions (in this case, the substitution mutants have an identical phenotype), ii) the modification of the residue produces steric hindrances which are the actual cause of the altered function (substitution mutations show no such effect), or iii) the residue is important for maintaining a proper conformation of the protein and cannot retain this position after being modified or mutated. With UCP1, case (i) is valid for its Arg 276, whereas case (ii) has been indicated for its cysteine residues.
When Arg 276 was either substituted in a mutated UCP1 protein (34) or modified by phenylglyoxal and 2,3-butadione (32), purine nucleotide binding and gating were absent. Since the proximal third matrix segment was photolabeled at three different positions with 8-azido-, 2-azido- and 3'-O-(5-fluoro-2,4-dinitrophenyl)adenosine 5'-triphosphate (FNDP-ATP) (35), and since the deletion of residues 261-269 resulted in the lack of nucleotide inhibition (36), it was concluded that the main location of the nucleotide-binding site in UCP1 was located between the fifth and sixth transmembrane segments. This site probably forms a water-filled cavity which penetrates deeply into the membrane close to the opposite surface (35). This cavity in UCP1 is lined with SH residues (C213, C224, C253, C287, C304, and possibly C188). Studies on these residues identified the case (ii) described above, since SH substitution mutants of UCP1 have no disrupted binding or transport (33).
The modification of UCP1 by hydrophobic and hydrophilic SH reagents drastically reduces inhibition by GDP (31). In contrast to UCP1, NEM did not prevent ATP inhibition of transport in StUCP. However, transport was inhibited by the arginine reagent 2,3-butadione. These findings suggest a probable difference between the purine nucleotide-binding site of UCP1 and StUCP and indicate that StUCP does not contain modifiable SH groups at or close to the nucleotide-binding site. Alternatively, SH groups may not be important for maintaining the integrity of StUCP conformation. These findings agree with the amino acid sequence of potato plant UCP (14,16). Thus, C188 of UCP1 is conserved in UCP2 and UCP3, but is substituted by A197 in StUCP (14). Of the two cysteines conserved in the fifth a-helix of UCP1, 2 and 3, the first, C234, is shifted two residues towards the matrix in StUCP such that its position in the a-helix is occupied by F231. The second SH (C213 of UCP1) is not conserved in StUCP and is substituted by T220. The similarity of the purine nucleotide-binding site in StUCP and UCP1 is reflected by the effect of 2,3-butadione, which probably interacts with the conserved arginines in UCPs (and in the mitochondrial carrier gene family as a whole), such as R276 of UCP1 (37), which corresponds to R281 and R278 in StUCP and AtUCP, respectively (14,15).
Hydrophilic, but not hydrophobic, SH reagents were good inhibitors of UCP1-mediated FA-induced H+ transport (30). Similarly, in StUCP only hydrophilic SH reagents inhibited StUCP-mediated transport of linoleic acid and hexanesulfonate, while hydrophobic SH reagents, arginine, lysine and other modifiers had no effect. Hence, inhibition by hydrophilic SH reagents is common to StUCP and UCP1. This inhibitory effect on UCP1 has not yet been fully explained. The SH groups which maintain the integrity of the translocation pathway or, alternatively, participate directly in the translocation mechanism, are probably distinct from those interacting with NEM (in UCP1) and interfere with nucleotide binding after modification (31). These SH groups are probably located at yet unknown similar positions in the StUCP sequence. In addition, the type of interference by SH reagents with the StUCP translocation mechanism is likely to be the same as for UCP1. A possible candidate for such a residue is C90, located in the second a-helix of StUCP, which does not have any counterpart in the sequences of UCP1, 2 and 3. Residue C24 of UCP1, absent in StUCP, may serve a similar function for C90 in StUCP.
Address for correspondence: A.E. Vercesi, Departamento de Patologia Clínica (NMCE), FCM, UNICAMP, Caixa Postal 6111, 13083-970 Campinas, SP, Brasil. Fax: +55-19-788-1118. E-mail: anibal@obelix.unicamp.br
Research supported by PRONEX, FAPESP, CNPq and PADCT/CNPq. A.D.T. Costa was the recipient of a FAPESP fellowship. Some chemicals were purchased through a grant (No. 301/98/0568) from the Grant Agency of the Czech Republic. Received April 28, 2000. Accepted September 13, 2000.
- 1. Vercesi AE, Martins IS, Silva MAP, Leite HMF, Cuccovia IM & Chaimovich H (1995). PUMPing plants. Nature, 375: 24.
- 2. Jezek P, Engstová H, Zácková M, Vercesi AE, Costa ADT, Arruda P & Garlid KD (1998). Fatty acid cycling mechanism and mitochondrial uncoupling proteins. Biochimica et Biophysica Acta, 1365: 319-327.
- 3. Vercesi AE, Chaimovich H & Cuccovia IM (1997). A plant uncoupling mitochondrial protein, PUMP. Recent Research Development in Plant Physiology, 1: 85-91.
- 4. Vercesi AE, Jezek P, Costa ADT, Kowaltowski AJ, Maia IG & Arruda P (1998). A plant uncoupling mitochondrial protein. In: Moller IM, Gardestrom P, Glimelius K & Glaser E (Editors), Plant Mitochondria: From Gene to Function Backhuys Publishers, Leiden, The Netherlands, 435-440.
- 5. Jezek P, Costa ADT & Vercesi AE (1996). Evidence for anion-translocating plant uncoupling mitochondrial protein in potato mitochondria. Journal of Biological Chemistry, 271: 32743-32748.
- 6. Jezek P, Costa ADT & Vercesi AE (1997). Reconstituted plant uncoupling mitochondrial protein allows for proton translocation via fatty acid cycling mechanism. Journal of Biological Chemistry, 272: 24272-24278.
- 7. Costa ADT, Nantes IL, Jezek P, Leite A, Arruda P & Vercesi AE (1999). Plant uncoupling mitochondrial protein activity in mitochondria isolated from tomatoes at different stages of ripening. Journal of Bioenergetics and Biomembranes, 31: 527-533.
- 8. Kowaltowski AJ, Costa ADT & Vercesi AE (1998). Activation of the potato plant uncoupling mitochondrial protein inhibits reactive oxygen species generation by the respiratory chain. FEBS Letters, 425: 213-216.
- 9. Nantes IL, Fagian MM, Catisti R, Arruda P, Maia IG & Vercesi AE (1999). Low temperature- and aging-promoted expression of PUMP in potato tuber mitochondria. FEBS Letters, 457: 103-106.
- 10. Sluse FE, Almeida AM, Jarmuszkiewicz W & Vercesi AE (1998). Free fatty acids regulate the uncoupling protein and alternative oxidase activities in plant mitochondria. FEBS Letters, 433: 237-240.
- 11. Jarmuszkiewicz W, Almeida AM, Sluse-Goffart C, Sluse FE & Vercesi AE (1998). Linoleic acid-induced activity of plant uncoupling mitochondrial protein in purified tomato fruit mitochondria during resting, phosphorylating, and progressively uncoupled respiration. Journal of Biological Chemistry, 273: 34882-34886.
- 12. Almeida AM, Jarmuszkiewicz W, Khomsi H, Arruda P, Vercesi AE & Sluse FE (1999). Cyanide-resistant, ATP-synthesis-sustained, and uncoupling-protein-sustained respiration during postharvest ripening of tomato fruit. Plant Physiology, 119: 1323-1330.
- 13. Saviani EE, da Silva Jr A & Martins IS (1997). Photoaffinity labelling of the uncoupling protein from potato tuber mitochondria. Plant Physiology and Biochemistry, 35: 701-706.
- 14. Laloi M, Klein M, Reismeier JW, Müller-Röber B, Fleury C, Bouillaud F & Ricquier D (1997). A plant cold-induced uncoupling protein. Nature, 389: 135-136.
- 15. Maia IG, Benedetti CE, Leite A, Turcinelli SR, Vercesi AE & Arruda P (1998). AtPUMP: an Arabidopsis gene encoding a plant uncoupling mitochondrial protein. FEBS Letters, 429: 403-406.
- 16. Ruzicka M, Novák P, Zácková M, Costa ADT, Vercesi AE, & Jezek P (1999). Plant uncoupling mitochondrial protein is the product of StUCP gene. Proceedings of the XXVIII Annual Meeting of the Brazilian Society for Biochemistry and Molecular Biology, Caxambu, MG, Brazil, May 22-25, A13, 3.
- 17. Klingenberg M (1990). Mechanism and evolution of the uncoupling protein of brown adipose tissue. Trends in Biochemical Sciences, 15: 108-112.
- 18. Nedergard J & Cannon B (1992). The uncoupling protein thermogenin and mitochondrial thermogenesis. In: Ernster L (Editor), Molecular Mechanisms in Bioenergetics Vol. 23. Elsevier Science, London, 385-420.
- 19. Jezek P & Garlid KD (1997). Mammalian mitochondrial uncoupling proteins. International Journal of Biochemistry and Cell Biology, 30: 1163-1168.
- 20. Fleury C, Neverova M, Collins S, Raimbault S, Champigny O, Levi-Meyrueis C, Bouillaud F, Seldin MF, Surwit RS, Ricquier D & Warden CH (1997). Uncoupling protein-2: a novel gene linked to obesity and hyperinsulinemia. Nature Genetics, 15: 269-272.
- 21. Boss O, Samec S, Paoloni-Giacobino A, Rossier C, Dullo A, Seydoux J, Muzzin P & Giacobino J-P (1997). Uncoupling protein-3: a new member of the mitochondrial carrier family with tissue specific expression. FEBS Letters, 408: 39-42.
- 22. Mao W, Yu XX, Zhong A, Li W, Brush J, Sherwood SW, Adams SH & Pan G (1999). UCP4, a novel brain-specific mitochondrial protein that reduces membrane potential in mammalian cells. FEBS Letters, 443: 326-330.
- 23. Sanchis D, Fleury C, Chomiki N, Goubern M, Huang Q, Neverova M, Gregoire F, Easlick J, Raimbault S, Levi-Meyrueis C, Miroux B, Collins S, Seldin M, Richard D, Warden C, Bouillaud F & Ricquier D (1998). BMCP1, a novel mitochondrial carrier with high expression in the central nervous system of humans and rodents, and respiration uncoupling activity in recombinant yeast. Journal of Biological Chemistry, 273: 34611-34615.
- 24. Jaburek M, Varecha M, Gimeno RE, Dembski M, Jezek P, Zhang M, Burn P, Tartaglia LA & Garlid KD (1999). Transport function and regulation of mitochondrial uncoupling proteins 2 and 3. Journal of Biological Chemistry, 274: 26003-26007.
- 25. Skulachev VP (1991). Fatty acid circuit as a physiological mechanism of uncoupling of oxidative phosphorylation. FEBS Letters, 294: 158-162.
- 26. Garlid KD, Orosz DE, Modriansky M, Vassanelli S & Jezek P (1996). On the mechanism of fatty acid-induced proton transport by mitochondrial uncoupling protein. Journal of Biological Chemistry, 271: 2615-2620.
- 27. Jezek P & Garlid KD (1990). New substrates and competitive inhibitors of the Cl- translocating pathway of the uncoupling protein of brown adipose tissue mitochondria. Journal of Biological Chemistry, 265: 19303-19311.
- 28. Jezek P & Borecký J (1998). Mitochondrial uncoupling protein may participate in futile cycling of pyruvate and other monocarboxylates. American Journal of Physiology, 275: C496-C504.
- 29. Kopecký J, Jezek P, Drahota Z & Houstek J (1987). Control of uncoupling protein in brown fat mitochondria by purine nucleotides. Chemical modification by diazobenzenesulfonate. European Journal of Biochemistry, 164: 687-694.
- 30. Jezek P (1987). Sulfhydryl groups are involved in H+ translocation via the uncoupling protein of brown adipose tissue mitochondria. FEBS Letters, 240: 89-93.
- 31. Jezek P & Drahota Z (1989). Sulfhydryl groups of the uncoupling protein of brown adipose tissue mitochondria. Distinction between sulfhydryl groups of the H+ channel and the nucleotide binding site. European Journal of Biochemistry, 183: 89-95.
- 32. Katiyar SS & Shrago E (1989). Reconstitution of purified brown adipose tissue mitochondria uncoupling protein: demonstration of separate identity of nucleotide binding and proton translocation sites by chemical probes. Proceedings of the National Academy of Sciences, USA, 86: 2559-2562.
- 33. Arechaga I, Raimbault S, Prieto S, Levi-Meyrueis C, Zaragoza P, Miroux B, Ricquier D, Bouillaud F & Rial E (1993). Cysteine residues are not essential for uncoupling protein function. Biochemical Journal, 296: 693-700.
- 34. Murdza-Inglis DL, Modriansky M, Patel HV, Woldegiorgis G, Freeman K & Garlid KD (1994). A single mutation in uncoupling protein of rat brown adipose tissue mitochondria abolishes GDP sensitivity of H+ transport. Journal of Biological Chemistry, 269: 7435-7438.
- 35. Mayinger P & Klingenberg M (1992). Labeling of two different regions of the nucleotide binding site of the uncoupling protein from brown adipose tissue mitochondria with two ATP analogs. Biochemistry, 31: 10536-10543.
- 36. Bouillaud F, Arechaga I, Petit PX, Raimbault S, Levi-Meyrueis C, Casteila L, Laurent M, Rial E & Ricquier D (1994). A sequence related to a DNA recognition element is essential for the inhibition by nucleotides of proton transport through mitochondrial uncoupling protein. EMBO Journal, 13: 1990-1997.
- 37. Nelson DR, Lawson JE, Klingenberg M & Douglas MG (1993). Site-directed mutagenesis of the yeast mitochondrial ADP/ATP translocator. Journal of Molecular Biology, 230: 1159-1170.
Correspondence and Footnotes
Publication Dates
-
Publication in this collection
29 Nov 2000 -
Date of issue
Dec 2000
History
-
Accepted
13 Sept 2000 -
Received
28 Apr 2000