Abstract
The opportunistic bacterium Proteus mirabilis secretes a metalloprotease, ZapA, considered to be one of its virulence factors due to its IgA-degrading activity. However, the substrate specificity of this enzyme has not yet been fully characterized. In the present study we used fluorescent peptides derived from bioactive peptides and the oxidized ß-chain of insulin to determine the enzyme specificity. The bradykinin- and dynorphin-derived peptides were cleaved at the single bonds Phe-Ser and Phe-Leu, with catalytic efficiencies of 291 and 13 mM/s, respectively. Besides confirming already published cleavage sites, a novel cleavage site was determined for the ß-chain of insulin (Val-Asn). Both the natural and the recombinant enzyme displayed the same broad specificity, demonstrated by the presence of hydrophobic, hydrophilic, charged and uncharged amino acid residues at the scissile bonds. Native IgA, however, was resistant to hydrolysis by ZapA.
Proteus mirabilis; metalloprotease; substrate specificity; fluorogenic peptides; IgA; insulin ß-chain
Braz J Med Biol Res, November 2001, Volume 34(11) 1397-1403
ZapA, a possible virulence factor from Proteus mirabilis exhibits broad protease substrate specificity
M.A.F. Anéas1, F.C.V. Portaro2, I. Lebrun2, L. Juliano3, M.S. Palma4 and B.L. Fernandes1
1Departamento de Microbiologia, Instituto de Ciências Biomédicas, Universidade de São Paulo, São Paulo, SP, Brasil
2Departamento de Bioquímica e Biofísica, Centro de Toxinologia Aplicada, CEPID-FAPESP, Instituto Butantan, São Paulo, SP, Brasil
3Departamento de Biofísica e Biologia Molecular, Centro de Toxinologia Aplicada, CEPID-FAPESP, Universidade Federal de São Paulo, São Paulo, SP, Brasil
4Departamento de Biologia, Instituto de Ciências Biológicas, Centro de Toxinologia Aplicada, CEPID-FAPESP, Universidade Estadual Paulista, Rio Claro, SP, Brasil
References
Correspondence and Footnotes Correspondence and Footnotes Correspondence and Footnotes
Abstract
The opportunistic bacterium Proteus mirabilis secretes a metalloprotease, ZapA, considered to be one of its virulence factors due to its IgA-degrading activity. However, the substrate specificity of this enzyme has not yet been fully characterized. In the present study we used fluorescent peptides derived from bioactive peptides and the oxidized ß-chain of insulin to determine the enzyme specificity. The bradykinin- and dynorphin-derived peptides were cleaved at the single bonds Phe-Ser and Phe-Leu, with catalytic efficiencies of 291 and 13 mM/s, respectively. Besides confirming already published cleavage sites, a novel cleavage site was determined for the ß-chain of insulin (Val-Asn). Both the natural and the recombinant enzyme displayed the same broad specificity, demonstrated by the presence of hydrophobic, hydrophilic, charged and uncharged amino acid residues at the scissile bonds. Native IgA, however, was resistant to hydrolysis by ZapA.
Key words:Proteus mirabilis, metalloprotease, substrate specificity, fluorogenic peptides, IgA, insulin ß-chain
Introduction
Bacterial metalloproteases, which are usually secreted (1), have been implicated in several physiological as well as pathogenic processes (1,2), acting on various steps related to nutrition acquisition, or hydrolyzing defense factors of their mammal or insect hosts. The serralysins, for instance, secreted by Serratia marcescens, have been described as virulence factors, having an important role in keratitis (3) and pulmonary infections (4), but they also seem to have a role in nutrient digestion/uptake (5). The secreted hemagglutinin protease of Vibrio cholerae cleaves several physiologically important substrates such as mucin, fibronectin and lactoferrin (6), while the immune inhibitor A, a metalloprotease secreted by Bacillus thuringiensis, degrades antibacterial proteins produced by the insect host (7).
It is known that metalloproteases generally possess low substrate selectivity (8), acting less as regulatory enzymes than as enzymes participating in processes of inactivation and catabolism. Therefore, it is expected that these metalloproteases should act upon a variety of substrates, and not display narrow substrate specificity as do the serine proteases, for instance, which often are processing enzymes. In fact, bacterial enzymes, which specifically act on IgA, are serine proteases (9), cleaving the IgA molecule at a single peptide bond in the hinge region. Classical IgA proteases are secreted by Neisseria meningitidis, N. gonorrhea, Haemophilus influenzae, and Streptococcus pneumoniae (9). One exception is the metalloprotease secreted by Streptococcus sanguis (10). Only a few examples of metalloproteases have been described, which display a narrow specificity in the processing of other proteins. These include Clostridium botulinum B and C. tetani neurotoxins (11).
ZapA, the metalloprotease secreted by Proteus mirabilis, has been considered to be an IgA protease (12,13). In order to investigate whether ZapA should be included among specific bacterial metalloproteases, or whether it displays broad substrate specificy, we employed the newly developed fluorogenic operational substrate (14) to characterize putative biological substrates of ZapA.
Material and Methods
Bacterial strains and growth conditions
The P. mirabilis strain N17-12 is prototrophic and stably secretes a metalloprotease (15). Escherichia coli strain DH5a [F-, F80dlac ZDM15 D(lacZYA-argF) U169 endA1 recA1 hsdR17 (rK- mK+) deoR thi-1 supE44 l- gyrA 96relA1)] carries plasmid pCW101, which contains the whole zap gene cluster (12). Luria Bertani medium (1% tryptone, 0.5% yeast extract, 1% NaCl) was used for growth of P. mirabilis and E. coli, supplemented with ampicillin (50 µg/ml) when required.
Protein and peptide substrates
Human secretory IgA from colostrum, oxidized ß-chain of insulin, and trypsin type III from bovine pancreas were from Sigma (St. Louis, MO, USA). The intramolecularly quenched fluorescent peptide substrates were synthesized, purified, and analyzed by previously described procedures (16) using the multiple automated peptide synthesizer PSSM-8 from Shimadzu Corporation (Tokyo, Japan). These peptides contain an Abz-group (ortho-aminobenzoyl) at the N-terminus, and EDDnp (2,4, dinitrophenyl-ethylenediamine) at the C-terminus, as a fluorescent and quencher pair, respectively. The peptides used in this study were the bradykinin-derived peptide (Abz-R-P-P-G-F-S-P-F-R-EDDnp) and the dynorphin-derived peptide (Abz-G-G-F-L-R-R-Q-EDDnp) (14).
Purification of the protease by phenyl-Sepharose affinity chromatography
The native and recombinant ZapA proteases were purified as described (12), with little modification. Briefly, the culture supernatants of overnight grown bacteria were centrifuged (7000 g, 30 min, 4ºC) and filtered through 0.45-µm Millipore filters. The filtrates were loaded at a flow rate of 1 ml/min at 4ºC onto columns (2 x 60 cm) of phenyl-Sepharose (Pharmacia, Uppsala, Sweden) equilibrated with 50 mM Tris-HCl, pH 8.0. Columns were then washed with 10 volumes of the same buffer. Bound protease was eluted with 50 mM Tris-HCl, pH 11.0, and the collected fractions (3.8 ml) were monitored for protein content in a spectrophotometer at 280 nm, after the pH had been adjusted to 8.0. The peak fractions were pooled and concentrated by differential centrifugation using Millipore Ultrafree filters (cut-off size, 30 kDa). Material retained by the filter was aliquoted at appropriate concentrations and kept at -20ºC in 15% glycerol. The homogeneity of the preparations was analyzed by SDS-PAGE (17).
Enzymatic assays
Hydrolysis of the intramolecularly fluorogenic quenched peptide substrates at 37ºC in 50 mM Tris-HCl, 2 mM CaCl2, pH 8.0, was monitored by measuring the fluorescence at 420 nm and after excitation at 320 nm in a Hitachi F-2000 spectrofluorometer, as previously described (18). Standard hydrolysis conditions were strictly maintained for different substrates. The enzyme concentration varied from 2.8 pM for the best substrates to 28 pM for the less susceptible ones. In most cases, substrate concentration ranged from 10 times lower than the Km to 10 times greater than the Km. The kinetic parameters were calculated by the method of Wilkinson (19). The activities of the natural and recombinant enzymes were determined using the operational substrate described by Fernandes et al. (14). One unit of ZapA activity is the amount of enzyme which hydrolyzes 1 µmol of the peptide substrate in 1 min. All enzyme assays were performed in triplicate.
HPLC analysis of substrates and of their enzymatic hydrolysis products
The peptide solutions (20-50 µM) in 50 mM Tris-HCl, pH 8.0, and 2 mM CaCl2 were incubated with 2.8 pM of the native and recombinant proteases at 37ºC for 3 h. Samples (500 µl) from the substrate and from the incubates were periodically removed for HPLC analysis until 100% hydrolysis was achieved. The hydrolysis products were separated by HPLC, collected manually, and submitted to mass spectrometry. The scissile bonds were deduced from the sequences of the substrate fragments. The HPLC conditions used for the analytical procedure were: 0.1% trifluoroacetic acid in water (solvent A) and acetonitrile-solvent A (9:1) as solvent B. The separations were performed at a flow rate of 1 ml/min using a J.T. Baker C-18 column (4.6 x 300 mm). Analytical HPLC was performed using an SPD-10AV Shimadzu UV/visible detector and an RF-10 AX fluorescence detector. For peptide purification, a semi-prep TSK C-18-120T UltraPak column (300 x 7.8 mm, 10-µm particles; LKB, Bromma, Sweden) was used with the same solvents as above, at a flow rate of 2 ml/min. In all cases, elution was followed by UV absorption (214 nm) and by fluorescence (at 420 and at 320 nm). IgA (54 µM) was incubated with 2.8 and 28 pM of ZapA for 24 h at 37ºC in the buffer described above, and with 15.6 pM of trypsin in 50 mM Tris-HCl, pH 8.0, for 3 h at 37ºC. The HPLC analyses were performed with a J.T. Baker C-8 column (4.6 x 300 mm) using the solvents described above. Absorbance was monitored at 280 nm.
Mass spectrometry
The mass spectrometric experiments were performed using a QUATTRO II triple quadruple mass spectrometer equipped with a standard electrospray probe (Micromass, Altrinchan, UK), adjusted to about 40 µl/min with the ODS-HG-5 microcolumn (0.3 x 150 mm, 5-µm particles). The mass spectrometer data acquisition and treatment system was equipped with MassLynx and MaxEnt software for handling spectra. The source temperature was maintained at 80ºC throughout all experiments and the needle voltage was kept at 3.6 kV by applying a drying gas flow (nitrogen) of 200 l/h and a nebulizer gas flow of 20 l/h. The mass spectrometer was calibrated with intact horse heart myoglobin and its typical cone-voltage-induced fragments. For reliable mass determination of the intact ß-chain of insulin and its proteolytic fragments, reduction and carboxymethylation were performed as previously described (20). Mass spectrometric detection was achieved with different parameters for each type of experiment. The peptide fragments separated by liquid chromatography were detected by scanning 50 to 2000 m/z at 6 s/scan, with a 31-V cone. Product ions from tandem mass spectrum experiments were detected by several scanning runs of the appropriate mass for each situation, using high energy (25 eV) for single charged and low collision energy (15 eV) for multiple charged precursor ions. No tandem mass spectra were recorded for peptides smaller than four amino acid residues.
Results
Purification and comparative specificity features of the native extracellular metalloprotease from P. mirabilis strain N17-12 and of the recombinant ZapA protease
The purified extracellular metalloprotease from P. mirabilis strain N17-12 was visualized as a single band (data not shown) of the same molecular mass as the purified recombinant enzyme (12). The ZapA activities, determined by a fluorimetric assay using the operational substrate peptide Abz-A-F-R-S-A-A-Q-EDDnp (14), were 1.0 and 1.21 mU/mg for the natural and the recombinant enzymes, respectively. The experiments described in this study were performed using both the natural and the recombinant ZapA, with similar results. Thus, only the data obtained with the recombinant enzyme are presented.
Hydrolysis of peptides by the recombinant metalloprotease from P. mirabilis
Fluorogenic peptides. The brakykinin-derived peptide (peptide 1) was cleaved at the Phe-Ser bond (Figure 1) with a Km of 13.6 µM and a kcat of 3.96 s-1, resulting in a catalytic efficiency of 291 mM/s. The dynorphin-derived quenched fluorescent substrate (peptide 2) was hydrolyzed at the Phe-Leu bond (Figure 1) with a Km of 2.3 µM and a kcat of 0.031 s-1, resulting in a catalytic efficiency of 13 mM/s.
Oxidized ß-chain of insulin. A number of fragments with different relative concentrations were generated and resolved by HPLC (Figure 2). No attempt was made to identify the most susceptible peptide bond within the oxidized ß-chain of insulin. The molecular mass of the major fragments derived from the incubation of the oxidized ß-chain of insulin with ZapA was determined by mass spectrometry (Table 1). The interpretation of the molecular masses of the reaction products indicated that the following peptide bonds were hydrolyzed: Val2-Asn3, Asn3-Gln4, Cys7-Gly8, His10-Leu11, Leu11-Val12, Tyr16-Leu17, Gly20-Glu21, Phe25-Tyr26, and Tyr26-Thr27 (Figure 2).
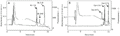 2, pH 8.0, at 37ºC for 3 h. The hydrolysis products were analyzed by HPLC as described in Material and Methods. The chromatograms in panels A and B show the elution profiles of the fragments generated by the incubation of peptide 1 and peptide 2 with ZapA, respectively. Dashed lines indicate fluorescence and continuous lines show the absorbance at 214 nm. Bk = bradykinin; Dyn = dynorphin.
2, pH 8.0, at 37ºC for 3 h. The hydrolysis products were analyzed by HPLC as described in Material and Methods. The chromatograms in panels A and B show the elution profiles of the fragments generated by the incubation of peptide 1 and peptide 2 with ZapA, respectively. Dashed lines indicate fluorescence and continuous lines show the absorbance at 214 nm. Bk = bradykinin; Dyn = dynorphin.
[View larger version of this image (16 K GIF file)]
 2, pH 8.0, at 37ºC for 3 h. The hydrolysis products were analyzed by HPLC as described in Material and Methods. The chromatogram in panel B shows the profile of the intact polypeptide, and panel C shows the production of the fragments generated by the incubation with ZapA. The bonds cleaved by ZapA, indicated by arrows in the sequence of the ß-chain of insulin (panel A), were determined by mass spectrometry as described in Material and Methods. ¯ = known cleavage sites; ß = newly detected cleavage site.
2, pH 8.0, at 37ºC for 3 h. The hydrolysis products were analyzed by HPLC as described in Material and Methods. The chromatogram in panel B shows the profile of the intact polypeptide, and panel C shows the production of the fragments generated by the incubation with ZapA. The bonds cleaved by ZapA, indicated by arrows in the sequence of the ß-chain of insulin (panel A), were determined by mass spectrometry as described in Material and Methods. ¯ = known cleavage sites; ß = newly detected cleavage site.
[View larger version of this image (16 K GIF file)]
Enzymatic activity of ZapA on secretory IgA
We were not able to observe efficient hydrolytic activity of ZapA towards IgA, even after long incubation times using an enzyme concentration ten-fold higher than the one used in the assays performed with the peptide substrates. HPLC of the incubation mixture showed that less than 2% of IgA hydrolysis had taken place after 24 h of incubation (Figure 3). Parallel control experiments (data not shown) demonstrated that i) the enzyme lost 11% of its initial activity after 24 h of incubation, and ii) when IgA was incubated at 37ºC in the presence of trypsin, less than 10% of IgA was detected after 3 h.
 2, pH 8.0, at 37ºC for 24 h without (panel A) or with (panel B) 28 pM ZapA. The incubation mixtures were analyzed by HPLC as described in Material and Methods.
2, pH 8.0, at 37ºC for 24 h without (panel A) or with (panel B) 28 pM ZapA. The incubation mixtures were analyzed by HPLC as described in Material and Methods.
[View larger version of this image (11 K GIF file)]
Discussion
Metalloproteases display a broad spectrum of action, which impairs the use of natural substrates in the study of their specificity. Therefore, we opted to use quenched fluorogenic substrates for the studies of ZapA, which had the additional advantage of being suitable for comparison to the recently developed quenched fluorogenic operational substrate of the enzyme (14).
Analysis of the hydrolysis of the peptide substrates used showed that single peptide bonds, Phe-Ser and Phe-Leu, were hydrolyzed in peptide 1 and peptide 2, respectively, with different catalytic efficiencies. While peptide 1 showed an efficiency only 23% lower than that obtained for the operational substrate, peptide 2 showed a very low efficiency, which may reflect the unfavorable influence of two highly hydrophobic and basic residues at positions P'2 and P'3.
Analysis of the products generated by the hydrolysis of the oxidized ß-chain of insulin indicates that ZapA can produce multiple cleavages in a large polypeptide, giving rise to a number of products. These data strongly suggest the broad specificity of ZapA. Except for the hydrolysis of the Val-Asn bond, all other cleavages had been previously reported (21). The peptide bonds hydrolyzed are consistent with the specificity deduced when fluorescent substrates were used (14).
Globular proteins, however, display a relatively small number of peptide bonds susceptible to the attack by metalloproteases. Thus, it was not surprising that native IgA was not well hydrolyzed by ZapA, even after long incubation times. Controls designed to measure ZapA stability and the susceptibility of IgA to hydrolysis by trypsin indicated that the resistance of IgA to hydrolysis was neither due to the loss of ZapA activity nor to the lack of susceptibility of IgA to proteolysis. These experiments assured that IgA is not a good substrate for ZapA. In contrast, serine proteases, secreted by several pathogenic bacteria such as Neisseria gonorrhea, N. meningitidis, and Haemophilus influenzae, among others, have been shown to catalyze the hydrolytic cleavage of the IgA molecule at one peptide bond in the hinge region (9). Furthermore, it is well known that the metzincin family of metalloproteases, to which ZapA belongs, requires extended substrates for optimum cleavage efficiency (22), a property not displayed by native immunoglobulins.
ZapA has been considered to have a role as virulence factor for P. mirabilis (13,23,24). It seems, however, that its role in the pathogenicity of this bacterium is not due to the hydrolysis of IgA, which would destroy an important component of the host defense system of mammalian mucous surfaces. It is possible, however, that the enzyme may be effective for microbial proliferation, destroying other bioactive molecules like defensins, involved in the innate defense, or structural components of the host cells like matrix proteins. In fact, we showed here that one bioactive peptide, bradykinin, released during the inflammatory process, was efficiently hydrolyzed by ZapA. Studies involving the hydrolysis of bioactive peptides, also important in the inflammatory process, have been reported for the alkaline metalloprotease of Pseudomonas aeruginosa, another member of the serralysin family of bacterial metalloproteases (25). A similar conclusion could be drawn from our preliminary results with a newly discovered insect defensin, which was efficiently hydrolyzed by ZapA (Palma MS, unpublished results). Moreover, we have recently shown that ZapA has a dose-dependent fibrinolytic activity on fibrin plates, hydrolyzing the a- and ß-chains but not the g-chain of fibrinogen, while it completely degrades collagen and partially hydrolyzes laminin, not being able, however, to degrade fibronectin (Fernandes BL, unpublished results).
Acknowledgments
We thank Dr. R. Belas, Biotechnology Institute, University of Maryland, Baltimore, MD, USA, for plasmid pCW101, and Dr. A.C.M. Camargo, Instituto Butantan, São Paulo, SP, Brazil, for useful suggestions concerning the biological activity of ZapA.
Address for correspondence: B.L. Fernandes, Departamento de Microbiologia, ICB II, USP, Av. Prof. Lineu Prestes, 1374, 05508-900 São Paulo, SP, Brasil. Fax: +55-11-3818-7420. E-mail: blfernan@usp.br
Research supported by FAPESP and PIBIC/CNPq/USP. Received January 11, 2001. Accepted August 9, 2001.
- 1. Häse CC & Finkelstein RA (1993). Bacterial extracellular zinc-containing metalloproteases. Microbiology Reviews, 57: 823-837.
- 2. Rao MB, Tanksale AM, Ghatge MS & Deshpande VV (1998). Molecular and biotechnological aspects of microbial proteases. Microbiology and Molecular Biology Reviews, 62: 597-635.
- 3. Kamata R, Matsumoto K, Okamura R, Yamamoto T & Maeda H (1985). The serratial 56K protease as a major pathogenic factor in serratial keratitis: clinical and experimental study. Ophthalmology, 92: 1452-1459.
- 4. Lyerly DM & Kreger AS (1983). Importance of serratia protease in the pathogenesis of experimental Serratia marcescens pneumonia. Infection and Immunity, 40: 113-119.
- 5. Bauman U (1998). Serralysin. In: Barrett AJ, Rawlings ND & Woessner JF (Editors), Handbook of Proteolytic Enzymes Academic Press, New York, 1147-1150.
- 6. Finkelstein RA, Boesman-Finkelstein M & Holt P (1983). Vibrio cholerae hemagglutinin/lectin/protease hydrolyses fibrinonectin and ovomucin: FM Burnet revisited. Proceedings of the National Academy of Sciences, USA, 80: 1092-1095.
- 7. Dalhammar G & Steiner H (1984). Characterization of inhibitor A, a protease from Bacillus thuringiensis, which degrades attacins and cecropins, two classes of antibacterial proteins in insects. European Journal of Biochemistry, 139: 247-252.
- 8. Morihara K (1974). Comparative specificity of microbial proteinases. Advances in Enzymology and Related Areas of Molecular Biology, 41: 179-243.
- 9. Plauth AG & Bachovchin WW (1994). IgA-specific prolyl endopeptidases: serine type. Methods in Enzymology, 244: 137-151.
- 10. Plauth AG & Wright A (1995). Immunoglobulin A-metallo-type specific prolyl endopeptidases. Methods in Enzymology, 248: 634-642.
- 11. Schiavo G, Benfenati F, Poulain B, Rossetto O, de Laureto PP, DasGupta BR & Montecucco C (1992). Tetanus and botulinum-B neurotoxins block neurotransmitter release by proteolytic cleavage of synaptobrevin. Nature, 359: 832-835.
- 12. Wassif C, Cheek D & Belas R (1995). Molecular analysis of a metalloprotease from Proteus mirabilis Journal of Bacteriology, 177: 5790-5798.
- 13. Walker KE, Moghaddame-Jafari S, Lockatell CV, Johnson D & Belas R (1999). ZapA, the IgA degrading metalloprotease of Proteus mirabilis, is a virulence factor expressed specifically in swarmer cells. Molecular Microbiology, 32: 825-836.
- 14. Fernandes BL, Anéas MF, Juliano L, Palma MS, Lebrun I & Portaro FCV (2000). Development of an operational substrate for ZapA, a metalloprotease secreted by the bacterium Proteus mirabilis Brazilian Journal of Medical and Biological Research, 33: 765-770.
- 15. Costa SOP, Bonato MCM & Bianco M (1982). Instability of the protease (gelatinase) trait in Proteus mirabilis I. A study of newly isolated strains. Revista de Microbiologia, 13: 35-45.
- 16. Hirata IY, Cezari MHS, Nakaie CR, Boschcov P, Ito AS, Juliano MA & Juliano L (1994). Internally quenched fluorogenic protease substrates: solid-phase synthesis and fluorescence spectroscopy of peptides containing ortho-aminobenzoyl/dinitrophenyl groups as donor-acceptor pairs. Letters in Peptide Sciences, 1: 299-308.
- 17. Laemmli UK (1970). Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature, 227: 680-685.
- 18. Chagas JR, Juliano L & Prado ES (1991). Intramolecularly quenched fluorogenic tetrapeptide substrates for tissue and plasma kallikreins. Analytical Biochemistry, 192: 419-425.
- 19. Wilkinson GN (1961). Statistical estimations in enzyme kinetics. Biochemical Journal, 80: 324-332.
- 20. van Baar BLM, Hulst AG & Wills ER (1999). Characterization of cholera toxin by liquid chromatography-electrospray mass spectrometry. Toxicon, 37: 85-98.
- 21. Maeda H & Morihara K (1995). Serralysin and related bacterial proteases. Methods in Enzymology, 248: 395-413.
- 22. Stöcker W, Grams F, Baumann U, Reinemer P, Gomis-Rüth F-X, McKay D & Bode W (1995). The metzincins - topological and sequential relations between the astacins, adamalysins, serralysins, and matrixins (collagenases) define a superfamily of zinc-peptidases. Protein Science, 4: 823-840.
- 23. Loomes LM, Senior BW & Kerr MA (1990). A proteolytic enzyme secreted by Proteus mirabilis degrades immunoglobulins of the immunoglobulin A1 (IgA1), IgA2, and IgG isotypes. Infection and Immunity, 58: 1979-1985.
- 24. Allison C, Lai H & Hughes C (1992). Co-ordinate expression of virulence genes during swarm cell differentiation and population migration of Proteus mirabilis Molecular Microbiology, 6: 1583-1591.
- 25. Louis D, Bernillon J & Wallach JM (1998). Specificity of Pseudomonas aeruginosa serralysin revisited, using biologically active peptides as substrates. Biochimica et Biophysica Acta, 1387: 378-386.
Correspondence and Footnotes
Publication Dates
-
Publication in this collection
07 Nov 2001 -
Date of issue
Nov 2001
History
-
Received
11 Jan 2001 -
Accepted
09 Aug 2001


