Abstract
PURPOSE: To investigate the effects of ischemic preconditioning (IPC) on the expression of pro and anti-apoptotic genes in rat endothelial cells undergoing enteric ischemia (I) and reperfusion (R). METHODS: Thirty rats underwent clamping of the superior mesenteric vessels. Sham group (GS) laparotomy only; Ischemia (GI): intestinal ischemia (60 min); Ischemia and Reperfusion (GIR): ischemia (60 min) and reperfusion (120 min); Ischemia and intestinal ischemic preconditioning (GI + IPC) : 5 minutes of ischemia followed by 10 min of reperfusion before sustained ischemia (60 min) ischemia and reperfusion and IPC (GIR + IPC): 5 min ischemia followed by 10 min of reperfusion before sustained ischemia (60min) and reperfusion (120 min). Rat Endothelial Cell Biology (PCR array) to determine the expression of genes related to endothelial cell biology. RESULTS: Gene expression of pro-apoptotic markers (Casp1, Casp6, Cflar, Fas, and Pgl) was down regulated in GI+IPC and in GIR + IPC. In contrast, the expression of anti-apoptotic genes (Bcl2 and Naip2), was up-regulated in GI + IPC and in GIR + IPC. CONCLUSION: Ischemic preconditioning may protect against cell death caused by ischemia and reperfusion.
Ischemia; Reperfusion Injury; Endothelial Cells; Gene Expression Profiling; Rats
2 - ORIGINAL ARTICLE
ISCHEMIA-REPERFUSION
Ischemic preconditioning and the gene expression of enteric endothelial cell biology of rats submitted to intestinal ischemia and reperfusion1 1 Research performed at Laboratory of Experimental Surgery and Operative Technique, Department of Morphology, Laboratory of Histology and Structural Biology, Department of Pharmacology and Department of Biochemistry, Federal University of Sao Paulo (UNIFESP), Brazil.
Murched Omar TahaI; Regiane Miranda FerreiraII; Nabiha Saadi Abrahão TahaIII; Hugo Pequeno MonteiroIV; Afonso Caricati-NetoV; Itamar Souza Oliveira-JúniorVI; Djalma José FagundesVII
IPhD, Associate Professor, Operative Technique and Experimental Surgery Division, Department of Surgery, UNIFESP, Sao Paulo-SP, Brazil. Main author. Conception, design, intellectual and scientific content of the study
IIBSc, PhD, Postdoctoral Fellow, Pharmacology Department, UNIFESP, Sao Paulo-SP, Brazil. Acquisition and interpretation of biochemical data, manuscript writing
IIIPhD, Postdoctoral Fellow, Gynecology Department, UNIFESP, Sao Paulo-SP, Brazil. Acquisition of pharmacological data, manuscript writing
IVBSc, PhD, Full Professor, Department of Biochemistry, UNIFESP, Sao Paulo-SP, Brazil. Interpretation of biochemistry data, manuscript writing
VBSc, PhD, Associate Professor, Pharmacology Department, UNIFESP, Sao Paulo-SP, Brazil. Interpretation of pharmacological data, manuscript writing
VIAssociate Professor, Division of Anesthesiology, Pain and Intensive Therapy, UNIFESP, São Paulo-SP, Brazil. Interpretation of biological data and manuscript writing
VIIPhD, Associate Professor, Operative Technique and Experimental Surgery Division, Department of Surgery, UNIFESP, Sao Paulo-SP, Brazil. Conception and critical revision of the study
Correspondence Correspondence: Murched Omar Taha Rua Joaquim Távora, 1599/16º andar 04015-003 São Paulo - Brasil Tel.: (55 11)5084-0100/99982-0001 Fax: (55 11)5571-0233 taha@uol.com.br
ABSTRACT
PURPOSE: To investigate the effects of ischemic preconditioning (IPC) on the expression of pro and anti-apoptotic genes in rat endothelial cells undergoing enteric ischemia (I) and reperfusion (R).
METHODS: Thirty rats underwent clamping of the superior mesenteric vessels. Sham group (GS) laparotomy only; Ischemia (GI): intestinal ischemia (60 min); Ischemia and Reperfusion (GIR): ischemia (60 min) and reperfusion (120 min); Ischemia and intestinal ischemic preconditioning (GI + IPC) : 5 minutes of ischemia followed by 10 min of reperfusion before sustained ischemia (60 min) ischemia and reperfusion and IPC (GIR + IPC): 5 min ischemia followed by 10 min of reperfusion before sustained ischemia (60min) and reperfusion (120 min). Rat Endothelial Cell Biology (PCR array) to determine the expression of genes related to endothelial cell biology.
RESULTS: Gene expression of pro-apoptotic markers (Casp1, Casp6, Cflar, Fas, and Pgl) was down regulated in GI+IPC and in GIR + IPC. In contrast, the expression of anti-apoptotic genes (Bcl2 and Naip2), was up-regulated in GI + IPC and in GIR + IPC.
CONCLUSION: Ischemic preconditioning may protect against cell death caused by ischemia and reperfusion.
Key words: Ischemia. Reperfusion Injury. Endothelial Cells. Gene Expression Profiling. Rats.
Introduction
Ischemia followed by reperfusion (I/R) promotes multiple biological responses that cause tissue injury and organ damage. These responses include the generation of reactive oxygen species (ROS), the activation of transcription factors, the production of cytokines and eicosanoids, the expression of inducible nitric oxide synthase (NOS) and adhesion molecules, injury to blood vessel endothelium, neutrophil infiltration, platelet adhesion, microscopic circulating deficit, and apoptosis1. In the intestine, I/R injury is the major factor responsible for intestinal dysfunction and is a determinant of patient prognosis after transplantation. Thus, novel strategies to prevent or ameliorate intestinal I/R injury are necessary to improve patient outcomes following organ transplantation2,3.
I/R injury is considered to be the principal trigger of systemic inflammatory response syndrome (SIRS) and is a crucial process associated with an increased incidence of chronic rejection after transplantation. SIRS often progresses to multiple organ failure, resulting in up to 80% mortality1. Reperfusion of ischemic tissues is often associated with microvascular dysfunction that manifests as impaired endothelium-dependent dilation in arterioles, enhanced fluid filtration and leukocyte plugging in capillaries, and leukocyte trafficking and plasma protein extravasations in post capillary venules1.
Endothelial cells modulate many important vascular functions, such as vascular tone, inflammation, homeostasis, and angiogenesis4. Activated endothelial cells in all segments of the microcirculation produce more oxygen radicals, but less nitric oxide, in the initial period following reperfusion. The resulting imbalance between superoxide and nitric oxide in endothelial cells leads to the production and release of inflammatory mediators (e.g., platelet-activating factor and tumor necrosis factor) and enhances the biosynthesis of adhesion molecules that mediate leukocyte-endothelial cell adhesion1.
The inflammatory mediators released as a consequence of reperfusion also appear to activate endothelial cells in remote organs that are not exposed to the initial ischemic insult1. This remote response to I/R can result in the leukocyte-dependent microvascular injury that is characteristic of multiple organ dysfunction syndrome1.
Intestinal I/R injury is multifactorial and involves impaired blood flow reconstitution, increased expression of adhesion molecules, neutrophil activation, activation of antigen-presenting cells (APCs), cytokine/chemokine release by infiltrating leukocytes, oxidative stress, and endothelial cell apoptosis2,3. It is assumed that the initial endothelial damage plays a pivotal role in intestinal I/R, but limited access to endothelial tissue is a major constraint when investigating the cellular mechanisms of vascular inflammation4.
In 1986, Murry et al.5 observed that ischemic preconditioning (IPC) for a short duration before long-term I/R reduced the severity of the subsequent I/R injury. This phenomenon has since been documented in a variety of organs. Because it is easy, efficient, and cost-effective, IPC has been used in clinical medicine. The duration of IPC necessary to acquire tolerance to ischemia differs for every internal organ. There are two temporally and mechanistically distinct types of protection afforded by this adaptive response: acute and delayed preconditioning. In general, the protective effects of acute preconditioning are independent of protein synthesis, whereas the effects of delayed preconditioning require protein synthesis6. Some factors, such as protein kinase C, are involved in both types of preconditioning responses6. Despite intensive research, the exact cellular and biochemical mechanisms by which IPC confers protection in the intestine have not been elucidated.
A clearer understanding of cellular signaling during intestinal I/R could provide better insight into the protective mechanism of IPC. Recent studies have shown that the use of the quantitative real time polymerase chain reaction method (qRT-PCR) to evaluate specific gene expression during I/R could be crucial to understanding the protective mechanism of IPC.6 In the present work, we used the Rat Endothelial Cell Biology PCR Array to characterize gene expression in endothelial cells in rats subjected to IPC prior to intestinal I/R injury. This array profiles the expression of 84 genes related to endothelial cell biology and includes genes involved in permissibility and vessel tone, angiogenesis, endothelial cell activation and endothelial cell injury.
Methods
The experimental protocol (0109/11) was approved by the Ethics Committee of the Federal University of Sao Paulo (UNIFESP), on according to the recommendations of the international legislations on animal protection. The study was designed as a randomized controlled trial with a blinded assessment of the outcome.
Thirty male Wistar-EPM1 rats (Center for the Development of Experimental Models for Medicine and Biology - CEDEME-UNIFESP) that weighed 250 to 300g were housed under controlled temperature and light conditions with a 12-h light-dark cycle. The animals had free access to water and standard pellet chow until 6 h prior to the surgical procedures. All procedures were conducted in the laboratories of Experimental Surgery, Department of Surgery (UNIFESP).
The animals underwent anesthesia, laparotomy, and exposure of the upper mesenteric vessels and were randomly assigned to one of five groups:
Sham group (SG): six animals with exposure of the upper mesenteric vessels without clamping of the upper mesenteric vessels;
Ischemia group (IG): six animals in which the upper mesenteric vessels were clamped for 60 min;
Ischemia and reperfusion group (IRG): six animals in which the upper mesenteric vessels were clamped for 60 min, followed by 120 min of reperfusion;
Ischemia and intestinal preconditioning group (IG+IPC): six animals exposed to 5 min of ischemia followed by 10 min of reperfusion before the sustained 60 min of ischemia;
Ischemia and reperfusion and IPC group (IRG+IPC): six animals subjected to 5 min of ischemia followed by 10 min of reperfusion before starting the sustained 60 min of ischemia and 120 min of reperfusion.
Anesthesia and surgical procedures
After 6h of fasting for solids and liquids, the animals were anesthetized with an intramuscular injection of a mixture containing 80 mg/kg ketamine and 10 mg/kg xylazine and provided with oxygen through a mask. Body temperature was maintained at 37.8oC using a homeothermic soft blanket (40 x 70 cm) and monitored with a rectal temperature probe.
Under aseptic conditions, all 30 animals underwent a midline laparotomy. The upper mesenteric vessels were carefully dissected, and the mesenteric artery and vein were identified and occluded with small vascular clamps. Ischemia was confirmed by observing the pale appearance of the clamped intestine and the lack of arterial beating. After 60 min of ischemia, the clamps were removed, and reperfusion was evaluated based on immediate color recovery and arterial beating. The surgical wounds remained covered with wet gauze wrappings throughout the experiment to minimize evaporative loss.
After each procedure, intestinal segments were removed 20 cm from the duodenum-jejunum flexure. A 3-cm jejunal segment was opened longitudinally, gently washed in saline solution, wrapped in aluminum foil, and placed in liquid nitrogen in less than 1.5 min.
After the samples were collected, the animals received an intravenous injection (0.3 ml/kg) of a combination of embutamide, mebezone and tetracain which promotes rapid death.
RNA preparation and quantitative real-time RT-PCR
The Rat Endothelial Cell Biology RT² Profiler PCR array from SABiosciences (Frederick, Maryland; catalog # PAMM-065) was performed according to the manufacturer 's protocol. Briefly, total RNA was extracted from S and I/R small bowel tissues using TRIzol reagent (Life Technologies, Grand Island, NY, USA) and purified using an RNeasy MiniKit (QIAGEN, USA). The concentration of each total RNA sample was determined by spectrophotometer, and the quality was assessed by electrophoresis on 2% agarose gels. The first strand complementary DNA (cDNA) was synthesized using 1 µg of total RNA and the RT² First Strand Kit (SABiosciences). Equal amounts of cDNA and the Master Mix SYBR® Green qPCR Mastermix (SABiosciences, QIAGEN Company) were distributed to each well of the PCR array plate containing the pre-dispensed gene-specific primer sets. PCR was performed according to the manufacturer 's instructions in 96-well plates to detect the expression of 84 genes related to oxidative stress, five housekeeping genes (ACTB, Gapdh, Hsp90ab1, Hprt1, Gusb) used for normalizing the PCR array data, and one negative control for genomic DNA contamination. The negative control primer set specifically detects non-transcribed, repetitive genomic DNA with a high level of sensitivity. Three wells of reverse transcription controls (RTCs) were used to verify the efficiency of the RT reaction with a qPCR assay that specifically detects template synthesized from the RNA control of the first strand synthesis kit. The replicate positive PCR controls (PPCs) were used to determine the efficiency of the polymerase chain reaction itself. These controls use a pre-dispensed artificial DNA sequence and a specific primer set to detect it. The two sets of replicate control wells (RTCs and PPCs) also test for inter-well and intra-plate consistency. The instrument 's software (MxPro Equipment Real Time Systems, Stratagene, GE, Co) calculates the threshold cycle (Ct) values for all of the genes in the array. Finally, it performs pair-wise comparisons by calculating fold changes in gene expression from the raw threshold cycle data using the ΔΔCt method. The method used in our study to determine the relative expression levels of the genes of interest in each sample is contained in the spreadsheet for PCR Array Data Analysis v3.3 (SABiosciences Qiagen, Co).
Statistical analysis
The analysis of gene expression by real-time PCR used in this work represents a relative quantification of genes of interest. To support the analysis six endogenous controls were part of the PCR reactions array for each of the samples tested; genes whose expression was used do not present a statistically significant variation between samples. The control samples for each experimental group were used as reference baseline. The results were transformed into log2 scale for the calculation of averages and standard errors and for statistical analysis. Fold-Change [2^ (- Delta Delta Ct)] is the normalized gene expression [2^ (- Delta Ct)] in the Test Sample divided the normalized gene expression [2^ (- Delta Ct)] in the Control Sample. Fold-Regulation represents fold-change results in a biologically meaningful way. It was considered that fold-change values greater than one indicates a positive or an up-regulation, and the fold-regulation is equal to the fold-change. The p values are calculated based on a Student's t-test of the replicate 2^ (- Delta Ct) values for each gene in the control group and treatment groups. P values less than 0.05 was considered significant in all experiments and indicated by symbols.
Results
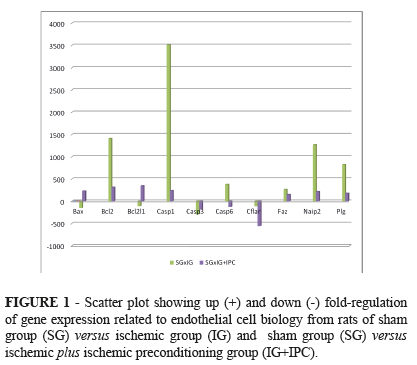
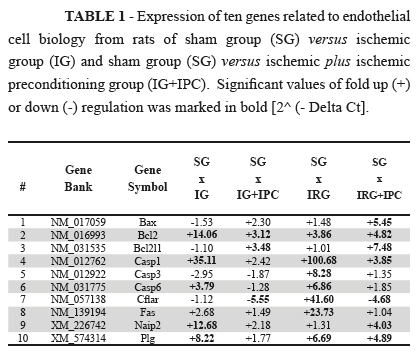
Table 1 shows the expression levels of ten genes related to endothelial cell biology in the rats belonging to the SG, IG, IG+IPC, IRG and IRG+IPC groups. This table lists the genes, gene abbreviations (alphabetical order), and gene descriptions along with the fold upregulation or down regulation of each gene and p values (bold). These data are plots in Figures 1 and 2.
Discussion
Our results clearly demonstrate a down regulation of the genes related to endothelial cell biology in the rats that underwent IPC. The down regulation of these genes was observed in the IG+IPC and IRG+IPC. These results are consistent with the hypothesis that IPC promotes the decreased expression of genes that modulate the lesions produced by I/R.
Whether the expression of certain genes corresponds to effective protein synthesis remains to be ascertained. Reports in the literature assume that proteins having matching or similar profiles tend to be functionally linked1. Those proteins that function together in a pathway or complex are likely to evolve in a correlated fashion. To enhance our analysis, we sought to group genes according to the similarity of their previously characterized cellular activity. We focused on two clusters that are relevant to intestinal I/R: the caspase activation cluster (Bax, Casp1, Casp3, Casp6, Cflar) and the induction of apoptosis cluster (Bax, Casp1, Casp3, Plg, Fas). In addition, we assessed the expression of anti-apoptotic genes (Bcl2, Bcl2l1, Naip2) that are important for intestinal I/R.
Induction of apoptosis cluster (Bax, Casp1, Casp3, Casp6, Plg, Fas)
It is well established that apoptosis plays an essential role in regulating development, homeostasis, and immune defense by removing redundant or abnormal cells from organisms7,8. A delicate balance between pro-apoptotic mechanisms and anti-apoptotic mechanisms determines whether a cell death signal can activate the execution of the apoptotic program9,10.
The cysteine-aspartic acid protease (caspase) family is involved in the signal transduction pathways of apoptosis, necrosis and inflammation.11 Caspases are the primary executioners of apoptosis. Death stimuli trigger the activation of initiator caspases, which in turn activate the effectors caspases12. The sequential activation of caspases plays a central role in the execution phase of apoptosis. There are two families of caspases: the inflammatory caspases and the apoptotic caspases, and each of these families can be further divided into initiator and effectors subgroups13. The initiator caspases (caspases 1, 4, 5, 8, 9, 10, 11, and 12) are activated by, and interact with, upstream adaptor molecules through protein-protein interaction domains known as CARD and DED13. Effectors caspases (caspases 3, 6, and 7) are responsible for cleaving downstream substrates and are sometimes referred to as the executioner caspases13. Initiator caspases, such as caspase 8, may be directly activated by death receptors such as Fas. Caspase 9 is recruited to the apoptosome formed during apoptosis, whereas caspases 1 and 5 can form part of the inflammasome, a key mediator of cytokine processing during inflammation.
Caspase 1 (Casp1) was identified based on its ability to proteolytically cleave and activate the inactive precursor of interleukin-1, a cytokine involved in processes such as inflammation, septic shock, and wound healing11. In the present study, Casp1 gene expression was up regulated in the IG (+35.11) and IRG (+100.68) but significantly reduced in the IG+IPC (+2.42) and IRG+IPC (+3.85). This decrease in Casp1 gene expression in the IPC groups can be interpreted as a lack of activation of the caspase cascade in response to cellular signaling stimulated in I/R injury, reinforcing the protective effect of IPC. The Casp3 protein cleaves and activates caspases 6, 7 and 9, and caspase 3 itself is processed by caspases 8, 9 and 1013. In our work, the Casp3 gene was up regulated in the IRG (+8.28) group, but its expression was significantly reduced in the IRG+IPC (+1.35). The caspase 6 protein is processed by caspases 7, 8 and 10 and is thought to function as a downstream enzyme in the caspase activation cascade13. We observed that the Casp6 gene was up regulated in the IG (+3.79) and IRG (+100.68), but the expression of this gene was significantly reduced in the IG+IPC (+2.42) and IRG+IPC (+3.85). These findings regarding Casp 1, 3 and 6 expressions support the idea that cellular signaling stimulated by I/R injury is altered by IPC.
The protein encoded by the Cflar gene is a regulator of apoptosis11 that is structurally similar to caspase 8, however, the encoded protein lacks caspase activity and appears to be cleaved into two peptides by caspase 8. The apoptosis regulator protein c-FLIP is a crucial link between cell survival and cell death pathways in mammalian cells as it inhibits TNFRSF6-mediated apoptosis11. Full length and shorter isoforms have been shown either to induce apoptosis or to reduce TNFRSF-triggered apoptosis12. We observed that the Cflar gene was down regulated in the IG + IPC (-5.55) and IRG+IPC (-4.68) groups when compared to the IG and IRG. In the IRG, Cflar gene expression was up regulated, accounting for the protective effect of IPC.
The BAX protein forms a heterodimer with Bcl2 and is an apoptotic activator. Bax increases the opening of the mitochondrial voltage-dependent anion channel (VDAC), leading to a loss in membrane potential and the release of cytochrome c, which in turn activates the apoptotic cascade13. In our study, Bax gene expression was not down regulated in the IG + IPC and IRG+IPC when compared to the IG and IRG. Similar results were obtained for Fas gene expression. These results suggest that cellular signaling stimulated by IPC was unable to interfere with Bax and Fas gene expression. The protein encoded by the Fas gene is a member of the TNF receptor superfamily14. Known as a death receptor, FAS protein is located on the surface of cells15. The interaction of this receptor with its ligand results in the formation of a death-inducing signaling complex that includes FAS-associated death domain protein (FADD), caspase 8, and caspase 1014.
The protein encoded by the Plg gene is a secreted blood zymogen that is activated by proteolysis and converted to plasmin and angiostatin16. Plasmin dissolves fibrin in blood clots and is an important protease in many other cellular processes, whereas angiostatin inhibits angiogenesis16. In the present study, the Plg gene expression was upregulated in the IG (+8.22) and IRG (+6.69) groups, but IPC did not interfere with the expression of this gene.
Anti-apoptotic genes (Bcl2, Bcl2l1, Naip2)
Following TNF-R1 and Fas activation in mammalian cells, a balance between pro-apoptotic (BAX, BID, BAK, or BAD) and anti-apoptotic (BCL-Xl and BCL-2) members of the Bcl-2 family is established17. In the present study, Bcl2l1 gene expression was up regulated in the IG+IPC (+3.48) and IRG+IPC (+7.48) groups, but not in the IG and IRG, suggesting that IPC stimulated the upregulation of the anti-apoptotic protein encoded by the Bcl2l1 gene. In contrast, Bcl2 gene expression was up regulated in the IG (+14.06), IRG (+3.86), IG+IPC (+3.12) and IRG+IPC (+4.82), suggesting that expression of the anti-apoptotic protein Bcl2 was not differentially stimulated by IPC. It is important to note that BCL-2 proteins can act as anti- or pro-apoptotic regulators that are involved in a wide variety of cellular activities10.
Neuronal apoptosis inhibitory protein (NAIP) is a protein that inhibits the apoptosis of neurons and other cell types induced by a variety of signals18. At least three Naip genes encode full-length mRNAs that correspond to possible functional proteins (NAIP1, 2 and 3)18. The NAIP mRNA transcripts are expressed in macrophage-rich tissues, such as spleen, lung, and liver, and are abundant in primary macrophages18. In the present study, Naip2 gene expression was up regulated in the IRG+IPC (+4.03) when compared to the IRG (+1.31), suggesting that IPC stimulated the upregulation of the anti-apoptotic protein NAIP2.
Importantly, these findings confirm the protective effect of IPC against ischemia-mediated cell injury. The protective effect of IPC appears be correlated with the regulation of cellular adhesion, improvement in energetic and substrate metabolism, and activation of intracellular signal transduction
Conclusion
Ischemic preconditioning may protect against cell death caused by ischemia and reperfusion.
Acknowledgments
The native English-speaking editors of American Journal Experts, Profa. Maria do Carmo Negrini Fagundes for assistance with translation and formatting of the article, biologist Marcos de Azevedo Junior for providing technical assistance for sequencing of RT-PCR and Prof. Ismael Dale Cotrim Guerreiro da Silva for helpful assistance to analyze our data.
Received: October 24, 2012
Review: December 20, 2012
Accepted: January 22, 2013
Conflict of interest: none
Financial source: Sao Paulo Research Foundation (FAPESP)
- 1. Carden DL, Granger DN. Pathophysiology of ischaemia-reperfusion injury. J Pathol. 2000;190:255-66.
- 2. Ritter T, Kupiec-Weglinski JW. Gene therapy for the prevention of ischemia/reperfusion injury in organ transplantation. Curr Gene Ther. 2005;5:101-9.
- 3. Ritter T, Nosov M, Griffin MD. Gene therapy in transplantation: Toward clinical trials. Curr Opin Mol Ther. 2009;11:504-12.
- 4. Onat D, Jelic S, Schmidt AM, Pile-Spellman J, Homma S, Padeletti M, Jin Z, Le Jemtel TH, Colombo PC, Feng L. Vascular endothelial sampling and analysis of gene transcripts: a new quantitative approach to monitor vascular inflammation. J Appl Physiol. 2007;103:1873-8.
- 5. Murry CE, Jennings RB, Reimer KA. Preconditioning with ischemia: a delay of lethal cell injury in ischemic myocardium. Circulation. 1986;74:1124-36.
- 6. Wang Z, Hernandez F, Pederiva F, Andrés AM, Leal N, Burgos E, Martínez MP, Molina M, Santamaría ML, Tovar JA. Ischemic preconditioning of the graft for intestinal transplantation in rats. Pediatr Transplant. 2011;15:65-9.
- 7. Kerr JF, Wyllie AH, Currie AR. Apoptosis: a basic biological phenomenon with wide-ranging implications in tissue kinetics. Br J Cancer. 1972;26:239-57.
- 8. Wei Y, Fan T, Yu M. Inhibitor of apoptosis proteins and apoptosis. Acta Biochim Biophys Sin. 2008;40:278-88.
- 9. Schimmer A. Inhibitor of apoptosis proteins: translating basic knowledge into clinical practice. Cancer Res. 2004;64:7183-90.
- 10. Nachmias B, Ashhab Y, Ben-Yehuda D. The inhibitor of apoptosis protein family (IAPs): an emerging therapeutic target in cancer. Semin Cancer Biol. 2004;14:231-43.
- 11. Danial NN, Korsmeyer SJ. Cell death: critical control points. Cell 2004;116:205-19.
- 12. Fan TJ, Han LH, Cong RS, Liang J. Caspase family proteases and apoptosis. Acta Biochem Biophys Sin. 2005;37:719-27.
- 13. Yuan J, Horvitz HR. A First insight into the molecular mechanisms of apoptosis. Cell. 2004;116:53-6.
- 14. Siegel RM, Chan FK, Chun HJ, Lenardo MJ. The multifaceted role of Fas signaling in immune cell homeostasis and autoimmunity. Nat Immunol. 2001;1:469-74.
- 15. Wajant H. The Fas signaling pathway: more than a paradigm. Science. 2002;296:16356.
- 16. Gutiérrez-Fernández A, Parmer RJ, Miles LA. Plasminogen gene expression is regulated by nerve growth factor. J Thromb Haemost. 2007;5:1715-25.
- 17. Murphy KM, Ranganathan V, Farnsworth ML, Kavallaris M, Lock RB. "Bcl-2 inhibits Bax translocation from cytosol to mitochondria during drug-induced apoptosis of human tumor cells. Cell Death Differ. 2000;7:102-11.
- 18. Endrizzi MG, Hadinoto V, Growney JD, Miller W, Dietrich WF. Genomic sequence analysis of the mouse Naip gene array. Genome Res. 2000;10:1095-102.
Publication Dates
-
Publication in this collection
06 Mar 2013 -
Date of issue
Mar 2013
History
-
Received
24 Oct 2012 -
Accepted
22 Jan 2013 -
Reviewed
20 Dec 2012