Abstracts
Nucleotide synthesis is of central importance to all cells. In most organisms, the purine nucleotides are synthesized de novo from non-nucleotide precursors such as amino acids, ammonia and carbon dioxide. An understanding of the enzymes involved in sugarcane purine synthesis opens the possibility of using these enzymes as targets for chemicals which may be effective in combating phytopathogen. Such an approach has already been applied to several parasites and types of cancer. The strategy described in this paper was applied to identify sugarcane clusters for each step of the de novo purine synthesis pathway. Representative sequences of this pathway were chosen from the National Center for Biotechnology Information (NCBI) database and used to search the translated sugarcane expressed sequence tag (SUCEST) database using the available basic local alignment search tool (BLAST) facility. Retrieved clusters were further tested for the statistical significance of the alignment by an implementation (PRSS3) of the Monte Carlo shuffling algorithm calibrated using known protein sequences of divergent taxa along the phylogenetic tree. The sequences were compared to each other and to the sugarcane clusters selected using BLAST analysis, with the resulting table of p-values indicating the degree of divergence of each enzyme within different taxa and in relation to the sugarcane clusters. The results obtained by this strategy allowed us to identify the sugarcane proteins participating in the purine synthesis pathway.
A via de síntese de purino nucleotídeos é considerada uma via de central importância para todas as células. Na maioria dos organismos, os purino nucleotídeos são sintetizados ''de novo'' a partir de precursores não-nucleotídicos como amino ácidos, amônia e dióxido de carbono. O conhecimento das enzimas envolvidas na via de síntese de purinas da cana-de-açúcar vai abrir a possibilidade do uso dessas enzimas como alvos no desenho racional de inibidores no combate a agentes fitopatogênicos, como esta sendo feita com diversos parasitos e células cancerosas. A seguinte estratégia esta sendo utilizada na identificação de genes de cana-de-açúcar para cada membro da via de síntese de purinas: Seqüências representativas dos genes que compões a via foram escolhidas do banco de dados NCBI. Essas seqüências de peptídeos estão sendo utilizadas em buscas ao banco de dados gerado pelo SUCEST pelo programa BLAST (implementação tBLASTn). Alinhamentos com os clusters de cana-de-açúcar são posteriormente analisados para sua significância estatística pela implementação PRSS3 do algoritmo conhecido como Monte Carlo shuffling. Para calibrar a análise dos resultados de PRSS3, foram empregadas seqüências conhecidas de diferentes taxas ao longo da arvore filogenética. Essas seqüências são comparadas duas a duas e com o cluster da cana-de-açúcar. A tabela de valores-p resultante indica o grau estatístico de similaridade e divergência entre as seqüências já descritas e entre essas e os clusters de cana-de-açúcar. Os resultados obtidos dessas análises estão descritos neste artigo.
Identification of sugarcane genes involved in the purine synthesis pathway
Mario A. Jancso1, Susana A. Sculaccio1,2 and Otavio H. Thiemann1*
1Laboratory of Protein Crystallography and Structural Biology, Physics Institute of São Carlos, University of São Paulo - USP, Av. Trabalhador Sãocarlense 400, 13566-590 São Carlos, SP, Brazil.
2Department of Genetics and Evolution, Federal University of São Carlos, São Carlos, Brazil.
Corresponding author: Otavio H. Thiemann. E-mail: thiemann@if.sc.usp.br.
ABSTRACT
Nucleotide synthesis is of central importance to all cells. In most organisms, the purine nucleotides are synthesized de novo from non-nucleotide precursors such as amino acids, ammonia and carbon dioxide. An understanding of the enzymes involved in sugarcane purine synthesis opens the possibility of using these enzymes as targets for chemicals which may be effective in combating phytopathogen. Such an approach has already been applied to several parasites and types of cancer. The strategy described in this paper was applied to identify sugarcane clusters for each step of the de novo purine synthesis pathway. Representative sequences of this pathway were chosen from the National Center for Biotechnology Information (NCBI) database and used to search the translated sugarcane expressed sequence tag (SUCEST) database using the available basic local alignment search tool (BLAST) facility. Retrieved clusters were further tested for the statistical significance of the alignment by an implementation (PRSS3) of the Monte Carlo shuffling algorithm calibrated using known protein sequences of divergent taxa along the phylogenetic tree. The sequences were compared to each other and to the sugarcane clusters selected using BLAST analysis, with the resulting table of p-values indicating the degree of divergence of each enzyme within different taxa and in relation to the sugarcane clusters. The results obtained by this strategy allowed us to identify the sugarcane proteins participating in the purine synthesis pathway.
INTRODUCTION
The purine nucleotide synthesis and recycling (or salvage) pathways are of central importance to all living organisms (Marr, 1991) because they provide the purines (adenine and guanine) necessary for nearly all biochemical processes such as DNA and RNA metabolism, the biosynthesis intermediates ATP, GTP and coenzymes as well as being an important metabolic regulator in cell signaling (Stryer, 1995). Due to their importance, purine synthesis pathways have been investigated as potential targets for chemotherapy in several different scenarios (Marr, 1991; Ullman and Carter, 1997; Marr and Ullman, 1995). Cancer cells are dependent on the purine salvage pathway to provide the large quantity of purine nucleotides for RNA synthesis because of the increased intracellular synthesis rates of cancerous cells. Due to differences between host and pathogens, the purine synthesis enzymes have been proposed as potential anti-parasitic targets since several parasites are purine auxotrophs, lacking the de novo synthesis pathway and relying entirely on the salvage pathway.
The de novo purine synthesis pathway is composed of eleven enzymatic steps leading to the formation of inosinate (Figure 1). The pathway initiates with the synthesis of 5-phosphoribosyl-1-pyrophosphate (PRPP) from ribose-5-phosphate and ATP, a reaction catalyzed by PRPP synthetases (PRS; EC: 2.7.6.1). PRS is an important enzyme involved in the salvage as well as de novo pathway because PRPP is a substrate for the major salvage enzymes adenine-phosphoribosyl-transferase (APRT) and hypoxanthine-guanine-phosphoribosyl-transferase (HGPRT).
Due to its cellular importance and potential as a chemotherapeutic target, the purine synthesis and salvage pathways have been extensively studied in several organisms. Most of the genes encoding for the proteins involved in these pathways have been cloned from several organisms. Interestingly, the genetic organization varies between organisms across diverse taxa and some remarkable gene fusions have been described for several of the enzymes participating on this pathway.
In this paper we describe the identification of the eleven enzymes from the sugarcane expressed sequence (SUCEST) tag project homologous to known de novo purine synthesis proteins. Several genes are represented by full-length expressed sequence tags (ESTs) and will allow further sequence and functional analysis. Better understanding of the enzymes involved in sugarcane purine synthesis will help elucidate the genetic components of this important pathway in sugarcane and may open the possibility of using such enzymes as a target for chemicals which may be effective in combating phytopathogens (Kubinyi, 1998).
MATERIAL AND METHODS
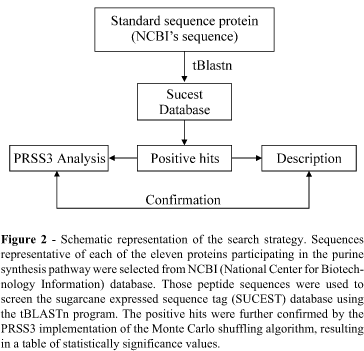
The search strategy employed is summarized in Figure 2. Representative protein sequences for each member of the purine de novo synthesis pathway (Figure 1) were chosen from the NCBI database (National Center for Biotechnology Information, www.ncbi.nlm.nih.gov). The first approach was to select protein sequences of organisms as closely related to sugarcane in the phylogenetic tree as possible, to avoid possible sequence divergence effects in our search methodology. In three cases (Glutamine-PRPP-amidotransferase, Adenylosuccinate lyase and 5-aminoimidazole-4-carboxamide ribonucleotide transformylase/inosine monophosphate cyclohydrolase bifunctional enzyme) sequences from chicken, protozoa and rat had to be used as representatives due to the lack of reliable full-length plant homologues. In those cases, sequence divergence was estimated to be low by comparison of the available homologues with the Monte Carlo shuffling strategy implemented by the PRSS3 program (http://fasta.bioch.virginia.edu/fasta/prss.htm).
The selected peptide sequences were used to search the entire translated SUCEST database with the available BLAST facility (tBLASTn) (Altschul et al. 1990; Madden et al. 1996; Zhang and Madden, 1997). Sugarcane clusters retrieved with the BLAST program were further tested for the statistical significance of the alignment by the PRSS3 program. To calibrate the Monte Carlo analysis, known protein sequences of divergent taxa along the phylogenetic tree were used. Those sequences were compared to each other and to the clusters using the PRSS3 program. The resulting table of p-values aided to estimate the degree of divergence of each enzyme between different taxa and in relation to the sugarcane clusters. The alignments and the results from the PRSS3 program analysis were taken into consideration to confirm a selected cluster as a homologue to a known purine synthesis pathway member as described in Figure 2. The PRSS3 comparison tables are not presented here due to lack of space but they are available at the sugarcane data-mining site (http://sucest.lad.ic.unicamp.br/cgi-bin/prod/mining-reports/mining-reports.pl).
The representative enzyme sequences for each member in the de novo pathway and the corresponding accession numbers are: PRPP synthetase (Arabidopsis thaliana, CAA63552) (Krath et al., 1999), Glutamine-PRPP-amidotransferase (Rattus norvegicus, A46088) (Iwahana et al., 1993), glycinamide ribonucleotide (GAR) synthetase (Arabidopsis thaliana, P52420) (Schnorr, et al., 1994), GAR transformylase (Arabidopsis thaliana, P52422) (Schnorr, et al., 1994), Formylglycinamide ribonucleotide (FGAM) synthetase (Glycine max, AAF21596, unpublished direct submission), 5-aminoimidazole ribonucleotide (AIR) synthetase (Arabidopsis thaliana, Q05728) (Senecoff and Meagher, 1993), AIR carboxylase (Arabidopsis thaliana, T02535, unpublished direct submission), 5-aminoimidazole-4-N-succinocarboxamide ribonucleotide (SAICAR) synthetase (Arabidopsis thaliana, P38025, unpublished direct submission), adenylosuccinate lyase (Leishmania major, CAC22697) (Ivens et al. 1998), with AICAR transformylase and IMP cyclohydrolase bifunctional enzymes (Gallus gallus, P31335) (Ni et al., 1991).
The clusters selected using the above strategy were further aligned to representative sequences of each enzyme of the pathway. The presence of a full-length EST sequence within a cluster is identified by a significant alignment in the translated N-terminal sequence of that EST, taking into consideration the natural heterogeneity of those regions.
This strategy allowed us to identify at least one potential case of polymorphism in sugarcane, the PRPP synthetase protein, an essential enzyme of the de novo purine synthesis pathway. This case is being further analyzed by the cloning and functional characterization of the sugarcane genes.
RESULTS
The search strategy adopted (Figure 2) allowed the estimation of a statistical support for the alignment of representative sequences to the sugarcane clusters using the Monte Carlo shuffling strategy. The initial analysis of several clusters identified as representatives for each member of the purine synthesis pathway indicated the degree of divergence within those enzymes. This result was compared to the statistical score of the Sugarcane cluster to each of the different representative enzyme sequences retrieved from the NCBI (National Center for Biotechnology Information) database. With such a strategy we were able to identify clusters representing each enzyme of the sugarcane purine synthesis pathway with a high degree of confidence. As shown in Table I, the analysis of the results obtained in each of the different libraries indicates that the purine synthesis pathway is present in every tissue of the plant. Fifty-three EST clones forming different clusters have been identified as unambiguously belonging to the purine synthesis pathway. In four libraries (normalized tissue, seeds, stem bark and leaves) we did not identify any EST sequence or cluster belonging to the purine synthesis pathway. However, these libraries represent the ones with the least number of ESTs clones sequenced in the project. Interestingly, the homologues of glutamine-PRPP-amidotransferase (GPA), AIR carboxylase (AIRcarb) and SAICAR synthetase were identified in only one library (Table I). The EST clone identified as having the highest frequency was the homologue to PRPP synthetase (PRS), this being consistent with this enzyme being central to several metabolic pathways.
The presence of EST clones within clusters representing full-length sequences is of great importance for the further investigation of the genes involved in this pathway. Such EST clones will allow the fast analysis of the entire sequence and its sub-cloning into appropriate vectors for functional investigation of the expressed protein. The alignment of the individual EST clone sequence and the reference sequences revealed the frequency of full length clones (Table II). From the 53 EST clones identified and distributed in the different clusters (Table I), 13 are full length ESTs and 40 represent partial length EST clones. The sequence for the bifunctional enzyme AICAR transformylase/IMP cyclohydrolase was identified in one instance as a full-length EST clone.
DISCUSSION
The investigation of central metabolic pathways is of great interest for the advancement of the knowledge of any organism. The purine de novo synthesis pathway (Figure 1) represents a central pathway affecting several metabolic routes of great importance and has been conserved phylogenetically in several taxa. However, some organisms (e.g. protozoan parasites) are purine auxotrophs and are dependent on the salvage pathway for providing the purine nucleotides necessary for intracellular metabolism. Differences in metabolic pathways or in the protein sequences that participate in such pathways can be exploited as potential targets for the development of inhibitory compounds, an approach which is being aggressively undertaken in various rational drug design programs.
We have applied a stringent search strategy (Figure 2) for the identification of homologue genes from the sugarcane clusters which participate in the purine synthesis pathway. The use of Monte Carlo shuffling analysis in our screening strategy was important to avoid the subjectivity normally involved in the evaluation of the alignment results of divergent sequences. Our results (Table I) show that all the enzymes necessary for the purine de novo synthesis pathway are present in sugarcane.
The identification of some clusters with a higher frequency of EST clones than others clusters (depending on the tissue or library analyzed) may be a result of the different number of sequences deposited in the SUCEST database for each library. At this point we can not conclude that these results reflect differential expression or mRNA stability for each gene. However, it is interesting to notice the high frequency of PRPP synthetase (PRS) ESTs identified because the product of PRS (PRPP) is a substrate for diverse metabolic pathways and Its abundance in the sugarcane libraries may reflect the importance of this enzymes. In several organisms, including mammals, isoforms of the PRS gene have been identified. In mammals at least three forms are known, PRS-I, PRS-II and PRS-III which act as a multienzyme complex. It is likely that in sugarcane the PRS gene is also present as a set of isoforms. We will further investigate such hypothesis.
The last enzyme in the metabolic cascade leading from Ribose-5-phosphate to IMP has also been identified in sugarcane as the bifunctional enzyme AICAR transformylase /IMP cyclohydrolase, but there is no evidence of any other multifunctional gene in this pathway in sugarcane. Four EST clones have been identified from the libraries of infected plants (Gluconacetobacter diazotroficans and Herbaspirillum rubrisubalbicans) as shown in Table I. These sequences have been identified as belonging to sugarcane genes and not the bacterial homologues, however only the full-length sequence will identify their origin with certainty.
The alignment of the identified sequence clusters with representatives of the purine metabolic pathway has allowed us to verify the presence of 13 full-length EST clones including one full-length bifunctional AICAR/ IMPcyc homologue. These findings do not directly reflect the expression of the sugarcane genes because they are not the result of direct mRNA quantification.
The present study identified all the genes of the purine de novo synthesis pathway in sugarcane. Several full-length EST clones are presently being investigated for their enzymatic characteristics. Such research may lead to the rational design and future development of anti-pathogenic agents and also provide insights into the evolutionary origin of this conserved metabolic pathway.
RESUMO
A via de síntese de purino nucleotídeos é considerada uma via de central importância para todas as células. Na maioria dos organismos, os purino nucleotídeos são sintetizados ''de novo'' a partir de precursores não-nucleotídicos como amino ácidos, amônia e dióxido de carbono. O conhecimento das enzimas envolvidas na via de síntese de purinas da cana-de-açúcar vai abrir a possibilidade do uso dessas enzimas como alvos no desenho racional de inibidores no combate a agentes fitopatogênicos, como esta sendo feita com diversos parasitos e células cancerosas. A seguinte estratégia esta sendo utilizada na identificação de genes de cana-de-açúcar para cada membro da via de síntese de purinas: Seqüências representativas dos genes que compões a via foram escolhidas do banco de dados NCBI. Essas seqüências de peptídeos estão sendo utilizadas em buscas ao banco de dados gerado pelo SUCEST pelo programa BLAST (implementação tBLASTn). Alinhamentos com os clusters de cana-de-açúcar são posteriormente analisados para sua significância estatística pela implementação PRSS3 do algoritmo conhecido como Monte Carlo shuffling. Para calibrar a análise dos resultados de PRSS3, foram empregadas seqüências conhecidas de diferentes taxas ao longo da arvore filogenética. Essas seqüências são comparadas duas a duas e com o cluster da cana-de-açúcar. A tabela de valores-p resultante indica o grau estatístico de similaridade e divergência entre as seqüências já descritas e entre essas e os clusters de cana-de-açúcar. Os resultados obtidos dessas análises estão descritos neste artigo.
ACKNOWLEDGMENTS
This work was supported by a research grant 99/02874-9 to O.H. Thiemann from the Fundação de Amparo a Pesquisa no Estado de São Paulo (FAPESP) as part of the ONSA network. We would like to thank the members of the Protein Crystallography and Structural Biology Group (IFSC-USP) for helpful discussions in the course of this work.
- Altschul, S.F., Gish, W., Miller, W., Myers, E.W. and Lipman, D.J. (1990). Basic local alignment search tool. J. Mol. Biol. 215: 403-410.
- Ivens, A.C., Lewis, S.M., Bagherzadeh, A., Zhang, L., Chan, H.M. and Smith, D.F. (1998). A physical map of the Leishmania major Friedlin genome. Genome Res. 8 (2): 135-145.
- Iwahana, H., Yamaoka,T., Mizutani, M., Mizusawa, N., Ii, S., Yoshimoto, K. and Itakura, M. (1993). Molecular cloning of rat amidophosphoribosyltransferase. J. Biol. Chem 268 (10): 7225-7237.
- Karlin S, Altschul SF. (1990). Methods for assessing the statistical significance of molecular sequence features by using general scoring schemes. Proc. Natl. Acad. Sci. U.S.A. 87 (6): 2264-8.
- Krath, B.N., Eriksen, T.A., Poulsen, T.S. and Hove-Jensen, B. (1999). Cloning and sequencing of cDNAs specifying a novel class of phosphoribosyl diphosphate synthase in Arabidopsis thaliana.Biochim. Biophys. Acta1430 (2): 403-408.
- Kubinyi, H. (1998). Structure-based drug design Curr. Opin. Drug. Disc. Dev. 1: 4-15.
- Madden, T.L., Tatusov, R.L. and Zhang, J. (1996). Applications of network BLAST server Meth. Enzymol 266: 131-141.
- Marr, J. J. and Ullman, B. (1995). Biochemistry and Molecular Biology of Parasites (Marr, J.J. and Muller, M., eds) 323-336.
- Marr, J. J. (1991). Purine analogs as chemotherapeutic agents in Leishmaniasis and American Trypanosomiasis J. Lab. Clin. Med. 118: 111-119.
- Ni, L., Guan, K., Zalkin, H. and Dixon, J.E. (1991). De novo purine nucleotide biosynthesis: cloning, sequencing and expression of a chicken PurH cDNA encoding 5-aminoimidazole-4-carboxamide-ribonucleotide transformylase-IMP cyclohydrolase. Gene 106 (2): 197-205.
- Pearson, W.R and Lipman, DJ. (1988). Improved tools for biological sequence comparison. Proc. Natl. Acad. Sci. U.S.A. 85 (8): 2444-8.
- Pearson, WR. (1990). Rapid and sensitive sequence comparison with FASTP and FASTA. Methods Enzymol 183: 63-98.
- Schnorr, K.M., Nygaard, P. and Laloue, M. (1994). Molecular characterization of Arabidopsis thaliana cDNAs encoding three purine biosynthetic enzymes. Plant J. 6 (1): 113-121.
- Senecoff, J.F. and Meagher, R.B. (1993). Isolating the Arabidopsis thaliana genes for de novo purine synthesis by suppression of Escherichia coli mutants. I. 5-Phosphoribosyl-5-aminoimidazole synthetase. Plant Physiol 102 (2): 387-399.
- Stryer, L. (1995). Biochemistry 4th edition: pp. 739-743.
- Ullman, B. and Carter, D. (1997). Molecular and biochemical studies on the hypoxanthine-guanine phosphoribosyltransferases of the pathogenic haemoflagellates. Int. J. Parasitol. 27: 203-213.
- Zhang, J. and Madden, T.L. (1997). PowerBLAST: A new network BLAST application for interactive or automated sequence analysis and annotation. Genome Res. 7: 649-656.
Publication Dates
-
Publication in this collection
27 June 2002 -
Date of issue
Dec 2001