Abstract
We established a specific genotyping assay for HLA-A*01, which is one of the most frequently found HLA-A alleles in the Caucasian population. This assay uses the polymerase chain reaction (PCR) with allele group-specific primers (ASP). HLA-A*01 group-specific primers were designed for exon 3 of the HLA-A gene, based on the recent HLA-sequence alignment. Both sense and anti-sense primers were designed with completely matched sequences to each specific HLA-A*01 allele, but mismatched by at least 1 nucleotide to all other known class I HLA alleles. By the use of these primers and stringent PCR conditions, we successfully genotyped the HLA-A*01 group alleles and achieved greater accuracy than previous methods.
HLA-A1; PCR; allele group-specific primers
HUMAN AND MEDICAL GENETICS
SHORT COMMUNICATION
Specific genotyping of human leukocyte antigen-A*01 by polymerase chain reaction using allele group-specific primers
Ikuma KasugaI, II; Peter D. ParéI; Andrew J. SandfordI
IUniversity of British Columbia, St Pauls Hospital, The James Hogg iCAPTURE Centre, Vancouver, BC, Canada
IIFirst Department of Internal Medicine, Tokyo Medical University, Tokyo, Japan
Send correspondence to Send correspondence to Ikuma Kasuga Tokyo Medical University, First Department of Internal Medicine 6-7-1 Nishi-shinjuku 160-0023 Shinjuku-ku, Tokyo, Japan E-mail: ikumakasuga@aol.com
ABSTRACT
We established a specific genotyping assay for HLA-A*01, which is one of the most frequently found HLA-A alleles in the Caucasian population. This assay uses the polymerase chain reaction (PCR) with allele group-specific primers (ASP). HLA-A*01 group-specific primers were designed for exon 3 of the HLA-A gene, based on the recent HLA-sequence alignment. Both sense and anti-sense primers were designed with completely matched sequences to each specific HLA-A*01 allele, but mismatched by at least 1 nucleotide to all other known class I HLA alleles. By the use of these primers and stringent PCR conditions, we successfully genotyped the HLA-A*01 group alleles and achieved greater accuracy than previous methods.
Key words: HLA-A1, PCR, allele group-specific primers.
The human leukocyte antigen (HLA) exhibits the highest degree of polymorphism in the human genome (Tokunaga et al., 1997; Prasad and Yang, 1996), and more than 950 HLA class I alleles have been reported to date (www.anthonynolan.org.uk/HIG/lists/class1list.html). Until recently, serological typing has been the primary technique used for HLA class I typing (Mittal et al., 1968). However, with the advent of PCR technologies, DNA-based HLA genotyping has developed (Sadler et al., 1994; Browning et al., 1993) during the last few years and has gradually replaced serological methods. Most commonly, DNA-based genotyping for the HLA-locus is performed by PCR with combinations of allele-specific primers (ASP) (Newton et al., 1989) designed from one of the hypervariable regions located in the second and third exons which encode the extracellular domains (Trowsdale et al., 1991). PCR-ASP methodology is based on the principle that completely matched oligonucleotide primers are more efficiently used in amplifying a target DNA sequence than mismatched oligonucleotide primers (Newton et al., 1989), and a mismatch at the 3 end of the primer to non-target sequence inhibits non-specific amplification. However, it is difficult to apply this technique for HLA group-specific genotyping, since HLA genes have a complex structure and have sequence homology among the different HLA class I loci, including the less polymorphic HLA-E, F, G genes and several pseudogenes (Trowsdale et al., 1991). Recently, the search for new class I alleles has advanced rapidly, and detection of new HLA alleles (Bunce et al., 2000; Guttridge and Darke, 2001) may render PCR-ASP techniques obsolete. For this reason, group-specific primers are often designed with a mismatch at different positions from the 3 end, and this reduces the amplification efficiency and specificity. In addition, many of the HLA class I group-specific primers are now designed to match the major alleles of each HLA group, but some less common alleles are mismatched to these primers. As a result, false-positive and false-negative amplification occur by the combination of these ASPs. Recently, HLA typing has contributed not only to the disease association studies, but also to the fields of legal medicine (Primorac et al., 1996) and anthropology (Ivanova et al., 2001; Arnaiz-Villena et al., 1999). Therefore, it is important to design primers which are completely matched to the target HLA loci, to provide accurate HLA group-specific genotyping.
In the present study, we established a new method of HLA-A*01 genotyping, which is one of the most frequent HLA-A allele groups in the Caucasian population. We designed the HLA-A*01 group-specific primer and PCR conditions based on the most up-to-date HLA-sequence alignment (http://www.ebi.ca.uk/imgt/hla/align.html). Both sense and antisense primers were designed in exon 3 of the HLA-A gene. The PCR product size is 101 bp. Although the sense primer is matched to all 11 HLA-A*01 alleles, this primer is also matched to some other HLA-A alleles (HLA-A*11 group, A*36 group, HLA-A*0244, HLA-A*24022 and HLA-A*6812) (Figure 1a). Other HLA-A alleles are mismatched by at least one nucleotide to the sense primer sequence, and HLA-B and C locus alleles are mismatched by at least two nucleotides. The antisense primer is only matched to all 11 HLA-A*01 alleles. Other HLA-A alleles are mismatched by at least one nucleotide at the 3 end or by two nucleotides in the centre of the primer, and HLA-B and Cw locus alleles are mismatched by at least 2 nucleotides (Figure 1b). With the exception of HLA-A*11 (2-nucleotide mismatch) and HLA-A*24022 (1 nucleotide mismatch at 3 end), these primers have at least 3 nucleotides mismatched to other HLA-A alleles. All HLA-B and HLA-Cw alleles have at least 4 mismatched nucleotides, and other class I HLA locus (HLA-E, F, G) alleles have more than 10 nucleotides mismatched to these primers. Finally, the sequences of these primers were completely matched to only the 11 HLA-A*01 alleles. In order to verify whether all DNA is successfully amplified or not, internal control primers which amplified a 394 bp fragment of the Coxsackie-adenovirus receptor gene (Bowles et al., 1999) (exon 2) were added in each reaction mixture. The sequence of the control sense primer is 5-CTGGGCAT CTCTTGAGTTTGGA-3, and the anti-sense primer is 5-ACTGGCAAGGTGATGGACACAT-3. Optimized PCR conditions for HLA-A*01 typing are as follows: the PCR reaction mixture in a final volume of 20 mL consisted of 100 ng genomic DNA, 1.5 mM MgCl2, 200 mM each of dATP, dCTP, dGTP and dTTP, 0.1 mM of each HLA-A*01 primer and 1.0 mM of each control primer, and 0.5 U DNA Taq Polymerase (Hotstar Taq®, Qiagen Inc, Mississauga, Ontario). PCR amplification was performed in a PCR EXPRESS Thermal Cycler (Thermo Hybrid, Ashford, Middlesex, UK). PCR conditions for cycling were optimized using the touchdown method (Hecker and Roux, 1996) as follows: initial denaturation step at 95 °C for 15 min, followed by 5 cycles of 94 °C for 30 s (denaturation), 70 °C for 30 s (annealing), and 72 °C for 45 s (extension), followed by 10 cycles with a decreased annealing temperature at 69 °C and 20 cycles at 68 °C, and a final extension step of 10 min at 72 °C.
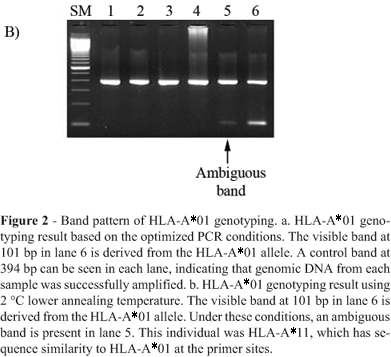
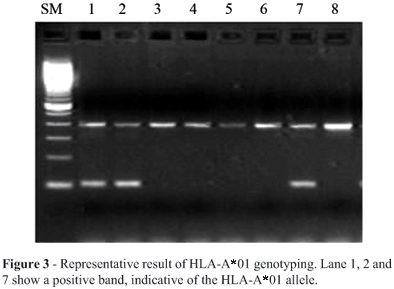
Figure 2 represents the amplified PCR fragment of HLA-A*01 and the internal control. The 101 bp band in lane 6 is specific for HLA-A*01 (Figure 2a and 2b). Changes in reagent and high DNA concentration or changes of annealing temperature may cause the ambiguous result shown in Figure 2b. In this case, a 2 °C lower annealing temperature causes false-positive bands such as in lane 5. This sample was actually HLA-A*11. We successfully genotyped the HLA-A*01 positive sample by using stringent PCR conditions and distinguished it from the other alleles even when the mismatch was not at the 3 end of the primers. After optimization of the PCR conditions, we performed HLA-A*01 genotyping using 200 Caucasian DNA samples. We could evaluate HLA-A*01 positive bands in this large number of samples without ambiguous amplification (Figure 3). Positive samples were selected for direct sequencing, which confirmed the genotypes as HLA-A*01. We calculated the allele frequency of HLA-A*01 in the 200 genotyped samples by assuming Hardy-Weinberg equilibrium. The estimated allele frequency of HLA-A*01 was 0.15, which was well matched to the reported Caucasian HLA-A*01 allele frequency. Thus, we consider that by the use of these primers and stringent PCR conditions, we successfully genotyped the HLA-A*01 group alleles and distinguished them from 952 HLA class I alleles.
PCR-ASP is one of the common methods for group-specific HLA typing, but it sometimes leads to typing errors (Schaffer and Olerup, 2001). In fact, we previously used the published group-specific primers for class I HLA genotyping, and occasionally found mistyping by direct sequencing. One of the reasons for mistyping is sub-optimal design of ASPs. For example, in the case of HLA-A*01 group-specific amplification, the primer sequences of HLA-A*01 are matched to the high-frequency alleles such as HLA-A*010101 and HLA-A*0102, whereas these are not completely complementary to HLA-A*0103, HLA-A*0106 or HLA-A0107. Moreover, other HLA-A group alleles (e.g., HLA-A*3603, HLA-A*8001) are matched to this sequence and amplified by these primers. In this study, however, we also found that minor changes of primer and enzyme concentration, or annealing temperature, could cause the typing error shown in Figure 2b even with the use of highly specific primers. Therefore, we emphasize that strictly controlled PCR conditions as well as the use of well-designed primers are required to achieve accurate HLA group-specific genotyping. On the other hand, HLA-A*01 is one of the most frequent serological types in the Caucasoid population, with an allele frequency of 0.13-0.18. From these data, it results that almost 25-34% of Caucasian people have HLA-A*01. Thus, precise HLA-A*01 typing would provide important information for disease and population studies.
In summary, we established a new and accurate genotyping method for HLA-A*01 by using PCR-ASP. The primers successfully amplified all 11 HLA-A*01 alleles and distinguished them from all other HLA class I alleles. Primer design and allele identification were according to the newly-aligned HLA sequence database. Thoroughly optimized, appropriate PCR conditions were also used to prevent both false-positive and false-negative results. Thus, we overcame the drawback of previous PCR-ASP methods for HLA typing as mentioned above, and provided a reliable HLA-A*01 genotyping method.
Received: December 14, 2004; Accepted: October 24, 2005.
Associate Editor: Emmanuel Dias
- Arnaiz-Villena A, Iliakis P, Gonzalez-Hevilla M, Longas J, Gomez-Casado E, Sfyridaki K, Trapaga J, Silvera-Redondo C, Matsouka C and Martinez-Laso J (1999) The origin of Cretan populations as determined by characterization of HLA alleles. Tissue Antigens 53:213-26.
- Bowles KR, Gibson J, Wu J, Shaffer LG, Towbin JA and Bowles NE (1999) Genomic organization and chromosomal localization of the human Coxsackievirus B-adenovirus receptor gene. Hum Genet 105:354-359.
- Browning MJ, Krausa P, Rowan A, Bicknell DC, Bodmer JG and Bodmer WF (1993) Tissue typing the HLA-A locus from genomic DNA by sequence-specific PCR: Comparison of HLA genotype and surface expression on colorectal tumor cell lines. Proc Natl Acad Sci USA 90:2842-2845.
- Bunce M, Procter J, Dunn PP, Day S, Ross J and Welsh KI (2000) Identification of the null HLA-A2 allele, A*0232N. Tissue Antigens 55:31-36.
- Guttridge MG and Darke C (2001) Identification and nucleotide sequence of HLA-A*03013. Tissue Antigens 57:546-547.
- Hecker KH and Roux KH (1996) High and low annealing temperatures increase both specificity and yield in touchdown and stepdown PCR. Biotechniques 20:478-485.
- Ivanova M, Spassova P, Michailova A and Naumova E (2001) Distributions of HLA class I alleles and haplotypes in Bulgarians - Contribution to understanding the origin of the population. Tissue Antigens 57:208-215.
- Mittal KK, Mickey MR, Singal DP and Terasaki PI (1968) Serotyping for homotransplantation. 18. Refinement of microdroplet lymphocyte cytotoxicity test. Transplantation 6:913-927.
- Newton CR, Graham A, Heptinstall LE, Powell SJ, Summers C, Kalsheker N, Smith JC and Markham AF (1989) Analysis of any point mutation in DNA. The amplification refractory mutation system (ARMS). Nucleic Acids Res 17:2503-2516.
- Prasad VK and Yang SY (1996) Allele assignment for HLA-A, -B, and -C genes to the Tenth International Histocompatibility Workshop cell lines. Tissue Antigens 47:538-546.
- Primorac D, Andelinovic S, Defins-Gojanovic M, Drmic I, Rezic B, Baden MM, Kennedy MA, Schanfield MS, Skadel SB and Lee HC (1996) Identification of war victims from mass graves in Croatia, Bosnia, and Herzegovina by use of standard forensic methods and DNA typing. J Forensic Sci 41:891-894.
- Sadler AM, Petronzelli F, Krausa P, Marsh SG, Guttridge MG, Browning MJ and Bodmer JG (1994) Low-resolution DNA typing for HLA-B using sequence-specific primers in allele- or group-specific ARMS/PCR. Tissue Antigens 44:148-154.
- Schaffer M and Olerup O (2001) HLA-AB typing by polymerase-chain reaction with sequence-specific primers: More accurate, less errors, and increased resolution compared to serological typing. Tissue Antigens 58:299-307.
- Tokunaga K, Ishikawa Y, Ogawa A, Wang H, Mitsunaga S, Moriyama S, Lin L, Bannai M, Watanabe Y, Kashiwase K, Tanaka H, Akaza T, Tadokoro K and Juji T (1997) Sequence-based association analysis of HLA class I and II alleles in Japanese supports conservation of common haplotypes. Immunogenetics 46:199-205.
- Trowsdale J, Ragoussis J and Campbell RD (1991) Campbell, Map of the human MHC. Immunol Today 12:443-446.
Send correspondence to
Publication Dates
-
Publication in this collection
12 June 2006 -
Date of issue
2006
History
-
Accepted
24 Oct 2005 -
Received
14 Dec 2004