Abstracts
The objective of the present trial was to characterize genetically strains of Campylobacter jejuni subsp. jejuni isolated from humans and several animal sources (bovines, swine, dogs, primates, wild boars and poultry). A total of 828 different animal samples (feces, carcass, aborted fetus and hysterectomized uterus) were analysed by means of routine bacteriological methods, and 36 C. jejuni strains were isolated. Thirty strains of human fecal origin were obtained in clinical analysis laboratories in the city of São Paulo. The 66 C. jejuni strains isolated were submitted to genetic characterization. Primers based on fla A gene were used in a polymerase chain reaction (PCR) and amplified a fragment of the 702 bp. PCR products were evaluated by means of sequencing and genealogic analysis. Genetic variability analysis of 66 strains showed 44 different subtypes of C. jejuni. One subtype was identical to a C. jejuni strain of human origin with the sequence in the GenBank (GENBANK accession number AF050186). Subtyping analysis of C. jejuni strains based on sequencing of the fla A gene variable region and analysis of sequence alignment by the Maximum Parsimony method showed to be highly discriminatory, providing the best conditions to differentiate strains involved in outbreaks from those sporadically isolated. This is the first study of molecular subtyping analysis of human and animal C. jejuni strains using sequencing technique and genealogic analysis in the state of São Paulo, Brazil.
Campylobacter jejuni; subtyping; sequencing, flagellin
O objetivo do presente trabalho foi caracterizar geneticamente estirpes de Campylobacter jejuni subsp. jejuni isoladas de humanos e de diferentes origens animais (bovinas, suínas, cães, primatas, javalis, suínos e aves de corte). Um total de 828 amostras (fezes, carcaças, fetos abortados e útero histerectomizado) foram analisadas por métodos de rotina bacteriológica e 36 estirpes de C. jejuni foram isoladas. Trinta estirpes de origem fecal humana foram obtidas de laboratórios de análises clínicas da cidade de São Paulo. As 66 estirpes de C. jejuni isoladas foram submetidas à caracterização genética. Oligonucleotídeos baseados no gene fla A foram usados na reação de polimerase em cadeia (PCR) e amplificou um fragmento de 702 pb. Os produtos obtidos pela PCR foram avaliados pelas técnicas de seqüenciamento e análise genealógica. Análise da variabilidade genética das 66 estirpes revelou 44 diferentes subtipos de C. jejuni. Um subtipo de origem humana apresentou seqüência idêntica à de C. jejuni depositada no GenBank (GENBANK acesso número AF050186). A subtipagem das estirpes de C. jejuni baseadas no seqüenciamento da região variável do gene fla A e na análise do alinhamento das seqüências pelo método da Máxima Parcimônia, mostraram-se altamente discriminatórios fornecendo melhores condições para a correta diferenciação entre estirpes originárias de surto e as isoladas esporadicamente. Este foi o primeiro estudo de subtipagem molecular de estirpes de C. jejuni de origem humana e animal utilizando a técnica do seqüenciamento com análise genealógica realizado no Estado de São Paulo, Brasil.
Campylobacter jejuni; subtipagem; seqüenciamento; flagelina
VETERINARY MICROBIOLOGY
Molecular subtyping of Campylobacter jejuni subsp. jejuni strains isolated from different animal species in the state of São Paulo, Brazil
Subtipagem molecular de estirpes de Campylobacter jejuni subsp. jejuni isoladas de diferentes espécies animais do Estado de São Paulo, Brasil
Eliana ScarcelliI* * Corresponding Author. Mailing address: Centro de Pesquisa e Desenvolvimento de Sanidade Animal, Instituto Biológico, Av. Conselheiro Rodrigues Alves, 1252. 04014-002, São Paulo, SP, Brasil. Tel.: (+5511) 5087-1770. E-mail: pinheiro@biologico.sp.gov.br ; Rosa Maria PiattiI; Ricardo HarakavaII; Simone MiyashiroI; Flora Maria de Campos FernandesIV; Fabíola Ribeiro CamposI; Waldemar FranciscoV; Margareth Élide GenovezI; Leonardo José RichtzenhainIII
ICentro de Pesquisa e Desenvolvimento de Sanidade Animal, Instituto Biológico, São Paulo, SP, Brasil
IICentro de Pesquisa e Desenvolvimento de Sanidade Vegetal, Instituto Biológico, São Paulo, SP, Brasil
IIIDepartamento de Medicina Veterinária e Preventiva e Saúde Animal, Faculdade de Medicina Veterinária e Zootecnia, Universidade de São Paulo, São Paulo, SP, Brasil
IVMuseu de Zoologia, Universidade de São Paulo, São Paulo, SP, Brasil
VInstituto de Ciências Biomédicas, Universidade de São Paulo, São Paulo, SP, Brasil
ABSTRACT
The objective of the present trial was to characterize genetically strains of Campylobacter jejuni subsp. jejuni isolated from humans and several animal sources (bovines, swine, dogs, primates, wild boars and poultry). A total of 828 different animal samples (feces, carcass, aborted fetus and hysterectomized uterus) were analysed by means of routine bacteriological methods, and 36 C. jejuni strains were isolated. Thirty strains of human fecal origin were obtained in clinical analysis laboratories in the city of São Paulo. The 66 C. jejuni strains isolated were submitted to genetic characterization. Primers based on fla A gene were used in a polymerase chain reaction (PCR) and amplified a fragment of the 702 bp. PCR products were evaluated by means of sequencing and genealogic analysis. Genetic variability analysis of 66 strains showed 44 different subtypes of C. jejuni. One subtype was identical to a C. jejuni strain of human origin with the sequence in the GenBank (GENBANK accession number AF050186). Subtyping analysis of C. jejuni strains based on sequencing of the fla A gene variable region and analysis of sequence alignment by the Maximum Parsimony method showed to be highly discriminatory, providing the best conditions to differentiate strains involved in outbreaks from those sporadically isolated. This is the first study of molecular subtyping analysis of human and animal C. jejuni strains using sequencing technique and genealogic analysis in the state of São Paulo, Brazil.
Key words:Campylobacter jejuni, subtyping, sequencing, flagellin
RESUMO
O objetivo do presente trabalho foi caracterizar geneticamente estirpes de Campylobacter jejuni subsp. jejuni isoladas de humanos e de diferentes origens animais (bovinas, suínas, cães, primatas, javalis, suínos e aves de corte). Um total de 828 amostras (fezes, carcaças, fetos abortados e útero histerectomizado) foram analisadas por métodos de rotina bacteriológica e 36 estirpes de C. jejuni foram isoladas. Trinta estirpes de origem fecal humana foram obtidas de laboratórios de análises clínicas da cidade de São Paulo. As 66 estirpes de C. jejuni isoladas foram submetidas à caracterização genética. Oligonucleotídeos baseados no gene fla A foram usados na reação de polimerase em cadeia (PCR) e amplificou um fragmento de 702 pb. Os produtos obtidos pela PCR foram avaliados pelas técnicas de seqüenciamento e análise genealógica. Análise da variabilidade genética das 66 estirpes revelou 44 diferentes subtipos de C. jejuni. Um subtipo de origem humana apresentou seqüência idêntica à de C. jejuni depositada no GenBank (GENBANK acesso número AF050186). A subtipagem das estirpes de C. jejuni baseadas no seqüenciamento da região variável do gene fla A e na análise do alinhamento das seqüências pelo método da Máxima Parcimônia, mostraram-se altamente discriminatórios fornecendo melhores condições para a correta diferenciação entre estirpes originárias de surto e as isoladas esporadicamente. Este foi o primeiro estudo de subtipagem molecular de estirpes de C. jejuni de origem humana e animal utilizando a técnica do seqüenciamento com análise genealógica realizado no Estado de São Paulo, Brasil.
Palavras-chave:Campylobacter jejuni, subtipagem, seqüenciamento, flagelina
INTRODUCTION
Bacteria in the Campylobacter genus are widely distributed and can be isolated from both domestic and wild animal species (1,3,20).
Campylobacter jejuni is the most common cause of diarrhea in children in developing countries, and it is the primary cause of enteritis in industrialized regions (4). Adittionally, the infection caused by C. jejuni may lead to a form of neuromuscular paralysis called Guillain-Barré syndrome (6).
In the United States, it is estimated that more than 2.5 milion cases of enteritis a year are caused by C. jejuni, a rate that is far over the number of salmonellosis and shigellosis cases (7).
In Brazil, the presence of Campylobacter spp. has been reported in both asymptomatic individuals and cases of chronic or acute diarrhea. Its incidence in cases of diarrhea in the state of São Paulo is around 25.9% (19).
Genotyping may enable adequate differentiation between C. jejuni strains, and the technique has been successfully used in phylogenetic and epidemiological studies (7,11).
The ability of PCR to amplify specific DNA regions has been used to identify Campylobacter strains. Sequencing of specific genes has shown to be highly efficient in the study of genetic variability of C. jejuni, specially gene fla A, which encodes the protein flagellin, the main monomeric subunit of the flagellum (7,9,14,15,21).
The objective of the present trial was the genotyping of Campylobacter jejuni subsp. jejuni strains isolated from humans and different animal species (bovines, swine, dogs, primates, wild boar and poultry) in the state of São Paulo, both by means of sequencing gene fla A variable region, and of genealogic analysis.
MATERIALS AND METHODS
Samples origin
A total of 828 different animal samples were sent to the Laboratório de Doenças Bacterianas da Reprodução at Instituto Biológico of São Paulo, and analysed by means of routine bacteriological methods and 36 C. jejuni strains were isolated (bovine, n=1/330; swine, n=1/24; dogs, n=4/26; non-human primates, n=18/366; wild boar, n=1/27 and poultry, n=11/55). Samples came from different cities of São Paulo state. Thirty strains of human fecal origin were obtained in clinical analysis laboratories in city of São Paulo. The patients' ages ranged from 1 to 77 years with a 16 year-old average, and analysed in the present study (Table 1).
DNA preparation and PCR of the fla A gene
C. jejuni strains were grown at 37ºC for 48 h under micraerophilic atmosphere (5% O2, 10% CO2, 85% N2) on blood Brucella Agar (Difco). DNA was extracted by boiling, according to Nishimura et al. (16) and On et al. (17). Colonies obtained from 2 to 3 day-old cultures were used to prepare C. jejuni suspensions in 1.0 mL of ultrapure water (Milli-Q, Millipore Inc.), corresponding to a reading of 8 on the McFarland turbidity scale (2.3 x 109 bact/mL). These suspensions were heated at 100ºC for 10 min. Five microliters (0.1 mg DNA/mL) of suspension supernatant were used as templates. The oligonucleotides used as PCR primers were forward fla A primer 5'-TA CTA CAG GAG TTC AAG CTT-3' and reverse fla A primer 5'-GT TGA TGT AAC TTG ATT TTG-3' that represented the variable (V1) region, according to Nishimura et al. (16). PCR was performed with 1 x PCR buffer (Gibco- BRL), 200 mM dNTPs, 2.5 mM MgCl2, 40 pmol of each primer, and 2.5 u Taq DNA polymerase (Gibco- BRL), and 5 ml of template DNA. A 30-cycles reaction was run in a PTC 200 thermocycler (MJ Research) with 48 sec denaturing at 94ºC, 36 sec annealing at 55ºC, 2 min extension at 72ºC and 10 min final extension at 72ºC. Resulting product was approximately 702 bp. Analysis of the product amplified from gene fla A was subjected in 2.0% agarose gel electrophoresis in 0.5 X TBE buffer (0.045M TRIS-Borate and 1 mM EDTA, pH 8.0). Gel was submitted to constant voltage equal to 5-6 V/cm, using as standard marker a 100bp ladder (Gibco- BRL, Gaithersburg, USA). Gel was stained by ethidium bromide 0.5 mg/mL, for 15 min and photographed under UV light (300-320 nm) using a Kodak Digital Camera DC/120 Zoom. Images were analyzed by 1D Image Analysis software (Kodak Digital Science).
Sequencing
PCR products were purified using Concert Rapid PCR Purification System (Gibco-Brl). Both DNA strands were sequenced using fla A primers and Big Dye Terminator 2.0 kit (Applied Biosystems). Sequencing reactions were conducted in a PTC 200 thermocycler (MJ Research), using 25 cycles at 96ºC for 10 sec, 50ºC for 5 sec and 60ºC for 4 min. Sequencing reactions were analysed in an automatic sequencer ABI 377 (Applied Biosystems).
Alignment and translation of the nucleotide sequences
Sequences were aligned with homologous C. jejuni sequences available in the GenBank (Accession number AF050186). Alignment was performed manually with the aid of SeqPup v. 0.6f (8) and Sequence Navigator v. 1.0.1 software. Translation was performed automatically with the aid of SeqPup v. 0.6f software, considering the universal genetic code (8).
Genealogical analysis
Aligned sequences were compared and dendrograms were generated by means of the Phylogenetic Analysis Using Parsimony (PAUP*) software v. 4.0 (22), using criterion-based heuristics with stepwise addition algorithm. The computation of genetic distances was performed using the same software, based on model TN93 (23). Bootstrap values were also obtained with PAUP*, and consistency and retention indexes were obtained with MacClade v. 3.0.3 software (13).
RESULTS
The sixty-six C. jejuni strains analysed in the present study were grouped in 44 different subtypes (Table 2).
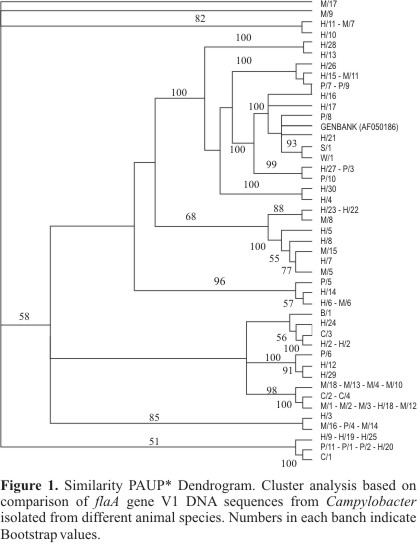
The Fig. 1 presents the dendrogram that was generated based on the 44 subtypes of the variable region V1 in flagellin A gene (fla A), obtained by PAUP* software (22).
The alignment of the 66 sequences of the gene fla A variable region V1, presented identity of several strains. It is showed in Table 2, where the 66 sequences were grouped in 44 subtypes.
One subtype (14) was identical to a C. jejuni strain of human origin with the sequence in the GenBank (GENBANK accession number AF050186).
Subtypes 15 through 44 corresponded to unique sequences of C. jejuni strains from different animal species.
DISCUSSION
The Centers for Disease Control and Prevention estimate that Campylobacter enteritis is a multibillion-dollar disease and that the consumption of poultry meet is a primary source of human infection (10).
Human infections caused by Campylobacter are far more common than those caused by Salmonella spp (5). However, few outbreaks could be traced to a particular foodstuff (5). The frequent contamination of foods with different strains of Campylobacter spp has made it difficult to determine a route of infection and the recognition even well characterized outbreaks (5,12).
Reproducible and discriminatory typing methods are increasingly necessary to identify and trace the routes of transmission of Campylobacter spp strains in the food chain (5). Due to the fact that the epidemiology of Campylobacter spp infections is not fully understood, new laboratory techniques are needed to differentiate isolates involved or not in outbreaks (7).
Serotyping is the best phenotypical method for the epidemiological typing of Campylobacter spp and has been available for many years. Although it is still useful in the investigation of clinical cases and outbreaks, results may be ambiguous, probably due to the occurrence of non-typable strains or cross-reactions between certain serotypes (18). Moreover, it is an expensive and laborious technique, demands a large number of standard strains and constant production of anti-serum. It also requires at least 5 to 7 days to be completed and repetitive subculture of the isolates before serological tests involving more than 74 different serotypes of C. jejuni may be performed (7,24).
Table 2 shows that the 66 C. jejuni sequences that came from state of São Paulo were grouped in 44 subtypes identical by means of alignment and translation of nucleotide sequences.
The results show that one animal species can be shared in the same subtype (subtypes 3, 4, 5, 8, 12 and 13). Table 2 also shows that 6 subtypes were common both to human and animal strains. Subtypes 7 and 9, are shared by human and poultry strains, and subtypes 1, 6, 10 and 11, by human and primates, suggesting that these animal species might be a possible source of infection for humans. Subtype 2 presented strains common to primates and to poultry. It should be emphasized that similar to what occurs with humans, poultry were part of the diet of primates kept in captivity. Sequence homology observed between strain isolated from a clinical case and a strain isolated from the probable infection source may be strong evidence of the participation of this source of infection in the enteritis case.
Subtypes 15 to 44 corresponded to individual sequences of the other 30 samples of C. jejuni analyzed. They corresponded to most of the isolates studied, involving 16 of human origin and 13 of animal origin. This extreme accuracy in the individualization of strains in unique subtypes may be greatly useful in the identification of strains or sources of infection related to these strains in the state of São Paulo.
Although a few subtypes were found exclusively in one animal specie, the limited number of samples does not allow to infer a host specialitation. In fact, subtypes grouping through phylogenetic analyses suggests the opposite.
Genotypic methods have usually greater discriminatory power than phenotypical methods, such as biotyping and phagotyping. However, the combination of a genotypic method with serotyping, may not lead only to a greater discriminating power, but also to a complete identification of the strain (24).
Subtyping of C. jejuni strains based on the sequencing of fla A gene variable region (V1) and on analysis of sequence alignment by means of the Maximum Parsimony method produced a dendrogram that was highly discriminatory, as it was observed by Meinersmann et al. (14) and Fitzgerald et al. (5). This method may be a useful option on the differentiation between strains related to outbreaks and those that are sporadically isolated.
This report describes the first study of molecular subtyping of C. jejuni strains obtained from human and animal samples by means of sequencing and genealogic analysis in the state of São Paulo. In Brazil, few studies on the epidemiology of human or animal campylobacteriosis using molecular techniques have been published. It is urgent the stablishment of this kind of methodology in order to support the investigation of foodborne outbreaks caused by Campylobacter.
Submitted: April 25, 2005; Returned to authors for corrections: August 09, 2005; Approved: October 20, 2005
- 1. Altekruse, S.F. Campylobacter jejuni in foods. J. Am. Vet. Med. Assoc. Rev., 213, 1734-1735, 1998.
- 2. Blaser, M.J. Campylobacter species In: Mandell, L.G.; Douglas, R.G.; Bennett, J.E., (eds). Principles and pratice of infections diseases New York, Churchill Livingstone, 1990, p. 1649-1658.
- 3. Carter, A.M.; Pacha, R.E.; Clarck, G.W.; Willians, E.A. Seasonal occurrence of Campylobacter spp. in surface waters and their correlation with standard indicator bacteria. Appl. Environ. Microbiol., 53, 523-526, 1987.
- 4. Carvalho, A.C.T.; Ruiz-Palacios, G.M.; Ramos-Cervantes, P.; Cervantes, L.; Jing, X.; Picckering, L.K. Molecular characterization of invasive and noninvasive Campylobacter jejuni and Campylobacter coli isolates. J. Clin. Microbiol., 39, 1353-1359, 2001.
- 5. Desai, I.M.; Logan, J.M.J.; Frost, J.A.; Stanley, J. Genome sequence-based fluorescent length polymorphism of Campylobacter jejuni, its relanonship to serotyping, and its implications for epidemiological analysis. J. Clin. Microbiol., 39, 3823-3829, 2001.
- 6. Duim, B.; Win Ang, C.; Van Bellkum, A.; Rigter, A.; Van Leeuwen, N.W.J.; Endtz, H.P.; Wagenaar, J.A. Amplified fragment length polymorphism analysis of Campylobacter jejuni strains isolated from chickens and patients with gastroenteritis or Guillain-Barré or Miller Fisher syndrome. Appl. Environ. Microbiol, 66, 3917-3923, 2000.
- 7. Fitzgerald, C.; Helsel, L.O.; Nicholson, M.A.; Olsen, S.J.; Swerdlow, D.L.; Flahart, R.; Sexton, J.; Fields, P.I. Evaluation of methods for subtyping Campylobacter jejuni during an outbreak involving a food handler. J. Clin. Microbiol., 39, 2386-2390, 2001.
- 8. Gilbert, D.G. SequPup. A biosequence editor and analysis application. v. 0.6f, 1995.
- 9. Harrington, C.S.; Thomson-Carter, F.M.; Carter, P.E. Evidence for recombination in the flagellin locus of Campylobacter jejuni: implications for the flagellin gene typing scheme. J. Clin. Microbiol., 35, 2386-2392, 1997.
- 10. Hiett, K.L.; Stern, N.J.; Fedorka-Cray, P.; Cox, N.A.; Musgrove, M.T.; Ladely, S. Molecular Subtype Analyses of Campylobacter spp. from Arkansas and California Poultry Operations. Appl. Environ. Microbiol, 68, 6220-6236, 2002.
- 11. Jimenez, A.; Barros-Velazquez, J.; Rodrigues, J.; Villa, T.G.; Restriction endonuclease analysis, DNA relatedness and phenotypic characterization of Campylobacter jejuni and Campylobacter coli isolates involved in food-borne disease. J. Appl. Microbiol, 82, 713-721, 1997.
- 12. Kramer, J.M.; Frost, J.A.; Bolton, F.J.; Wareing, D.R. Campylobacter contamination of raw meat and poultry at retail sale: identification of multiple types and comparison with isolates from human infection. J. Food. Prot, 63, 1654-1659, 2000.
- 13. Maddison, W.P.; Maddison, D.R. MacClade Massachusetts: Sinauer, v. 3.0.3., 1992.
- 14. Meinersmann, R.J.; Helsel, L.O.; Fields, P.I.; Hiett, K.L. Discrimination of Campylobacter jejuni isolates by fla gene sequencing. J. Clin. Microbiol., 35, 2810-2814, 1997.
- 15. Nielsen, E.M.; Engberg, J.; Fussing, V.; Petersen, L.; Brogren, C.H.; On, S.L. W. Evaluation of phenotypic and genotypic methods for subtyping Campylobacter jejuni isolates from humans, poultry, and cattle. J. Clin. Microbiol., 38, 3800-3810, 2000.
- 16. Nishimura, M.; Nukina, M.; Yuan, J.M.; Shen, B.Q.; M.A.J.J.; Ohta, M.; Saida, T.; Uchiyama, T. PCR-based restriction fragment lengt polymorphism (RFLP) analysis and serotyping of Campylobacter jejuni isolates from diarrheic patients in China and Japan. FEMS Microbiol. Lett., 142, 133-138, 1996.
- 17. On, S.; Harrington, C.S. Evaluation of numerical analysis of pfge-dna profiles for differentiating Campylobacter fetus subspecies by comparison with phenotypic, PCR and 16S rDNA sequencing methods. J. Appl. Microbiol., 90, 285-293, 2001.
- 18. Patton, C.M.; Barret, T.J.; Morris, G.K. Comparison of the Penner and Lior methods for serotyping Campylobacter spp. J. Clin. Microbiol., 22, 558-565, 1985.
- 19. Pinheiro, E.S. Caracterização genética de estirpes de Campylobacter jejuni subsp. jejuni isoladas de diferentes espécies animais. São Paulo, 2003, 69p. (Ph. D. Thesis. Instituto de Ciências Biomédicas. USP).
- 20. Scarcelli, E.; Genovez, M.E.; Cardoso, M.V.; Souza, M.C.A.M.; Grasso, L.M.P.S.; Souza, C.A.I.; Torres, A.P. Avaliação do potencial de disseminação de Campylobacter spp por diferentes espécies animais. Arq. Inst. Biol, São Paulo, 65, 55-61, 1998.
- 21. Shi, F.; Chen, Y.Y.; Wassenaar, T.M.; Woods, H.W.; Coloe, P.J.; Fry, B.N. Development and application of a new scheme for typing Campylobacter jejuni and Campylobacter coli by PCR-based restriction fragment length polymorphism analysis. J. Clin. Microbiol., 40, 1791-1797, 2002.
- 22. Swofford, D.L. Paup*, Phylogenetic Analysis Using Parsimony (*and Other Methods). Sunderland, Massachusetts, v. 40, 2000.
- 23. Tamura K.; Nei, M. Estimation of the number of nucleotide substitutions. Mol. Biol. Evol., 11, 154-157, 1993.
- 24. Wassenaar, T.M., Newell, D. Genotyping of Campylobacter spp. Appl. Environ. Microbiol., 66, 1-9, 2000.
Publication Dates
-
Publication in this collection
10 May 2006 -
Date of issue
Dec 2005
History
-
Accepted
20 Oct 2005 -
Reviewed
09 Aug 2005 -
Received
25 Apr 2005