Abstract
Effects of various factors, such as pH, ionic strength, glutaraldehyde concentration, enzyme amount and immobilization time, on enzyme activity were investigated. The immobilization conditions were optimized by orthogonal experiments. Characterizations of immobilized nuclease p1 were also evaluated. Through orthogonal optimization, the optimal immobilization conditions were as follows: pH 5.6, ionic strength 0.125, glutaraldehyde concentration 0.20% and immobilization time 2.0 h. Optimal pH of immobilized enzyme was 5.8. Optimal temperature of immobilized enzyme was 70ºC. Thermal, operational and storage stabilities of the enzyme were improved after it was immobilized on DEAE cellulose. Michaelis constant Km of immobilized enzyme at 69ºC was found to be 27.21 g/l by the Lineweaver-Burk plot.
DEAE cellulose; Nuclease p1; Immobilization; Characterization
BIOPROCESS ENGINEERING
Nuclease p1 immobilized on deae cellulose
Lu-E ShiI; Yu YiII; Zhen-Xing TangII; Wen-Yue XiongII; Jiang-Feng MeiII; Guo-Qing YingII, * * To whom correspondence should be addressed
ICollege of Life and Environmental Sciences, Hangzhou Normal University, Hangzhou, 310036, Zhejiang, China
IICollege of Pharmaceutical Science, Zhejiang University of Technology, Hangzhou, 310032, Zhejiang, China. E-mail: gqying@zjut.edu.cn, E-mail: shilue@126.com
ABSTRACT
Effects of various factors, such as pH, ionic strength, glutaraldehyde concentration, enzyme amount and immobilization time, on enzyme activity were investigated. The immobilization conditions were optimized by orthogonal experiments. Characterizations of immobilized nuclease p1 were also evaluated. Through orthogonal optimization, the optimal immobilization conditions were as follows: pH 5.6, ionic strength 0.125, glutaraldehyde concentration 0.20% and immobilization time 2.0 h. Optimal pH of immobilized enzyme was 5.8. Optimal temperature of immobilized enzyme was 70ºC. Thermal, operational and storage stabilities of the enzyme were improved after it was immobilized on DEAE cellulose. Michaelis constant Km of immobilized enzyme at 69ºC was found to be 27.21 g/l by the Lineweaver-Burk plot.
Keywords: DEAE cellulose; Nuclease p1; Immobilization; Characterization.
INTRODUCTION
Nuclease p1 (EC 3.1.30.1), an extra-cellular enzyme, was first identified by Kuninaka et al. from Penicillium citrinum (Kuninaka et al, 1961). It can cleave 5'-nucleotides (e.g., 5'-AMP, -GMP, -CMP and -UMP) successively from the 3'-hydroxy termini of 3'-hydroxy-terminated oligonucleotides derived from RNA (Steensma et al., 2004; Kuninaka, 1996). When nuclease p1 is added during the production of bakers' yeast, the enzyme hydrolyses the yeast RNA efficiently into 5'-GMP. An interesting feature of 5'-GMP is that it acts synergistically with monosodium glutamate and thus can largely replace monosodium glutamate in various food products. 5'-Nucleotides have been widely used in food processing and the pharmaceutical industry (Gerald and Tilak, 1991). They can be used to synthesize antivirus and anticancer drugs like an acridine (Ying et al., 2004). Nucleotide derivatives have been applied in the treatment of illnesses of the human central nervous system and circulatory system.
Immobilization of enzymes on insoluble supports is a significant process due to its promising potential for improving enzyme thermal or pH stability, simplifying product purification, and facilitating enzyme recycling (Blanco et al., 2004; Caramori et al.,2004; Fernandes et al.,2004; Cheng et al., 2006; Goradia et al., 2005; Lai and Lin, 2005; Panya et al., 2005). Therefore, immobilized enzymes have been widely applied in the fields of biocatalysis and biosensors (Dai et al., 2005; Luo et al., 2004; Xu et al., 2006; Xu et al, 2004; Zhao et al., 2007),
To the best of our knowledge, studies of nuclease p1 immobilized on DEAE cellulose rarely have been reported. In this paper, nuclease p1 was immobilized on DEAE cellulose. Preparation of immobilized enzyme and its characterizations were studied systematically. All the results obtained in this paper would provide a sound basis for further exploration.
MATERIALS AND METHODS
Materials
DEAE cellulose was purchased from Wilenman (USA). Groundnut meal was a gift from Hangzhou Zhongmeihuadong Pharmaceutical Co., Ltd. (Hangzhou, China). Glucose kit was purchased from Ningbo Cicheng Reagent Factory (Ningbo, China). Nuclease p1 -producing strain of Penicillium citrinum, used throughout the study, was kindly provided by Hangzhou Meiya Biotechnical Co. Ltd. (Hangzhou, China). All other reagents were of analytical grade.
Production of Nuclease p1
For nuclease p1 production, Penicillium citrinum was grown at 30ºC in a medium (pH 7.0) consisting of (w/v): sucrose 3.0%, potassium dihydrogen phosphate 0.10%, ferrous sulfate 0.010%, sodium nitrate 0.30%, magnesium sulfate 0.050%, potassium chloride 0.050%, and potato extract 20%. This 24 h grown mother culture (10 ml) was used to inoculate 50 ml of production medium containing (w/v): glucose 6.0%, peptone 0.20%, groundnut meal 0.50%, zinc sulfate 0.030%, calcium carbonate 0.040%, and potassium dihydrogen phosphate 0.10%. The pH of the medium was adjusted to 5.4 with HCl. Erlenmeyer flasks (500 ml) containing 50 ml of medium were incubated at 28ºC in an orbital shaker at 240 rpm for 49 h. The nuclease p1 solution was harvested by centrifugation at 3028g at 4ºC for 10 min, and the supernatant thus obtained was used as the crude enzyme preparation. Crude enzyme activity was 360 U/ml.
Immobilized Method
1.0 g DEAE cellulose was added to acetate buffer (100 ml, pH 5.4, 0.125 mol/l). Then it was mixed with glutaraldehyde solution (4.0 ml, 25%) for 2.0 h at room temperature. This support was washed using acetate buffer. After that, cross-linked DEAE cellulose could be obtained.
Cross-linked DEAE cellulose was mixed with nuclease p1 solution for 2.0 h at room temperature. After that, immobilized enzyme support was washed using acetate buffer.
The effect of several factors, such as solution pH, ionic strength, glutaraldehye concentration, immobilization time and enzyme amount, on immobilized enzyme activity were investigated systematically.
Characterizations of Immobilized Enzyme
Determination of pH Optimum and pH Stability
Effect of pH on free enzyme and immobilized enzyme was studied by assaying the enzymes at different pH values (4.07.5). pH stability was studied by pre-incubating the enzyme in buffers of different pH values (4.07.5), at 65ºC for 30 min. The remaining activities were determined by the method described in the item "Nuclease p1 assay".
Determination of Temperature Optimum and Thermal Stability
To determine the optimum temperature for free enzyme and immobilized enzyme, the enzyme activities of nuclease p1 were measured at various temperatures using RNA as the substrate. The thermal stability was studied by incubating the enzyme at 30, 40, 50, 60, 65, 70, 75, 80 and 90ºC for 30 min. The residual activities were determined by the method described in the item "Nuclease p1 assay".
Determination of Km and Vmax
Km and Vmax values of free enzyme and immobilized enzyme were determined by measuring enzyme activities with various concentrations (0.75, 1.5, 2.25, 3.00, 3.75, 4.50, 5.25, 6.00, 7.50, 9.00, 10.5 mg/ml) of RNA substrate. Kinetic constants were calculated as described by the LineweaverBurk method.
Nuclease p1 Assay
The activity of nuclease p1 was measured by the method of Fujishima et al. (Fujishima et al., 1972) and (Fujimoto et al., 1974). The nuclease p1 activity was measured in terms of the amount of acid-soluble nucleotides produced by RNA hydrolysis, which is catalyzed by nuclease p1. Enzyme solution (0.10 ml) or immobilized enzyme (0.10 mg) was incubated with substrate solution (1.0% RNA (w/v), 0.125 M acetate buffer, pH 5.4, 3.0 mM Zn2+) at 69ºC for 15 min. The reaction was stopped by adding 2.0 ml of ice-cold nucleic acid precipitator (0.25% ammonium molybdate dissolved in 2.5% perchloric acid, w/v). The mixture was mantained in an ice-bath for more than 10 min. The precipitated RNA was removed by centrifugation at 3028 g at 4ºC for 10 min. The supernatant liquid was diluted 50-fold with distilled water. The absorbance at 260 nm of the diluted solution was read with a blank incubation without enzymes. The activity (U) of nuclease p1 was calculated according to the following formula:
enzyme activity (U/ml)=OD260×a×4.0×50/(0.1×15)= OD260×a×133.3;
where a is a dilution factor of the enzyme before the enzyme activity assay.
One unit of enzyme activity was defined as the amount of enzyme that produced an increase in the optical density of 1.0 in 1 min at 260 nm.
Protein Assay
Protein was measured by the method of Bradford (Bradford, 1976). The rapid and sensitive method for quantification of microgram quantities of protein used the principle of protein dye-binding, with bovine serum albumin (BSA) as standard. The concentration of protein during purification studies was calculated from the standard curve.
RESULTS AND DISCUSSION
Preparation of Immobilized Enzyme
Effect of pH on Immobilized Enzyme Activity
Effect of pH on immobilized enzyme activity was studied. The results are shown in Fig. 1. Immobilized enzyme activity was the highest when pH of acetate buffer was 5.8 (Fig. 1). Immobilized enzyme activity was very low when the pH was below 5.0. This was mainly due to different pH, resulting in the difference of charge distribution of the carrier and enzyme, thereby affecting spatial structure of the immobilized enzyme.
Effect of Ionic Strength on Immobilized Enzyme Activity
Effect of ionic strength on immobilized enzyme activity was investigated. The results are shown in Fig. 2.
When ionic strength was below 0.125, immobilized enzyme activity increased with the increase of ionic strength (Fig. 2). But, when ionic strength was higher than 0.125, immobilized enzyme activity began to decrease with the increase of ionic strength. When ionic strength reached 0.50, immobilized enzyme activity was only half of the maximum. Competitive adsorption may be existent when ionic strength is high. Thus, immobilized enzyme activity was decreased. Another reason may be enzyme coagulation at high ionic strength. Thus, the buffer solution (ionic strength 0.125) was used for enzyme immobilization.
Effect of Glutaraldehyde Concentration on Immobilized Enzyme Activity
Effect of glutaraldehyde concentration on immobilized enzyme activity was studied. The results are shown in Fig. 3.
Immobilized enzyme activity (DEAE preparation activity) changed greatly with the increase of glutaraldehyde concentration (Fig. 3). When glutaraldehyde concentration was 0.20%, immobilized enzyme activity reached the maximum. However, when glutaraldehyde concentration was more than 0.60%, enzyme activity decreased with the increase of glutaraldehyde concentration. Glutaraldehyde is a cross-linking agent and also a denaturant. When excessive NH2 groups in the enzyme molecule reacted with glutaraldehyde, overcrowding of the enzyme molecule resulted in a change of the enzyme structure.
Effect of Immobilization Time on Immobilized Enzyme Activity
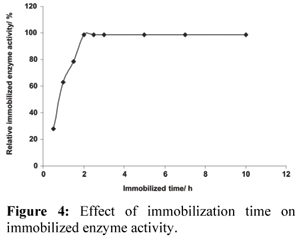
Effect of immobilization time on immobilized enzyme activity was investigated. The results are shown in Fig. 4.
Immobilized enzyme activity was increased with the increase of immobilization time. When the immobilization time reached 2.0 h, the maximum enzyme activity was obtained. Immobilized enzyme activity remained unchanged even if the immobilization time was increased. This indicates that the available space on the immobilization carrier becomes saturated with the increase of the immobilization time.
e) Effect of Enzyme Amount on Immobilized Enzyme Activity
Effect of enzyme amount on immobilized enzyme activity was investigated. The results are shown in Fig. 5. Immobilized enzyme activity increased with the increase of enzyme amount from 0.5 ml to 6.0 ml and remained unchanged with the increase of enzyme amount (Fig. 5). Before the binding sites on the surface of DEAE cellulose were saturated, immobilized enzyme activity increased with enzyme amount. However, once it was saturated, immobilized enzyme activity can not be increased by an increase of enzyme amount. When the enzyme amount was 6.0 ml, enzyme activity recovery was only 50% (Fig. 5). So, an enzyme amount of 4.0 ml was chosen for enzyme immobilization.
f) Optimum Immobilization Conditions
According to the results of One-factor-at-a-time method, four factors that mainly affected the enzyme activity were selected to determine which factors were the significant factors. These factors were pH, ionic strength, glutaraldehyde concentration and immobilization time. Orthogonal design is shown in Tab. 1. Orthogonal experimental results and analysis of the results are shown in Tab. 2 and 3, respectively.
The factors were ranked according to R (contribution percentage), namely, B>A> D>C. The optimal condition was A3B3C2D1 (Tab. 1 and 2). That was pH 5.6, ionic strength 0.125 glutaraldehyde concentration 0.20% and immobilization time 2.0 h.
Significant factors affecting enzyme activity were pH and ionic strength (Tab. 3). Therefore, during preparation of immobilized enzyme, the most important thing was to control pH and ionic strength.
To confirm the optimal conditions, a set of four replicate experiments with the optimal conditions were carried out. The average relative enzyme activity was 100.3%.
CHARACTERIZATIONS OF IMMOBILIZED ENZYME
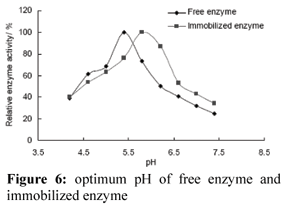
Optimum pH
Compared with free enzyme, the optimum pH of immobilized enzyme increased 0.5 units (Fig. 6). This result is in agreement with that of peroxidase immobilized on DEAE cellulose. The pH-optimum of immobilized peroxidase shifted to 5.6, while free enzyme showed a pH-optimum at pH 5.0 (Kulshrestha and Husain, 2006). These differences in the behaviors of free and immobilized enzyme could be explained by the poly-cationic nature of a support like DEAE- cellulose that attracts more OH- ions around the immobilized enzyme, thus making the pH of the enzyme's micro-environment higher than the bulk solution. Immobilized enzyme therefore requires a higher pH for optimal activity than free enzyme.
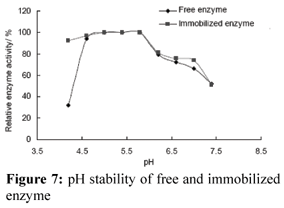
pH Stability of Enzyme
The pH stability of the enzyme was enhanced when the enzyme was immobilized (Fig. 7). It was not sensitive to pH change. This result is the same as in other work where we found that the pH stability of nuclease p1 was increased when it was immobilized on chitosan microspheres (not published). Enzyme may be encapsulated in the micro-environment of DEAE-cellulose. The change of solution pH did not affect the protection layer of the supports or maybe there is an interaction between the support and nuclease p1, making the enzyme conformation more stable.
Optimum Temperature
The effect of temperature on enzyme activity is shown in Fig. 8. Optimum temperature of immobilized enzyme was increased 10ºC. This result is similar to that reported by Fahmy et al. (Fashmy et al., 1998). In their study, urease was immobilized on DEAE-cellulose. The optimum temperature of immobilized enzyme was shifted to 65ºC, whereas free enzyme had an optimum temperature of approximately 55ºC. Similarly, optimum temperature of immobilized single-strand specific nuclease on gelatine-alginate was increased to 65ºC (Reddyand Shankar, 1987). This means that immobilized enzyme has a higher resistance to the change of temperature than free enzyme. At low temperatures, the activity of free enzyme was higher than that of immobilized enzyme. This may reflect molecular diffusion resistance and the conformational change of enzyme immobilized on the supports. At high temperatures, the enzyme activity of both free enzyme and immobilized enzyme decreased sharply with the increase of temperature. A possible explanation is that, with the increase of temperature, the concentration of the enzyme molecules opened up and they ultimately lost their biocatalytic activity.
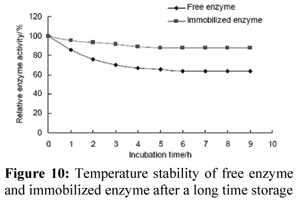
Temperature Stability of Enzyme
Free and immobilized enzyme was incubated at different temperatures for half an hour, respectively (Fig. 9). The activity of both free enzyme and immobilized enzyme decreased sharply with increasing temperature up to 70ºC. The activity of immobilized enzyme on DEAE cellulose remained more than 85% after it was incubated at 70ºC for 9.0 h (Fig. 10). However, only 63% of the activity of free enzyme remained under the same conditions. Compared with free enzyme, the thermal stability of enzyme immobilized on DEAE cellulose was greatly increased, probably because of the greater stability of the three-dimensional structures of immobilized enzyme. The improvement of the optimum temperature and thermal stability of the enzyme is favorable for its industrial application. Kulshresha and Hussin studied the thermal denaturation behavior of free and immobilized peroxidase on DEAE cellulose (Karboune et al., 2005). In their study, free peroxidase incubated at 60ºC for 2 h retained 43% of its initial enzyme activity, while the immobilized enzyme incubated under similar conditions was significantly more stable to heat inactivation. The immobilized peroxidase exhibited 60% of the original activity after 2 h of heat treatment. Similar results were also reported by Fahmy et al. (Fashmy et al., 1998).
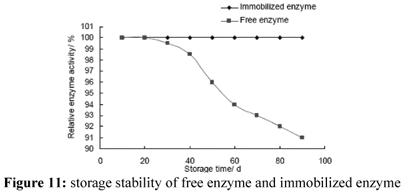
Storage Stability of Enzyme
Storage stability of immobilized enzyme was quite good (Fig. 11). The activity remained unchanged at 4ºC after 90 days. However, more than 10% activity of free enzyme was lost after 90 days. Generally, the combination of immobilized carrier with enzyme increases enzyme stability. The increase of storage stability of immobilized enzyme is very favourable for its industrial application. Fahmy et al. studied storage stability of free and immobilized enzyme on DEAE cellulose at 4ºC and 25ºC, respectively. Their results shown storage stability of immobilized enzyme was better than that of free enzyme (Fashmy et al., 1998). This result is consistent with our result.
Measurement of the Michaelis Constant
Determination of free and immobilized nuclease p1 Km and Vmax, was studied. The results are shown in Fig. 12 and Tab. 4. The Michaelis constant of the immobilized enzyme was a little higher than that of the free enzyme (Tab. 4 and Fig. 12). This increase may have resulted from conformational and steric modifications introduced by the attachment of the enzyme to the support and to mass transfer resistances inherent in the morphology of the support used, thus reducing the affinity of the substrate for the active site of the enzyme. This increase in Km of the immobilized enzyme is in agreement with the result reported by Fahmy et al. Km was about 3.8 times higher for immobilized enzyme than for free enzyme (Fashmy et al., 1998).
The enzyme activity was determined under standard assay conditions. Symbols: S RNA concentration (mg/ml); V reaction velocity (mg/mlmin)
CONCLUSIONS
Nuclease p1 immobilized on DEAE cellulose was studied in this paper. Optimal immobilization conditions were as follows: pH 5.6, ionic strength 0.125, glutaraldehyde concentration 0.20% and immobilization time 2.0 h. Characterizations of the immobilized enzyme showed that the optimum pH, optimum temperature and storage stability are significantly higher than those of free enzyme. This made immobilized enzyme better adapted to the change of pH and temperature, more conducive to industrial production.
(Submitted: February 26, 2009 ; Revised: May 5, 2009 ; Accepted: July 6, 2009)
- Blanco, R. M., Terreros, P., Fernández-Pérez, M., Otero, C. and Diáz-González, G., Functionalization of mesoporous silica for lipase immobilization: Characterization of the support and the catalysts. Journal of Molecular Catalysis B: Enzymatic, v. 30, 83-93 (2004).
- Bradford, M., A rapid and sensitive method for quantitation of microgram quantities of protein using the principle of protein dye-binding. Analytical Biochemistry, v. 72, 248-254 (1976).
- Caramori, S. S. and Fernandes, K. F., Covalent immobilisation of horseradish peroxidase onto poly(ethylene terephthalate)poly(aniline) composite. Process Biochemistry, v. 39, 883-888 (2004).
- Cheng, J., Yu, S. M. and Zuo, P., Horseradish peroxidase immobilized on aluminum-pillared interlayered clay for the catalytic oxidation of phenolic wastewater. Water Research, v. 40, 283-290 (2006).
- Dai, Z., Ju, H. and Chen, H., Mesoporous Materials Promoting Direct Electrochemistry and Electrocatalysis of Horseradish Peroxidase. Electroanalysis, v. 17, 862-868 (2005).
- Fashmy, A. S., Bagos, V. B. and Mohammed, T. M., Immobilization of Citrullus vulgaris urease on cyanuric chloride DEAE-cellulose ether, preparation and properties. Bioresource Technology, v. 64, 121-129 (1998).
- Fernandes, K. F., Lima, C. S., Lopes, F. M. and Collins, C. H., Properties of horseradish peroxidase immobilised onto polyaniline. Process Biochemistry, v. 39, 957-962 (2004).
- Fujimoto, M., Kuninaka, A., Yoshino, H., Purification of a nuclease from Penicillium citrinum Agricultural & Biological Chemistry, v. 38, 777-783 (1974).
- Fujishima, T., Uchida, K. and Yushino, H., Enzyme production by molds in sponge culture. Journal of Fermentation Technology, v. 50, 724-730 (1972).
- Gerald, R. and Tilak, W., Yeast derived products in Yeast technology. (2nd ed.), New York (1991).
- Goradia, D., Cooney, J., Hodnett, B. K. and Magner, E., The adsorption characteristics, activity and stability of trypsin onto mesoporous silicates. Journal of Molecular Catalysis B, Enzymatic, v. 32, 231-239 (2005).
- Karboune, S., Archelas, A., Furstoss, R., Baratti, J., Immobilization of epoxide hydrolase from Aspergillus niger onto DEAE-cellulose: enzymatic properties and application for the enantioselective resolution of a racemic epoxide. Journal of Molecular Catalysis B, Enzymatic, v. 32, 175-183 (2005)
- Kulshrestha, Y. and Husain, Q., Direct immobilization of peroxidase on DEAE cellulose from ammonium sulfate fractionated proteins of bitter gourd (Momordica charantia). Enzyme and Microbial Technology, v. 38, 470-477 (2006).
- Kuninaka, A., Nucleotides and related compounds. Germany (1996).
- Kuninaka, A., Kibi, M., Yoshino, H. and Sagaguchi, K., Studies on 50-phosphodiesterase in microorganisms. Part II, Properties and application of Penicillium citrinum 50-phosphodiesterase. Agricultural & Biological Chemistry, v. 25, 693-701 (1961).
- Lai, Y. C. and Lin, S. C., Application of immobilized horseradish peroxidase for the removal of p-chlorophenol from aqueous solution. Process Biochemistry, v. 40, 1167-1174 (2005).
- Luo, X. L., Xu, J. J., Zhao, W. and Chen, H. Y., Glucose biosensor based on ENFET doped with SiO2 nanoparticles. Sensors and Actuators B, Chemical, v. 97, 249-255 (2004).
- Panya, P. H., Jasra, R. V., Newalkar, B. L. and Bhatt, P. N., Studies on the activity and stability of immobilized α-amylase in ordered mesoporous silicas. Microporous and Mesoporous Materials, v. 77, 67-77 (2005).
- Reddy, L. G. and Shankar, V., Immobilization of single-strand specific nuclease (S1 nuclease) from Aspergillus oryzae Applied Biochemistry and Biotechnology, v. 14, 231-240 (1987).
- Steensma, A., van Dijck, P. W. M, and Hempenius, R. A., Safety evaluation of phosphodiesterase derived from Leptographium procerum. Food and Chemical Toxicology, v. 42, 935-944 (2004).
- Xu, J. Z., Zhang, Y., Li, G. X. and Zhu, J. J., An electrochemical biosensor constructed by nanosized silver particles doped solgel film. Materials Science and Engineering, C. v. 24, 833-836 (2004).
- Xu, Q., Mao, C., Liu, N. N., Zhu, J. J. and Shen, J., Immobilization of horseradish peroxidase on O-carboxymethylated chitosan/solgel matrix. Reactive and Functional Polymers, v. 66, 863-870 (2006).
- Ying, G. Q., Shi, L.E., Tang, Z.X. and Shan, J. F., The applications of mononucleotides and produing methods. Food Research and Development, v. 25, 120-123 (2004).
- Zhao, Z. X., Qiao, M. Q., Yin, F., Shao, B., Wu, B. Y., Wang, Y. Y., Wang, X. S., Qin, X., Li, S., Yu, L. and Chen, Q., Amperometric glucose biosensor based on self-assembly hydrophobin with high efficiency of enzyme utilization. Biosensors and Bioelectronics, v. 22, 3021-3027 (2007).
Publication Dates
-
Publication in this collection
14 Apr 2010 -
Date of issue
Mar 2010
History
-
Reviewed
05 May 2009 -
Received
26 Feb 2009 -
Accepted
06 July 2009