Abstract
INTRODUCTION:
The aim of this study was to evaluate some virulence factors in Candida albicans isolates from patients with onychomycosis and determine the correlation between these factors and the antifungal resistance profile.
METHODS:
Seventy species of C. albicans were confirmed using polymerase chain reaction amplification of the HWP1 gene. According to the Clinical & Laboratory Standards Institute guidelines, the susceptibility profile of four antifungal agents was investigated, and the production of aspartyl protease, phospholipase, haemolysin, and biofilm was determined. The correlation between these profiles was also investigated.
RESULTS:
The isolates indicated different levels of resistance and production of virulence factors. Significant correlations were observed between the minimum inhibitory concentration (MIC) of fluconazole/itraconazole and biofilm production, between phospholipase production and fluconazole/itraconazole MIC, and between fluconazole MIC and hemolytic activity in C. albicans isolates. The results also showed significant correlations between phospholipase activity and biofilm production.
CONCLUSIONS:
Our findings will contribute to a better understanding of the pathogenesis of C. albicans and characterize the relationship between virulence factors and antifungal resistance, which may suggest new therapeutic strategies considering the possible involvement of the virulence mechanism in the effectiveness of treatment.
Keywords:
Candida albicans; Virulence factors; Biofilm; Antifungal agents
INTRODUCTION
Candida species are opportunistic fungi that are involved in a wide range of superficial to systemic diseases. Among non-invasive infections, onychomycosis, which is a fungal nail infection caused by Candida species, is the most common cause of onychomycosis following dermatophytes11. Maraki S, Mavromanolaki VE. Epidemiology of onychomycosis in Crete, Greece: a 12-year study. Mycoses. 2016;59(12):798-802.,22. Otašević S, Barac A, Pekmezovic M, Tasic S, Ignjatović A, Momčilović S, et al. The prevalence of Candida onychomycosis in Southeastern Serbia from 2011 to 2015. Mycoses. 2016;59(3):167-72.. Due to the increase in the number of patients with immunodeficiency, as well as changes in fungal pathogenicity and antifungal resistance, Candida species have gained considerable attention as important pathogenic organisms33. Gupta AK, Daigle D, Carviel JL. The role of biofilms in onychomycosis. J Am Acad Dermatol. 2016;74(6):1241-6..
Secretory proteinases, as Candida virulence factors, can improve the potential of fungal organisms to colonize and penetrate into the host tissue and disrupt the immune system44. Naglik J, Albrecht A, Bader O, Hube B. Candida albicans proteinases and host/pathogen interactions. J Cell Microbiol. 2004;6(10):915-26.. Phospholipase production is another major virulence factor of C. albicans, which binds the fungus to the target tissue and generates a pathway to enter the tissue following the hydrolysis of phospholipids and degradation of cell membranes55. Schaller M, Borelli C, Korting HC, Hube B. Hydrolytic enzymes as virulence factors of Candida albicans. Mycoses. 2005;48(6):365-77.. In addition, haemolysin, as another extracellular enzyme, contributes to the invasion of yeasts through absorption of iron66. Sacristan B, Blanco M, Galan-Ladero M, Blanco J, Perez-Giraldo C, Gomez-Garcia A. Aspartyl proteinase, phospholipase, hemolytic activities and biofilm production of Candida albicans isolated from bronchial aspirates of ICU patients. J Med mycol. 2011;49(1):94-7..
Biofilm formation, which is another virulence factor of C. albicans, plays a pivotal role in the pathogenesis of fungi through the mass production of pseudohyphae77. Sav H, Rafati H, Öz Y, Dalyan-Cilo B, Ener B, Mohammadi F, et al. Biofilm formation and resistance to fungicides in clinically relevant members of the fungal genus fusarium. J Fungi. 2018;4(1):16.
8. Mukherjee PK, Chandra J, Kuhn DM, Ghannoum MA. Mechanism of fluconazole resistance in Candida albicans biofilms: phase-specific role of efflux pumps and membrane sterols. J Infect immun. 2003;71(8):4333-40.-99. Seidler MJ, Salvenmoser S, Müller F-MC. Aspergillus fumigatus forms biofilms with reduced antifungal drug susceptibility on bronchial epithelial cells. J Antimicrob Agents chemother. 2008;52(11):4130-6.. Biofilm contains dense yeast cells and hyphae, which can lead to antifungal resistance, as well as frequent recurrence of Candida infections. Studies indicate that C. albicans uses the nail as the substrate for biofilm formation. In fact, the production of biofilms in the nail acts as a source of continuous infection and leads to limited penetration of antifungal agents into the Candida biofilm33. Gupta AK, Daigle D, Carviel JL. The role of biofilms in onychomycosis. J Am Acad Dermatol. 2016;74(6):1241-6.,1010. Vila TVM, Rozental S, de Sá Guimarães CMD. A new model of in vitro fungal biofilms formed on human nail fragments allows reliable testing of laser and light therapies against onychomycosis. Lasers Med Sci. 2015;30(3):1031-9.. Furthermore, long-term treatment and excessive exposure to antifungals lead to variable levels of resistance in Candida species. The purpose of this study was to determine the antifungal susceptibility profile of four antifungals in vitro, examine the production of aspartyl protease, phospholipase, haemolysin, and biofilm, and assess the relationship between these characteristics and the minimum inhibitory concentration (MIC) of antifungals for C. albicans isolates from patients with clinical onychomycosis in Iran.
METHODS
Fungal strains
This study was conducted on C. albicans isolates (n = 70), obtained from patients with onychomycosis, who were admitted to the Dermatology Clinic of Razi Skin Hospital in Tehran, Iran from November 2017 to October 2018. The physiological characteristics of the isolates were identified using the germ tube test1111. Mackenzie D. Serum tube identification of Candida albicans. J Clin Pathol. 1962;15(6):563-5. and colony color analysis on CHOROM agar Candida medium (CHOROMagar Candida, France)1212. Al-Wasify RS. Sensitivity and specificity of chromogenic media for detection of some pathogens in water. Int J Environ Sustain. 2013;2(1).. All isolates were storedat−20°C untiluse. Before the experiments, these isolates were grownon Sabouraud dextrose agar (SDA, Acumedia, Baltimore, Maryland, USA) supplemented with 100 μg/ml cycloheximide (Sigma, Steinheim,Germany) and incubated at a temperature ranging from 25-30°C.
Next, genomic DNA was extracted using glass beads and phenol:chloroform:isoamyl alcohol (25:24:1) method1313. Yamada Y, Makimura K, Merhendi H, Ueda K, Nishiyama Y, Yamaguchi H, et al. Comparison of different methods for extraction of mitochondrial DNA from human pathogenic yeasts. Jpn J Infect Dis. 2002;55(4):122-5.. All clinical C. albicans isolates were confirmed through PCR amplification of the hyphal wall protein 1 (HWP1) gene, using forward (5′GCTACCACTTCAGAATCATCATC-3′) and reverse (5′ GCACCTTCAGTCGTAGAGACG-3′) primer pairs1414. Romeo O, Criseo G. First molecular method for discriminating between Candida africana, Candida albicans, and Candida dubliniensis by using hwp1 gene. Diagn Microbiol Infect Dis. 2008;62(2):230-3..
Antifungal susceptibility test
The antifungal susceptibility profiles were determined in vitro using the broth microdilution method, according to the M27-A3 guidelines of the Clinical and Laboratory Standards Institute (CLSI)1515. Clinical and Laboratory Standards Institute (CLSI). Reference method for Broth Dilution antifungal susceptibility testing of yeasts; approved standard -third edition. Document M27-A3.Wayne, PA: CLSI; 2008. 13p.. All isolates were evaluated against amphotericin B (AMB), fluconazole (FLC), itraconazole (ITC), and voriconazole (VRC) (Sigma-Aldrich Chemical Corporation, St. Louis, MO, USA). Briefly, each well of the 96-well round-bottom microtiter plate consisted of 100 μL of RPMI-1640 medium, 64-0.06 μg/mL of FLC, and 16-0.016 μg/mL of AMB, ITC, and VRC with 1-5×103 cells/mL of yeast. The plates were then incubated at 35°C for 48 h. Candida parapsilosis ATCC 22019 and Candida krusei ATCC 6258 were also used as quality controls.
Production of hydrolytic enzymes
The activity of aspartyl protease was determined using bovine serum albumin (BSA) (pH, 5.0) agar assay, as previously described1616. Chakrabarti A, Nayak N, Talwar P. In vitro proteinase production by Candida species. Mycopathol. 1991;114(3):163-8.. The assay was performed by adding 10 µL of cell suspension (McFarland turbidity, 0.5) to a sterile paper disk, which was placed on the surface of BSA medium, followed by incubation at 37°C for six days. Next, the plates were fixed with 20% trichloroacetic acid (TCA) and were stained with 1.25% Amido Black dye1717. Gokce G, Cerikcioglu N, Yagci A. Acid proteinase, phospholipase, and biofilm production of Candida species isolated from blood cultures. Mycopathol. 2007;164(6):265.. C. albicans ATCC 10231 was used as the positive control.
The production of phospholipases by C. albicans isolates was achieved using egg yolk agar plate method1818. Price MF, Wilkinson ID, Gentry LO. Plate method for detection of phospholipase activity in Candida albicans. Sabouraudia: J Med Vet Mycol. 1982;20(1):7-14.. The test was conducted by adding 10 μL of cell suspension (0.5 McFarland turbidity), inoculated onto SDA plates (containing 1 M NaCl, 0.005 M CaCl2, and 8% sterile egg yolk emulsion), followed by incubation at 37°C for four to five days. C. albicans SC 5314 strain was used as the positive control. In addition, the hemolytic activity of C. albicans isolates was evaluated in the SDA medium with 7% blood and 3% glucose1919. Manns JM, Mosser DM, Buckley HR. Production of a hemolytic factor by Candida albicans. Infect Immun. 1994;62(11):5154-6.. Similarly, 10 µL of the yeast suspension (108 cells) was inoculated onto the surface of the medium, and each plate was incubated at 37°C for two to three days. C. albicans ATCC 10261 was used as the control strain.
Furthermore, enzyme activity (Prz, Pz, and Hz) was determined for each isolate by measuring the ratio of colony diameter to the colony diameter plus the area of zone. According to the results, four classes of isolates were identified with respect to the enzyme activity zone: 1, negative activity; 0.999-0.700, weak producer; 0.699-0.400, moderate producer; and < 0.399, strong producer1818. Price MF, Wilkinson ID, Gentry LO. Plate method for detection of phospholipase activity in Candida albicans. Sabouraudia: J Med Vet Mycol. 1982;20(1):7-14.. Each isolate was tested in duplicate.
Biofilm formation
Biomass was assessed using crystal violet assay2020. Chatzimoschou A, Katragkou Α, Simitsopoulou M, Antachopoulos C, Georgiadou E, Walsh TJ, et al. Activities of triazole-echinocandin combinations against Candida species in biofilms and as planktonic cells. J Antimicrob Agents chemother . 2011;55(5):1968-74.. Briefly, C. albicans isolates, which were grown on SDA at 37°C for 24 h, were washed, and suspension was adjusted to 3×107 CFU/mL. Next, 20 µL of the yeast cell suspension and 180 µL of Sabouraud dextrose broth (SDB) supplemented with 60 g of glucose, were inoculated into flat-bottom 96-well microplates. The plates were then incubated at 35°C for 24 h. After incubation and washing, the plate walls were stained with 1% crystal violet for 15 min, following which 200 μL of 95% ethanol was added to each well2121. Aggarwal P, Kashyap B. Biofilm production by clinically isolated Candida: Comparative analysis based on specimen, methodology, and various Candida species. Indian J Med Speciali. 2018;9(2):69-72..
The optical density (OD) was measured using an ELISA reader at 590 nm. The OD cut-off point (ODc) for biofilm formation was determined as three standard deviations above the mean absorbance of the negative control2222. Stepanović S, Vuković D, Dakić I, Savić B, Švabić-Vlahović M. A modified microtiter-plate test for quantification of staphylococcal biofilm formation. J Microbiol Methods. 2000;40(2):175-9.. The isolates were classified into four groups according to OD590: non-biofilm producer, OD590 ≤ ODc; weak producer, ODc < OD590 ≤ 2 × ODc; moderate producer, 2 × ODc < OD590 ≤ 4 × ODc; and strong producer, OD590 nm > 4 × ODc (1, 2). The ODc measured in this study was 0.365.
Scanning electron microscopic (SEM) study of Candida biofilms
In this study, C. albicans isolates with the highest biofilm-forming ability among the ones tested were selected and analyzed through scanning electron microscopy (SEM), according to a previously described method2323. Brilhante RSN, Bittencourt PV, de Souza Collares Castelo-Branco D, de Melo Guedes GM, de Oliveira JS, Alencar LP, et al. Biofilms of Candida spp. from the ocular conjunctiva of horses with reduced azole susceptibility: a complicating factor for the treatment of keratomycosis? Vet Ophthalmol. 2017;20(6):539-46.. After incubation and removal of free cells by washing, biofilms and sticky cells were fixed with glutaraldehyde and incubated at 4°C overnight. After the biofilms were washed with PBS, they were dehydrated with various dilutions of ethanol (50%, 70%, 80%, 95%, and 100%), and the plates were dried in air. Finally, the biofilms were coated with gold and studied through SEM.
Statistical analysis
Statistical analysis was performed using SPSS version 22.0 (SPSS Inc., Chicago, IL, USA). Normality assumption was evaluated using the Kolmogorov-Smirnov test, which showed that the data were not normally distributed. Moreover, Spearman's rank-order correlation test was used to evaluate the correlation between antifungal MICs, extracellular enzymes, and biofilm production. The median values of variables and 95% confidence interval for median (95% CI) were compared for each group of antifungal profile, using Mann-Whitney U test. P-value < 0.05 was considered statistically significant.
RESULTS
Identification and antifungal susceptibility profile of C. albicans isolates
The results of germ tube test in the serum and green colony formation on CHOROM agar candida medium were confirmed using PCR-HWP1 assay for C. albicans isolates (~1000 bp). The antifungal susceptibility test was performed in vitro based on the CLSI M27-A3 guidelines, shown in Table 1. All C. albicans isolates (100%) were susceptible to AMB and VRC at concentrations ≤ 1 μg/mL, whereas eight isolates (11.4%) were resistant to FLC (MIC ≥ 64 μg/mL), and 19 isolates (27.1%) were susceptible-dose dependent (SDD). Furthermore, six isolates (8.6%) showed the highest MIC to ITC (MIC ≥ 1 μg/mL), while 18 isolates (25.7%) were SDD.
Production of hydrolytic enzymes and biofilm formation
Evaluation of proteinase production showed that 72.85% of C. albicans isolates produced aspartyl protease (Prz range, 0.32 to 0.69). In addition, phospholipase activity was detected in 61.42% of C. albicans isolates (Pz range, 0.22 to 0.96). Overall, 17 isolates (24.3%) were considered as strong producers (Pz range, 0.22 to 0.39), 15 isolates (21.4%) were classified as moderate producers (Pz range, 0.43 to 0.69), and 11 isolates (15.7%) were considered as weak producers (Pz range, 0.73 to 0.96). In this study, we observed that 94.28% of C. albicans isolates from the nail bed of patients with onychomycosis were able to induce hemolytic activity (Hz range, 0.33 to 0.69). In total, 22.9% of isolates were considered as strong producers (Hz range, 0.33 to 0.38), while 71.4% of isolates were classified as moderate producers (Hz range, 0.47 to 0.69). Additionally, biofilm production of 70 C. albicans isolates were classified into four groups according to OD590: non-biofilm producers (n = 47; 67.14%), weak (n = 7; 10%), moderate (n = 13; 18.6%), and strong (n = 3; 4.3%). Furthermore, Figure 1 shows the SEM results of the network of dense yeast cells and hyphae by C. albicans with strong biofilm producer (OD590 > 1.46).
Biofilm structure of the network of dense yeast cells and hyphae for C. albicans with strong biofilm production (OD590 > 1.46). Scanning electron microscopy images acquired after biofilm growth in SDB after 24 h with 1000X, 4000X, and 15000 X, magnification.
The results also showed significant correlations between phospholipase activity and biofilm production in C. albicans isolates (P = 0.004, rs = 0.339). In addition, there was no significant relationship between aspartyl protease, hemolytic activities, and biofilm production.
Correlation between antifungal susceptibility profile and virulence factors
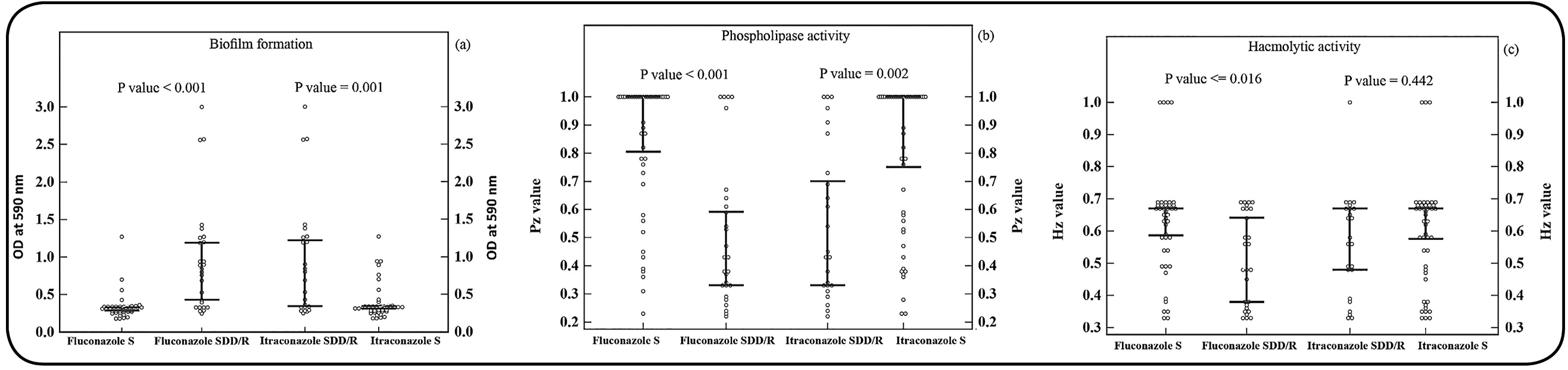
Evaluation of the correlation between antifungal MICs and virulence factors revealed significant relationships between FLC MIC and biofilm formation (P = 0.05, rs = 0.725), ITC MIC and biofilm formation (P = 0.002, rs = 0.498), FLC MIC and phospholipase production (P = 0.05, rs = 0.455), ITC MIC and phospholipase production (P = 0.003, rs = 0.345), and FLC MIC and hemolytic production (P = 0.011, rs = 0.302) (Table 2). The differences in the median production value of phospholipase, haemolysin, and biofilm formation by C. albicans isolates with their susceptibility profile against fluconazole and itraconazole antifungal are shown in Figure 2.
Differences in the median (95% CI for median) phospholipase index Pz values, hemolytic activity Hz values, and optical density at 590 nm of formed biofilms between strains with different fluconazole and itraconazole susceptibility profiles with Mann-Whitney test. (a) Absorbance value (OD 590 nm) of formed biofilms with FLC and ITC susceptibility profiles. (b) Median Pz values of phospholipase of strains with FLC and ITC susceptibility profiles. (c) Median Hz values of hemolytic activity of strains with FLC and ITC susceptibility profiles. In all the cases, the differences between the medians of groups with different susceptibility profiles were statistically significant (P < 0.05), except for Hz values of hemolytic activity with itraconazole susceptibility profile (P>0.05). S: susceptible; SDD: susceptible-dose dependent; R: resistant; 95% CI: 95% confidence interval.
DISCUSSION
Candida species are regarded as the second most common cause of onychomycosis after dermatophytes. However, the growing prevalence of onychomycosis due to different varieties of Candida species has been reported in several parts of the world in recent years22. Otašević S, Barac A, Pekmezovic M, Tasic S, Ignjatović A, Momčilović S, et al. The prevalence of Candida onychomycosis in Southeastern Serbia from 2011 to 2015. Mycoses. 2016;59(3):167-72.. The increasing resistance of C. albicans to antifungals possibly indicates the relationship between the antifungal resistance mechanisms and production of biofilms and extracellular enzymes33. Gupta AK, Daigle D, Carviel JL. The role of biofilms in onychomycosis. J Am Acad Dermatol. 2016;74(6):1241-6.,1010. Vila TVM, Rozental S, de Sá Guimarães CMD. A new model of in vitro fungal biofilms formed on human nail fragments allows reliable testing of laser and light therapies against onychomycosis. Lasers Med Sci. 2015;30(3):1031-9..
The antifungal susceptibility pattern identified in this study indicated that all isolates were highly susceptible to AMB (100%) and VRC (100%). According to the present results, which are supported by previous studies, susceptibility to AMB and VRC was detected in C. albicans isolates2424. Silva-Dias A, Miranda IM, Branco J, Monteiro-Soares M, Pina-Vaz C, Rodrigues AG. Adhesion, biofilm formation, cell surface hydrophobicity, and antifungal planktonic susceptibility: relationship among Candida spp. Front microbiol. 2015;6:205.,2525. Sav H, Baris A, Turan D, Altinbas R, Sen S. The frequency, antifungal susceptibility and enzymatic profiles of Candida species in cases of onychomycosis infection. Microb Pathog. 2018;116:257-62.. In contrast, Pakshir et al. reported that 43% of C. albicans isolates from patients with onychomycosis were resistant to VRC in Iran2626. Pakshir K, Zomorodian K, Zakaei A, Motamedi M, Ghiasi MR, Karamitalab M. Molecular identification and in-vitro antifungal susceptibility testing of Candida species isolated from patients with onychomycosis. Curr med mycol. 2015;1(4):26.. In addition, our results showed that 11.4% and 8.6% of C. albicans isolates were resistant to FLC and ITC, respectively. These findings are consistent with those of previous studies conducted in Portugal and China2323. Brilhante RSN, Bittencourt PV, de Souza Collares Castelo-Branco D, de Melo Guedes GM, de Oliveira JS, Alencar LP, et al. Biofilms of Candida spp. from the ocular conjunctiva of horses with reduced azole susceptibility: a complicating factor for the treatment of keratomycosis? Vet Ophthalmol. 2017;20(6):539-46.,2424. Silva-Dias A, Miranda IM, Branco J, Monteiro-Soares M, Pina-Vaz C, Rodrigues AG. Adhesion, biofilm formation, cell surface hydrophobicity, and antifungal planktonic susceptibility: relationship among Candida spp. Front microbiol. 2015;6:205..
Moreover, Khosravi et al. reported that 85.7% of Candida species isolated from the nails were susceptible to FLC in Iran2727. Khosravi A, Shokri H, Mansouri P, Katiraee F, Ziglari T. Candida species isolated from nails and their in vitro susceptibility to antifungal drugs in the department of Dermatology (University of Tehran, Iran). J Mycol Med. 2008;18(4):210-5.. The cause of the difference in antifungal resistance patterns of Candida species is related to clinical samples, immune system status, extensive use of azoles, and different distribution patterns of isolates in different geographical areas.
The extracellular enzymes of C. albicans play a pivotal role in binding, invasion, and destruction of the host cellular structure. C. albicans has the ability to synthesize aspartyl proteinase through the destruction of elastin, which is involved in the invasion of host tissues. Based on our results, the highest proteinase activity was observed in 72.85% of C. albicans. Previous studies have reported the rate of proteinase production to be 40%2828. El-Houssaini HH, Elnabawy OM, Nasser HA, Elkhatib WF. Correlation between antifungal resistance and virulence factors in Candida albicans recovered from vaginal specimens. Microb Pathog . 2019;128:13-9., 82.1%2929. Sachin C, Ruchi K, Santosh S. In vitro evaluation of proteinase, phospholipase and haemolysin activities of Candida species isolated from clinical specimens. Inter j Med Biomed res. 2012;1(2):153-7., and 97%3030. Mattei AS, Alves SH, Severo CB, Guazzelli LdS, Oliveira FdM, Severo LC. Determination of germ tube, phospholipase, and proteinase production by bloodstream isolates of Candida albicans. Rev Soc Bras Med Trop. 2013;46(3):340-2.. Additionally, phospholipases are responsible for the hydrolysis of phospholipids, which help Candida invade the cells and destroy the host cell membrane.
According to our results, 61.42% of C. albicans isolates produced phospholipases. In a study conducted by Sav et al., the highest phospholipase activity was reported in C. albicans isolates from nail samples2525. Sav H, Baris A, Turan D, Altinbas R, Sen S. The frequency, antifungal susceptibility and enzymatic profiles of Candida species in cases of onychomycosis infection. Microb Pathog. 2018;116:257-62.. Ramos Lde et al. reported that all C. albicans isolates (100%) from cutaneous candidiasis produced both aspartyl protease and phospholipase3131. de Souza Ramos L, Barbedo LS, Braga-Silva LA, Dos Santos ALS, Pinto MR, da Graça Sgarbi DB. Protease and phospholipase activities of Candida spp. isolated from cutaneous candidiasis. Rev Iberoam Micol. 2015;32(2):122-5.. In addition, El-Houssaini et al. reported that 42% of C. albicans isolates from vaginal swab specimens produced phospholipase2828. El-Houssaini HH, Elnabawy OM, Nasser HA, Elkhatib WF. Correlation between antifungal resistance and virulence factors in Candida albicans recovered from vaginal specimens. Microb Pathog . 2019;128:13-9..
Iron uptake through haemolysin is an essential requirement for the growth and invasion of fungi55. Schaller M, Borelli C, Korting HC, Hube B. Hydrolytic enzymes as virulence factors of Candida albicans. Mycoses. 2005;48(6):365-77.. In this study, 66 C. albicans isolates (94.28%) produced haemolysin, which is in agreement with the findings of recent studies66. Sacristan B, Blanco M, Galan-Ladero M, Blanco J, Perez-Giraldo C, Gomez-Garcia A. Aspartyl proteinase, phospholipase, hemolytic activities and biofilm production of Candida albicans isolated from bronchial aspirates of ICU patients. J Med mycol. 2011;49(1):94-7.,2929. Sachin C, Ruchi K, Santosh S. In vitro evaluation of proteinase, phospholipase and haemolysin activities of Candida species isolated from clinical specimens. Inter j Med Biomed res. 2012;1(2):153-7.,3232. Pakshir K, Zomorodian K, Karamitalab M, Jafari M, Taraz H, Ebrahimi H. Phospholipase, esterase and hemolytic activities of Candida spp. isolated from onychomycosis and oral lichen planus lesions. J Mycol Med . 2013;23(2):113-8., suggesting that hemolytic activity is an important virulence factor of C. albicans. Moreover, biofilm production, as another virulence factor, plays a major role in the pathogenesis of Candida species33. Gupta AK, Daigle D, Carviel JL. The role of biofilms in onychomycosis. J Am Acad Dermatol. 2016;74(6):1241-6.. Biofilm formation on the nail bed shows that C. albicans uses the nail as a substrate for producing thick biomasses of biofilm1010. Vila TVM, Rozental S, de Sá Guimarães CMD. A new model of in vitro fungal biofilms formed on human nail fragments allows reliable testing of laser and light therapies against onychomycosis. Lasers Med Sci. 2015;30(3):1031-9..
In the present study, 23 isolates (32.85%) from 70 C. albicans isolates from patients with onychomycosis showed the ability to form biofilms at weak, moderate, and strong levels in vitro. Other researchers have reported the ability to form biofilms in 42.9%3333. Girish Kumar C, Menon T. Biofilm production by clinical isolates of Candida species. Sabouraudia. 2006;44(1):99-101., 40.3%3434. Pannanusorn S, Fernandez V, Römling U. Prevalence of biofilm formation in clinical isolates of Candida species causing bloodstream infection. Mycoses. 2013;56(3):264-72., and 33.3%2121. Aggarwal P, Kashyap B. Biofilm production by clinically isolated Candida: Comparative analysis based on specimen, methodology, and various Candida species. Indian J Med Speciali. 2018;9(2):69-72. of C. albicans and indicated the relationship between high biofilm production and pathogenesis. Furthermore, multiple studies have shown that the rate of proteinase, phospholipase, haemolysin, and biofilm production is variable, depending on the location of infection and number of Candida isolates. Analysis of the effect of extracellular enzyme activity on biofilm production showed a significant correlation between phospholipase activity and biofilm production in C. albicans isolates (P< 0.05).
Similar results have been reported by El-Houssaini et al. from Egypt2828. El-Houssaini HH, Elnabawy OM, Nasser HA, Elkhatib WF. Correlation between antifungal resistance and virulence factors in Candida albicans recovered from vaginal specimens. Microb Pathog . 2019;128:13-9. and Deorukhkar et al. from India3535. Sachin D, Santosh S. Evaluation of phospholipase activity in biofilm forming Candida species isolated from intensive care unit patients. Br Microbiol Res J. 2013;3(3):440-7.. They revealed a significant positive correlation between biofilm formation and phospholipase production. Some of the proposed hypotheses in this area suggest that the importance of biofilm formation by Candida species is related to the reduction of azole drug penetration into the biofilm matrix, reduced metabolism of biofilm cells, and increased expression of gene resistance, associated with antifungal resistance3636. Vediyappan G, Rossignol T, d’Enfert C. Interaction of Candida albicans biofilms with antifungals: transcriptional response and binding of antifungals to beta-glucans. . J Antimicrob Agents chemother . 2010;54(5):2096-111.,3737. Baillie GS, Douglas LJ. Effect of Growth Rate on Resistance of Candida albicans Biofilms to Antifungal Agents. J Antimicrob Agents chemother . 1998;42(8):1900-5..
The present results showed a significant positive correlation of FLC/ITC MICs with biofilm formation and phospholipase production in C. albicans in vitro. Furthermore, there was a significant association between FLC MIC and haemolysin production. Therefore, biofilm formation can play a significant role in the increased azole resistance of C. albicans isolates from clinical specimens.
The current study confirmed the effect of extracellular enzymes on biofilm production and determined the relationship between virulence factors and antifungal resistance in C. albicans isolates from patients with clinical onychomycosis in vitro. These results can contribute to a better understanding of biofilm formation and extracellular enzymes as potential targets against antifungal resistance. Further studies on genetic and molecular levels are required to better understand the association between virulence factors and antifungal resistance of C. albicans clinical strains. Moreover, the relationship between virulence factors and antifungal resistance may highlight new therapeutic strategies, based on the involvement of the virulence mechanism in the effectiveness of treatment.
The code of ethics for this study is IR.QUMS.REC.1396.283.
ACKNOWLEDGEMENTS
The authors would like to thank all the patients for their kind cooperation.
REFERENCES
-
1Maraki S, Mavromanolaki VE. Epidemiology of onychomycosis in Crete, Greece: a 12-year study. Mycoses. 2016;59(12):798-802.
-
2Otašević S, Barac A, Pekmezovic M, Tasic S, Ignjatović A, Momčilović S, et al. The prevalence of Candida onychomycosis in Southeastern Serbia from 2011 to 2015. Mycoses. 2016;59(3):167-72.
-
3Gupta AK, Daigle D, Carviel JL. The role of biofilms in onychomycosis. J Am Acad Dermatol. 2016;74(6):1241-6.
-
4Naglik J, Albrecht A, Bader O, Hube B. Candida albicans proteinases and host/pathogen interactions. J Cell Microbiol. 2004;6(10):915-26.
-
5Schaller M, Borelli C, Korting HC, Hube B. Hydrolytic enzymes as virulence factors of Candida albicans Mycoses. 2005;48(6):365-77.
-
6Sacristan B, Blanco M, Galan-Ladero M, Blanco J, Perez-Giraldo C, Gomez-Garcia A. Aspartyl proteinase, phospholipase, hemolytic activities and biofilm production of Candida albicans isolated from bronchial aspirates of ICU patients. J Med mycol. 2011;49(1):94-7.
-
7Sav H, Rafati H, Öz Y, Dalyan-Cilo B, Ener B, Mohammadi F, et al. Biofilm formation and resistance to fungicides in clinically relevant members of the fungal genus fusarium. J Fungi. 2018;4(1):16.
-
8Mukherjee PK, Chandra J, Kuhn DM, Ghannoum MA. Mechanism of fluconazole resistance in Candida albicans biofilms: phase-specific role of efflux pumps and membrane sterols. J Infect immun. 2003;71(8):4333-40.
-
9Seidler MJ, Salvenmoser S, Müller F-MC. Aspergillus fumigatus forms biofilms with reduced antifungal drug susceptibility on bronchial epithelial cells. J Antimicrob Agents chemother. 2008;52(11):4130-6.
-
10Vila TVM, Rozental S, de Sá Guimarães CMD. A new model of in vitro fungal biofilms formed on human nail fragments allows reliable testing of laser and light therapies against onychomycosis. Lasers Med Sci. 2015;30(3):1031-9.
-
11Mackenzie D. Serum tube identification of Candida albicans J Clin Pathol. 1962;15(6):563-5.
-
12Al-Wasify RS. Sensitivity and specificity of chromogenic media for detection of some pathogens in water. Int J Environ Sustain. 2013;2(1).
-
13Yamada Y, Makimura K, Merhendi H, Ueda K, Nishiyama Y, Yamaguchi H, et al. Comparison of different methods for extraction of mitochondrial DNA from human pathogenic yeasts. Jpn J Infect Dis. 2002;55(4):122-5.
-
14Romeo O, Criseo G. First molecular method for discriminating between Candida africana, Candida albicans, and Candida dubliniensis by using hwp1 gene. Diagn Microbiol Infect Dis. 2008;62(2):230-3.
-
15Clinical and Laboratory Standards Institute (CLSI). Reference method for Broth Dilution antifungal susceptibility testing of yeasts; approved standard -third edition. Document M27-A3.Wayne, PA: CLSI; 2008. 13p.
-
16Chakrabarti A, Nayak N, Talwar P. In vitro proteinase production by Candida species. Mycopathol. 1991;114(3):163-8.
-
17Gokce G, Cerikcioglu N, Yagci A. Acid proteinase, phospholipase, and biofilm production of Candida species isolated from blood cultures. Mycopathol. 2007;164(6):265.
-
18Price MF, Wilkinson ID, Gentry LO. Plate method for detection of phospholipase activity in Candida albicans Sabouraudia: J Med Vet Mycol. 1982;20(1):7-14.
-
19Manns JM, Mosser DM, Buckley HR. Production of a hemolytic factor by Candida albicans Infect Immun. 1994;62(11):5154-6.
-
20Chatzimoschou A, Katragkou Α, Simitsopoulou M, Antachopoulos C, Georgiadou E, Walsh TJ, et al. Activities of triazole-echinocandin combinations against Candida species in biofilms and as planktonic cells. J Antimicrob Agents chemother . 2011;55(5):1968-74.
-
21Aggarwal P, Kashyap B. Biofilm production by clinically isolated Candida: Comparative analysis based on specimen, methodology, and various Candida species. Indian J Med Speciali. 2018;9(2):69-72.
-
22Stepanović S, Vuković D, Dakić I, Savić B, Švabić-Vlahović M. A modified microtiter-plate test for quantification of staphylococcal biofilm formation. J Microbiol Methods. 2000;40(2):175-9.
-
23Brilhante RSN, Bittencourt PV, de Souza Collares Castelo-Branco D, de Melo Guedes GM, de Oliveira JS, Alencar LP, et al. Biofilms of Candida spp. from the ocular conjunctiva of horses with reduced azole susceptibility: a complicating factor for the treatment of keratomycosis? Vet Ophthalmol. 2017;20(6):539-46.
-
24Silva-Dias A, Miranda IM, Branco J, Monteiro-Soares M, Pina-Vaz C, Rodrigues AG. Adhesion, biofilm formation, cell surface hydrophobicity, and antifungal planktonic susceptibility: relationship among Candida spp. Front microbiol. 2015;6:205.
-
25Sav H, Baris A, Turan D, Altinbas R, Sen S. The frequency, antifungal susceptibility and enzymatic profiles of Candida species in cases of onychomycosis infection. Microb Pathog. 2018;116:257-62.
-
26Pakshir K, Zomorodian K, Zakaei A, Motamedi M, Ghiasi MR, Karamitalab M. Molecular identification and in-vitro antifungal susceptibility testing of Candida species isolated from patients with onychomycosis. Curr med mycol. 2015;1(4):26.
-
27Khosravi A, Shokri H, Mansouri P, Katiraee F, Ziglari T. Candida species isolated from nails and their in vitro susceptibility to antifungal drugs in the department of Dermatology (University of Tehran, Iran). J Mycol Med. 2008;18(4):210-5.
-
28El-Houssaini HH, Elnabawy OM, Nasser HA, Elkhatib WF. Correlation between antifungal resistance and virulence factors in Candida albicans recovered from vaginal specimens. Microb Pathog . 2019;128:13-9.
-
29Sachin C, Ruchi K, Santosh S. In vitro evaluation of proteinase, phospholipase and haemolysin activities of Candida species isolated from clinical specimens. Inter j Med Biomed res. 2012;1(2):153-7.
-
30Mattei AS, Alves SH, Severo CB, Guazzelli LdS, Oliveira FdM, Severo LC. Determination of germ tube, phospholipase, and proteinase production by bloodstream isolates of Candida albicans Rev Soc Bras Med Trop. 2013;46(3):340-2.
-
31de Souza Ramos L, Barbedo LS, Braga-Silva LA, Dos Santos ALS, Pinto MR, da Graça Sgarbi DB. Protease and phospholipase activities of Candida spp. isolated from cutaneous candidiasis. Rev Iberoam Micol. 2015;32(2):122-5.
-
32Pakshir K, Zomorodian K, Karamitalab M, Jafari M, Taraz H, Ebrahimi H. Phospholipase, esterase and hemolytic activities of Candida spp. isolated from onychomycosis and oral lichen planus lesions. J Mycol Med . 2013;23(2):113-8.
-
33Girish Kumar C, Menon T. Biofilm production by clinical isolates of Candida species. Sabouraudia. 2006;44(1):99-101.
-
34Pannanusorn S, Fernandez V, Römling U. Prevalence of biofilm formation in clinical isolates of Candida species causing bloodstream infection. Mycoses. 2013;56(3):264-72.
-
35Sachin D, Santosh S. Evaluation of phospholipase activity in biofilm forming Candida species isolated from intensive care unit patients. Br Microbiol Res J. 2013;3(3):440-7.
-
36Vediyappan G, Rossignol T, d’Enfert C. Interaction of Candida albicans biofilms with antifungals: transcriptional response and binding of antifungals to beta-glucans. . J Antimicrob Agents chemother . 2010;54(5):2096-111.
-
37Baillie GS, Douglas LJ. Effect of Growth Rate on Resistance of Candida albicans Biofilms to Antifungal Agents. J Antimicrob Agents chemother . 1998;42(8):1900-5.
-
Financial support: This research was financially supported by the Research Deputy of Qazvin University of Medical Sciences (grant No. 14814).
Publication Dates
-
Publication in this collection
07 Feb 2020 -
Date of issue
2020
History
-
Received
13 May 2019 -
Accepted
17 Dec 2019