Abstracts
A reverse transcriptase polymerase chain reaction (RT-PCR) to detect mouse hepatitis virus (MHV) in hepatic tissue was developed. To circumvent possible failures in RT-PCR amplifications, a second round of PCR with internal primers was used to confirm the specificity and increase the sensitivity of the test. Using this method specific amplification of MHV sequences was observed in 18 out of 20 mouse colonies examined.
Mouse hepatitis virus; RT-PCR
Desenvolveu-se a técnica de transcrição reversa - reação em cadeia pela polimerase para detectar o vírus da hepatite do camundongo em tecido hepático. Para se eliminar possíveis falhas na reação de amplificação RT-PCR usou-se um segundo ciclo de amplificação com primers internos para confirmar a especificidade e aumentar a sensibilidade do teste. Após a optimização do protocolo, descreve-se a detecção da amplificação específica da seqüência-alvo em camundongos de 18 das 20 colônias examinadas.
Vírus da hepatite murina; RT-PCR
Detection of mouse hepatitis virus in mouse colonies using the nested polymerase chain reaction
[Detecção do vírus da hepatite murina utilizando-se a reação em cadeia pela polimerase]
A.B. Cecílio, A.L. Cândido, M. Resende*, E.D. Bontempo, A.S. Martins
Departamento de Microbiologia - Instituto de Ciências Biológicas UFMG
Caixa Postal 486
30161-970 - Belo Horizonte, MG
*Autor para correspondência
E-mail: mresende@mono.icb.ufmg.br
ABSTRACT
A reverse transcriptase polymerase chain reaction (RT-PCR) to detect mouse hepatitis virus (MHV) in hepatic tissue was developed. To circumvent possible failures in RT-PCR amplifications, a second round of PCR with internal primers was used to confirm the specificity and increase the sensitivity of the test. Using this method specific amplification of MHV sequences was observed in 18 out of 20 mouse colonies examined.
Keywords: Mouse hepatitis virus, RT-PCR
RESUMO
Desenvolveu-se a técnica de transcrição reversa - reação em cadeia pela polimerase para detectar o vírus da hepatite do camundongo em tecido hepático. Para se eliminar possíveis falhas na reação de amplificação RT-PCR usou-se um segundo ciclo de amplificação com primers internos para confirmar a especificidade e aumentar a sensibilidade do teste. Após a optimização do protocolo, descreve-se a detecção da amplificação específica da seqüência-alvo em camundongos de 18 das 20 colônias examinadas.
Palavras-Chave: Vírus da hepatite murina, RT-PCR
INTRODUCTION
Mouse hepatitis virus (MHV) is an enveloped virus which has a 31 Kb single-strand positive RNA genome (Lai, 1990; Spaan et al., 1990). MHV belongs to the Coronaviridae family and replicates in the cytoplasm of infected cells using a viral RNA-dependent RNA polymerase which is translated from the genomic RNA (Adami et al., 1995).
As with other members of the coronavirus family, MHV infection varies in relation to tissue tropism and pathogenic potential. MHV strains are classified as respiratory tropic or enterotropic groups based on tissue distribution of primary infection although the patterns of infection can vary (Compton et al., 1993). Strains of MHV vary considerably in virulence, however most of them cause brief non apparent infections in mice and expression of the disease is influenced by host age, genotype and immune status (Barthold & Smith, 1990).
MHV is well known to be the most common virus of laboratory mice (Kagiyama et al., 1986). Natural infections with MHV remain widespread in most laboratory mouse populations despite the efforts to detect and eradicate this agent. Current data based on serological tests estimate that 60 to 80% of laboratory animal colonies are infected with MHV (Parker, 1979).
Since its first description by Cheever in the late 1940s (Cheever et al., 1949), MHV has been shown to alter the results of in vivo experiments using other infectious and non-infectious agents (Boorman et al., 1982; Barthold & Smith, 1990). Concomitant infection with MHV has been correlated with altered responses to tumours (Akimaru et al., 1981) and to other viruses (Smith et al., 1987; Smith et al., 1991). Also immune system-modulation experiments were noted to potentiate MHV infection and disease (Boorman et al., 1982; Barthold & Smith, 1990). All these data show that inapparent MHV infection can lead to important influences on biomedical research.
MHV is able to spread rapidly in mouse colonies because of its high contagiousness (Yamada et al., 1993). Therefore an early detection of MHV infection is very important. Current methods used to detect MHV infection include enzyme-linked immunosorbent assays (ELISA) and immunofluorescent techniques. The diagnosis of MHV infection is mainly performed by serologic assays because of the difficulties in finding recognized histological lesions and in isolating the virus in tissue culture (van Herk et al., 1993; Percy & Barthold, 1993; FELASA, 1994). However, the seroconversion of the animal sentinels or the newly infected ones requires a waiting period before a serologic assay can be used. The direct detection of viral nucleic acid using molecular bio1ogy methods in clinical or necropsy specimens would be a quick and powerful means to detect an outbreak or sub-clinical conditions affecting the animals.
Reverse transcription and polymerase chain reaction (RT-PCR) have been used to detect MHV in mouse colonies and tissue culture (Homberger et al., 1991; Yamada et al., 1993). The aim of this study was to develop a RT-PCR assay sensitive enough to search for MHV RNA in naturally infected mice.
MATERIALS AND METHODS
Twenty animals showing no classical MHV signs originated from 20 different research colonies having various sanitary status were used for this study. The mice were ether anesthetised and killed by cervical dislocation according to recommendations of the Animal Welfare Act (Holden, 1986). Livers were taken out as soon as arrival in the laboratory premises in order to avoid virus cross contamination.
The MHV-3 strain (kindly provided by Dr. Rovilson Gilioli, CEMIB, University of Campinas, Sao Paulo, Brazil) was propagated on NCTC 929 cells grown in DMEM (Life Tecnologies, Inc.) supplemented with 10% fetal calf serum, and kept under 5% CO2 at 37ºC. Titers of virus stocks were determined by plaque assay on NCTC 929 cells as described previously (Taguchi et al., 1980). Cells were infected at a multiplicity of one. Under these condictions, the synsytia formation reached 50% on the third day post infection. The harvested culture media showed titers of 103 to 104 TCID50/ml.
Cells of cultures showing a synsytia formation of about 50% were lysed culture frasks (1.5´106 cells in one ml lysis buffer containing 10mM Tris-HCl pH 7.5, 10mM EDTA, 40mM dithiothreitol, 1% SDS and 0.2 mg/ml proteinase K) prior to extraction of viral RNA. The viruses in the clarified culture supernatant were collected using 40% saturation of NH4SO4 (AmS). AmS was s1ow1y added to the mixture at 4ºC. Precipitation was extended overnight at 4ºC. Suspensions containing the virus were centrifuged at 2000´g for 20 min at 4ºC and the sediment was dissolved in ultrapure water, dialysed against TEN (10mM Tris-HCl, pH 8.0, 0.1mM EDTA and 10mM NaCl) overnight and kept at 70ºC until needed.
RNA from viruses produced in cell culture and concentrated with AmS or from lysed cell culture was extracted using a method based on the protocol described by Chomczynski & Sacchi (1987). One ml of concentrated virus or of lysed infected cells was treated in 4ml of a guanidine isothiocyanate solution containing 25mM sodium citrate and 0.5% sarkosyl (lysis solution). Subsequently 0.1 volume 2M sodium acetate pH 4.0 was added. The lysate was thoroughly mixed with 0.2 volume chloroform/isoamyl alcohool (49:1), vortexed for 10 sec, incubated on ice for 15 min, and centrifuged at 10000 g for 20 min at 4ºC. The upper aqueous layer was transferred to a fresh tubes and an equal volume of isopropanol was added. The mixture was centrifuged at 10000´g to pellet the RNA which was ressuspended in 30ml 0.1% (DEPEC-water) and stored at 20ºC until use.
From clinical samples 100mg of hepatic tissue was homogenized in the presence of 1ml lysis solution using a tissue homogenizer (Turrax). Samples were centrifuged at 15000´g for 20 min, and RNA was harvested under the same conditions described above.
The RNA was quantified by optical density measurements (Gene Quant, Amersham Pharmacia Biotech).
The oligonucleotide primers were designed for cDNA synthesis and PCR amplification of the N gene based on the published sequences of MHV-A59 (Armstrong et al., 1983) and MHV-JHM strains (Skinner & Siddel, 1983). Primers LVC 08 (dCACATTAGAGTCATCTTCAT) and LVC 09 (dGAAGTAGAGATAATGTAAGCGT) were designed and used in RT according to Kunita et al. (1992). Primers LVC 28 (dACGCTTACATTATCAACTTC) and LVC 29 (dGATCTAAATTAGAATTGGTC) were designed considering the conserved regions of the N gene of MHV. The predicted fragment sizes from amplifications with primers LVC 08 and LVC 09 is a 147bp fragment, 256bp with primers LVC 28 and LVC 29 and 383bp with primers LVC 08 and 29.
Ten ml of the total RNA extraction mixture were reverse-transcribed in the presence of 200U murine Moloney transcriptase (Life-Technologies, Inc.), 20U Rnasin (Promega Corp.), 50pM primer complementary to the positive strand (LVC 09 or LVC 29), 2mM dithiotreitol and 200mM each dNTP. The reaction was carried out in a final volume of 20ml at 42ºC for 60 min.
The non nested PCR was performed in a total volume of 100ml containing 10ml of the cDNA transcript, 2U Taq DNA polymerase, 0.2mM each dNTP, 0.5mM each primer, 10ml 10´ PCR buffer (500mM KCl, 100mM Tris-HCl, pH 8.8, 15mM MgCl2 and 0.1% gelatin). For the primers pair LVC 08 and 09 conditions previously described were used (Kunita et al.,1992). For the second (LVC 28-29) and the third (LVC 08-29) set of primers two rounds of PCR cycle reactions were performed. The first round was carried out using five cycles of 94ºC, 50ºC and 72ºC for 60 sec each and a second round of 30 cycles of 90ºC, 50ºC and 72ºC also for 60 sec each. For the nested reaction two microliters of the amplification reaction with primers LVC 08-29 were used in another cycle of amplification using the internal primers LVC 08-09 or LVC 28-29 under the same conditions and cycle parameters as described above for the specific primers.
The sensitivity of the PCR assay was determined using as the template MHV-3 RNA extracted from virus produced in tissue culture and purified from the cell supernatant. Tenfold serial dilutions (5ng to 0.05fg) of the target RNA were done in sterile DEPC-water and 5.0ml of each dilution were used in the RT-PCR amplification assay described above with primers LVC 08 and 09.
One tenth (10ml) of each reaction volume was analysed in a 2% agarose gel in TBE buffer (Tris-borate 22mM, 0.5mM EDTA pH 8.0). The DNA molecular weight marker used was a 100bp ladder3. The gels were stained with ethidium bromide (0.5 mg/ml) and bands were visua1ized and photographed under UV light.
RESULTS
Cells infected with MHV and showing approximately 50% cytopathic effect were used as indicated before. The analysis of the amplification products in agarose gels indicated the presence of products of 147bp with primers LVC 08 and 09, 257bp with primers LVC 28 and 29 and 383bp with primers LVC 08 and 29. These products were clearly visualized as a sharp band on agarose gels (Fig. 1). At an annealing temperature of 50ºC all three sets of primers showed a band of the expected size. The negative controls BHV-1 BH83 strain and DNA of NTCT 929 cells yielded no visible PCR product. In Fig. 2 the results are shown for RT-PCR using MHV-3/CEMIB cDNA with nested primers LVC 08 and 09 (147bp fragment) and LVC 28-29 (257bp fragment).
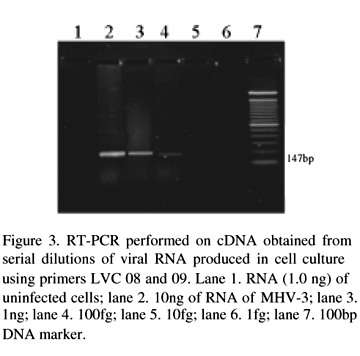
To assess the sensitivity of the RT-PCR the fragment of 147bp was amplified in serially RNA extracted from viruses purified from cell supernatant. Specific bands were visible in samples containing 100 fg of RNA in a ethidium bromide stained gel (Fig. 3).
The expected 383bp fragment from amplification with primers LVC 08-29 was detected as a faint band in mice of 18 out of 20 colonies examined (fig. not shown). The use of a second round of PCR with nested primers pair LVC 08 and 09 (results of nine colonies are shown in Fig. 4) or with primers LVC 28 and 29 increased the sensitivity up to 100-fold. All 18 PCR positive mice (from 18 different colonies) were healthy and had no anamnestic report of mild clinical disease that could be tracked to persistent MHV infection. The characteristic band of 383bp was not observed when RNA from mice of two colonies was amplified. In addition, reamplification with nested primers failed to detect MHV RNA in these two samples.
DISCUSSION
The increasing use of PCR methods for diagnostic purpose led us to develop a nested RT-PCR assay for detection of MHV in persistent infected murine colonies. Thus, one of the major aims of this study was to apply RT-PCR to detect natural infections with MHV in mice colonies and to determine the sensitivity and specificity of the assay. Using known amounts of purified viral RNA as little as 100fg of genomic RNA could be detected by RT-PCR. An additional nested RT-PCR increased 100-fold the sensitivity of MHV detection of MHV RNA extracted from in vitro infected cells up to and possibly less than 0.1 plaque forming unit of infected cell lysate. This RT-PCR protocol can be a sensitive alternative to viral isolation. Most diagnostic criteria for MHV isolation have emphasized hepatical and intestinal lesions (Ishida et al., 1978; Hierholzer et al., 1979) and that was the reason for using hepatic tissue. It is interesting to note that usually mice accepted for scientific research and for other kind of studies, although infected with MHV, were serologically negative and usually failed to show significant histopathologic alterations (Boorman et al., 1982).
Strain differences within MHV isolates such as base substitution at primer sites may lead to absence of amplification. To circumvent this possibility primers for amplification of the N gene have been used in this experiment. They were chosen from the highly conserved region near the 3end. The N gene code for a basic phosphoprotein that is associated with the viral genome. In pairwise comparisons of published data, the sequences of five strains were at least 91% identical and the differences which occurred in this region were at 12nt in strain MHV-A59, 26nt in MHV-JHM, 38nt inmMV-2, 9nt in MHV-5 and 13nt in MHV-Nu67 (Parker & Masters, 1990). Using the optimized protocol described here, specific amplification of the target sequence was confirmed in 18 mice out 20 colonies examined. This suggests that MHV is present and may circulate undetected in many Brazilian mouse colonies. Subclinical or latent infection could be a problem for the scientific investigator utilizing mice. This is especially true for research involving the mouse immune system (Virelizier et al., 1976). The RT-PCR assay described here confirmed the presence of MHV infection in colonies of different sanitary status, ranging from germ-free to conventional animals.
Finally, the nested RT-PCR principle described in this paper will be a powerful tool for the detection of MHV in mouse colonies. In conclusion the test could prove useful in monitoring free colonies with increased risk of contamination because of the circulation of wild-free mice in the area. In addition, the test showed excellent specificity and sensitivity and is considerably less time-consuming than other methods described so far for detecting MHV such as virus isolation, the infant mouse bioassay, and the mouse antibody production test.
ACKNOWLEGMENTS
This work was supported by grants of PADCT II, CNPq and FAPEMIG. We wish to thank Marília N. dos Santos and Tânia M. G. de Pinho for their technical assistance.
- ADAMI, C., POOLEY, J., GLOMB, J. et al. Evolution of mouse hepatitis virus (MHV) during chronic infection: quasispecies nature of the persisting MHV RNA. Virology, v.209, p.337-346, 1995.
- AKIMARU, K., STUHLMILLER, G.M., SEIGLER, H.F. Influence of mouse hepatitis virus on the growth of human melanoma in the peritoneal cavity of the athymic mice. J. Surg. Oncol., v.17, p.327-339, 1981.
- ARMSTRONG, J., SMEEKENS, S., ROTTIER, P. Sequence of the nucleocapsid gene from murine coronavirus MHV-A59. Nuc. Ac. Res., v.11, p.883-891, 1983.
- BARTHOLD, S.W., SMITH, A.L. Duration of mouse hepatitis virus infection: studies in immunocompetent and chemically immunosuppressed mice. Lab. Anim. Sci., v.40, p.133-137, 1990.
- BOORMAN, G.A., LUSTER M.I., CAMPBELL, M.L. et al. Peritoneal macrophage alterations caused by naturally occuring mouse hepatitis virus. Am. J. Pathol., v.106, p.110-117, 1982.
- CHEEVER F.S., DANIELS J.B., PAPPENHEIMER, A.M. A murine virus (JHM) causing disseminated encephalomyelitis with extensive destruction of the myelin: I. Isolation and biological properties of the virus. J. Exp. Med., v.90, p.181-194, 1949.
- CHOMCZYNSKI. P., SACCHI, N. Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Ana. Biochem., v.162, p.156-159.1987.
- COMPTON S.R., BARTHOLD, S.W., SMITH, A.L. The cellular and molecular pathogenesis of coronaviruses. Lab. Anim. Sci., v.43, p.15-28, 1993.
- FELASA. Recommendations for the health monitoring of mouse, rat, hamster, guineapig and rabbit breading colonies: Report of the FELASA working group. Anim. Health Lab. Anim, v.28, p.1-12, 1994.
- HIERHOLZER, J.C., BRODERSON, J.R., MURPHY, F.A. New strain of mouse hepatitis virus as the cause of lethal enteritis in infant mice. Infect. Immunol, v.24, p.508-522, 1979.
- HOLDEN C. New animal regulations causing scientists pain and distress. Science, v.233, p.619-626, 1986.
- HOMBERGER, F.R.A., SMITH. A.L., BARTHOLD, S.W. Detection of rodent coronaviruses in tissues and cell cultures using the polymerase chain reaction. J. CIin. Microbiol., v.29, p.2789-2793, 1991.
- ISHIDA, T., TAGUCHI, F., LEE, Y.S. et al. Isolation of mouse hepatitis virus from infant mice of fatal diarrhea. Lab. Anim. Sci., v.28, p.269-278, 1978.
- KAGIYAMA, N., TAKAKURA, A., ITOH, T. A serological survey on 15 murine pathogens in mice and rats. Exp. Anim., v.35, p. 531-536, 1986.
- KUNITA, S., TERADA, E., GOTO, K. et al. Sequence analysis and molecular detection of mouse hepatitis virus using the polymerase chain reaction. Lab. Anim. Sci, v.42, p.593-598, 1992.
- LAI, M.M.C. Coronavirus: Organization, replication and expression of genome . Annu. Vet. Microbiol, v.44, p.303-333, l 990.
- PARKER, J.C. The possibilities and limitations of virus control in laboratory animals. In: 7th ICLAS Symposium, Utrecht, Animal Quality and Models in Biomedical Research Stuttgard: Gustav Verlag, 1979. p.161-172.
- PARKER, M.N., MASTERS, P.S. Sequence comparison of the N genes of five strains of the coronavirus mouse hepatitis suggests a three-domain structure for the nucleocapsid protein. Virology, v.179, p.463-468, 1990.
- PERCY, D.H., BARTHOLD, S.W.(Ed.) Pathology of laboratory rodents and rabbits Ames: Iowa State Untversity Press, 1993. p.20-23.
- SKINNER, M.A., SIDDEL, S.G. Coronavirus JHM: nucleotide sequence of the mRNA that encodes nucleocapsid protein. Nucl. Ac. Res., v.11, p.5045-5054, 1983.
- SMITH, A.L., BOTTOMLY, K., WINOGRAD, D.F. Altered splenic T cell function of BALB/cByJ mice infected with mouse hepatitis virus or Sendai virus. J. Immunol, v.138, p.3426-3430, 1987.
- SMITH, A.L., WINOGRAD, D.F., DE SOUZA, M.S. In vitro splenic T cell responses of diverse mouse genotypes after oronasal exposure to mouse hepatitis virus, strain JHM. Lab. Anim. Sci, v.41, p.106-111, 1991.
- SPAAN, W., CAVANAGH, D., HORZINEK, M.C. The basis for serodiagnosis and vaccines. In: VAN REGENMORTEL, NEURATH, A.R. (Ed.) Immunochemistry of viruses. New York: Elsevier, 1990. p.359-379.
- TAGUCHI, F., YAMADA, A., FUJIWARA, K. Resistance to highly virulent mouse hepatitis virus acquired by mice after low-virulence infection: Enhanced antiviral activity of macrophages. Infect. Immunol., v.29, p.42-49, 1980.
- Van HERK, W.A., MULLINK, J.W.M.A., BOSLAND, M.C. Principles of laboratory animal science: a contribution to the humane use and care of animals and to the quality of experimental results. In: Van ZUTPMEN, L.F.M., BAUMANS, V., BEYNEN, A.C. (Ed.) Diseases in laboratory animals. Amsterdam: Elsevier, 1993. p.167-188.
- VIRELIZIER, J.L., VIRELIZIER, E.M., ALLISON, A.C. The role of circulating interferon in the modification of immune responsiveness by mouse hepatitis virus (MHV-3). J. Immunol., v.117, p.748-753, 1976.
- YAMADA, Y.K., YABE, M., YAMADA, A. et al. Detection of mouse hepatitis virus by polimerase chain reaction and its application to the rapid diagnosis of infection. Lab. Anim. Sci, v.43, p.285-290, 1993.
Publication Dates
-
Publication in this collection
21 Sept 2000 -
Date of issue
Aug 2000
History
-
Received
25 Feb 2000