Abstracts
The purpose of this review was to carry out an analysis of the liver regenerative process focusing on the molecular interactions involved in this process. The authors undertook a review of scientific publications with a focus on the liver regeneration.The cellular processes involved in liver regeneration require multiple systematic actions related to cytokines and growth factors. These interactions result in the initiation of mitogenic potential of the hepatocytes. The action of these modulators in the regenerative process require a processing in the extra-cellular matrix. Serines and metal proteins are responsible for the bio availability of cytokines and growth factors so that they can interact as receptors in the cellular membrane generating signaling events for the beginning and end of the liver regenerative process. The exact mechanism of interaction between cells, cytokines and growth factors is not well established yet. A series of ordered events that result in the hepatic tissue regeneration has been described. The better understanding of these interactions should provide a new approach of the treatment for liver diseases, aiming at inducing the regenerative process.
Liver Regeneration; Cytokines; Grow Factors; Hepatocytes
O objetivo desta revisão foi desenvolver uma análise do processo regenerativo do fígado, focando as interações moleculares envolvidas neste processo.Os processos celulares envolvidos na regeneração hepática requerem múltiplas ações sistemáticas relacionadas com citoquinas e fatores de crescimento. Estas interações resultam na iniciação do potencial mitogênico dos hepatócitos. A ação destes moduladores do processo regenerativo necessita de processamento no meio extra celular. As serinas e metaloproteínas são responsáveis pela biodisponibilização de citoquinas e fatores de crescimento, para que então possam interagir com receptores na membrana celular gerando os eventos sinalizadores para o inicio e o término do processo regenerativo hepático.O exato mecanismo de interação entre células, citoquinas e fatores de crescimento não está bem estabelecido. Tem-se descrito uma série de eventos ordenados que resulta na regeneração do tecido hepático. O melhor entendimento destas interações leva a uma nova abordagem de tratamento para doenças hepáticas, objetivando a indução do processo regenerativo.
Regeneração Hepática; Citocinas; Fatores de Crescimento; Hepatócitos
REVIEW ARTICLE
A molecular view of liver regeneration1 1 Study performed in the Laboratory of Cellular Proliferation of the Department of Pathology of the Faculty of Medicine of Ribeirão Preto, University of São Paulo, (FMRPUSP), Brazil.
Uma visão molecular da regeneração hepática
Marissa Rabelo TarláI; Fernando Silva RamalhoII; Leandra Naira Zambelli RamalhoIII; Tiago Castro e SilvaIV; Daniel Ferracioli BrandãoI; Juliana FerreiraI; Orlando Castro e SilvaV; Sérgio ZucolotoVI
IFellow PhD degree of the Department of Pathology, (FMRP-USP), Brazil
IIPhD, Professor of the Department of Surgery and Anatomy, (FMRP-USP), Brazil
IIIPhD, Professor of the Department of Pathology, (FMRP-USP), Brazil
IVGraduate student of the Physical Institute of São Carlos University of São Paulo (IFSC-USP), Brazil
VFull Professor, Head of Division of Gastroenterology of the Department of Surgery and Anatomy, Coordinator of the Liver Transplant Program, (FMRP-USP), Brazil
VIFull Professor of the Department of Pathology, (FMRP-USP), Brazil
Correspondence Correspondence: Orlando de Castro e Silva Jr. Rua Campos Salles, 809 - 9º andar CEP: 14015-110 Centro, Ribeirão Preto - SP - Brazil. Email: orlando@fmrp.usp.br
ABSTRACT
The purpose of this review was to carry out an analysis of the liver regenerative process focusing on the molecular interactions involved in this process. The authors undertook a review of scientific publications with a focus on the liver regeneration.The cellular processes involved in liver regeneration require multiple systematic actions related to cytokines and growth factors. These interactions result in the initiation of mitogenic potential of the hepatocytes. The action of these modulators in the regenerative process require a processing in the extra-cellular matrix. Serines and metal proteins are responsible for the bio availability of cytokines and growth factors so that they can interact as receptors in the cellular membrane generating signaling events for the beginning and end of the liver regenerative process. The exact mechanism of interaction between cells, cytokines and growth factors is not well established yet. A series of ordered events that result in the hepatic tissue regeneration has been described. The better understanding of these interactions should provide a new approach of the treatment for liver diseases, aiming at inducing the regenerative process.
Key words: Liver Regeneration. Cytokines. Grow Factors. Hepatocytes.
RESUMO
O objetivo desta revisão foi desenvolver uma análise do processo regenerativo do fígado, focando as interações moleculares envolvidas neste processo.Os processos celulares envolvidos na regeneração hepática requerem múltiplas ações sistemáticas relacionadas com citoquinas e fatores de crescimento. Estas interações resultam na iniciação do potencial mitogênico dos hepatócitos. A ação destes moduladores do processo regenerativo necessita de processamento no meio extra celular. As serinas e metaloproteínas são responsáveis pela biodisponibilização de citoquinas e fatores de crescimento, para que então possam interagir com receptores na membrana celular gerando os eventos sinalizadores para o inicio e o término do processo regenerativo hepático.O exato mecanismo de interação entre células, citoquinas e fatores de crescimento não está bem estabelecido. Tem-se descrito uma série de eventos ordenados que resulta na regeneração do tecido hepático. O melhor entendimento destas interações leva a uma nova abordagem de tratamento para doenças hepáticas, objetivando a indução do processo regenerativo.
Descritores: Regeneração Hepática. Citocinas. Fatores de Crescimento. Hepatócitos.
Introduction
It is well-known that the liver is an organ of unequalled regenerative capacity. It is well-established that the hepatocytes are able to reconstruct the liver tissue after the occurrence of damages such as surgical and toxic lesions. The elucidation of the events that make up this restorative capacity becomes crucial for the understanding of the hepatocyte dynamics during the liver regeneration. The regenerative process requires a close interplay between the signals of the extracellular matrix and the intracellular means. A response from the remanescent cells, such as the genes transcription and the new protein codification, must be established early resulting in the beginning of the cellular proliferation. A series of events regulated by cytokines and growth factors. These regulators of the regenerative process require a processing in the extracellular matrix. Metalproteins process cytokines and factors associated with the regeneration, making them available so that they signal with the extracellular membrane, starting and finishing an intracellular event cascade that will result in the reestablishment of the liver tissue.
Regulating Factors in Hepatocyte Division
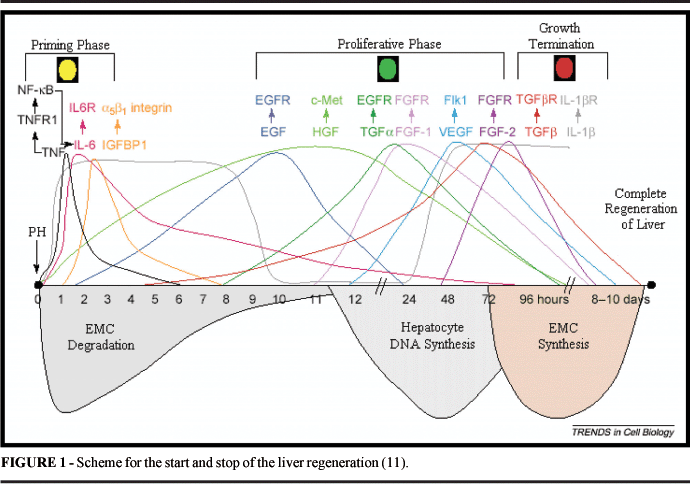
Liver regeneration is an event that involves multiple cellular processes and a complex interaction with cytokines and growth factors. Related factors that start the regeneration and make the regenerated liver functional, is a well-elucidated process with animal models after partial hepatectomy. More specifically, the liver regeneration may be divided into three phases: (1) Initiation; (2) Proliferation; and (3) Inhibition. The mitogenic response divides into two phases, first due to cytokines that stimulate the G0-G1 transition, and then the progression until phase S. In rats, after hepatectomy, hepatocytes go into G1 around 4 hours after the procedure and progress to G1-S with DNA synthesis peak within 24 hours 1. After hepatectomy, non-parenchimatous liver cells and remanescent hepatocytes increase the production of tumoral necrosis factor (TNF)-a, linfotoxine (LT)-b and interleukins (IL)-1 and 6. (TNF)-a bonds to the soluble receptor TNFR1, resulting in the translocation of the nuclear factor, Kappa-B (NF-kB). The transcription factor NF-kB controls the expression of the cytokine-codifying genes, regulate the cellular cycle and is an antagonist of the apoptosis in the liver regenerative process. NF-kB consists of different proteins with distinct biological activities. An activity form of NF-bK is found retained in the cytoplasm, and its activation occurs a few minutes after the HP process and lags for 1 to 2 hours 2. (TNF)-a also induces the expression of IL-6, which is a signal for the transduction and activation of (STAT) 3 transcription, possibly attaching to C/Erbb protein in the liver remanescent cells 3. STAT3 is important in the inhibition of hepatocyte apoptosis by inhibition of the caspasis 3 and 8 activation, in addition to improving the survival rate of hepatocytes through the inhibition of TGFâ mediated by Fas activation 4. TNFR1 or IL-6 deficiency diminishes the liver regeneration in mice 3. In tnf 0/0 mice, there is no significant blockage of regeneration, with the chance of initialization compensating factor occurence 5 As a result of the activation of these transcription factors (NF-kB, STAT3 and C/EBPb), the hepatocytes program the response of primary growth (immediate genes, activation of AP-1 and pro-oncogenesis: c-Myc, c-fos, c-Has, c-met, c-Erb 2. Nitric oxide takes part in these events through the generation of nitric oxide sintetase (NOS-2) that prevents the pro-apoptotic activation mediated by TNFa (via caspase 3), and protects the hepatocytes against cellular death mediated by procaspases S-nitrosilantes 6. In a second phase, the hepatocyte growth factor (HGF) and its receptor MET are considered the central stimulation for the G1-S progression in the remanescent hepatocytes 7. Hepatocytes are sensibilized by TNFa, IL-6 and nitric oxide (NO) in order to respond to stimulations of growth factors. HGF, a potent mitogenic agent, is a peptide secreted by non-parenchymatous cells after hepatectomy and induces early genes in the regeneration 2. It has been shown that HGF may be able to induce the differentiation of stem cells of the bone marrow in hepatocyte 8. The c-met deficiency, receptor HGF, results in the necrosis generation of hepatocytes and esteatosis 9. Other mitogens involved in the proliferative phase include the transforming growth factor (TGFa) and the epidermal growth factor (EGF). The expression of TGFa is induced in the hepatocytes around 2-3 hours after PH and suggests that it stimulates the proliferation of both hepatocytes and endothelial cells 10. TGFa and EGF signal to the cell through a receptor of tirosine kinase in common, a receptor EGFR. Endothelial cells can also be induced by the growth factor of the vascular endothelium (VEGF), angipoetinas 1 and 2 and growth factor of fibroblast (FGF) 11. The growth factor similar to insulin (IGF) is an auxiliary element that bonds to receptor IGFP-1 that is induced in hepatocytes by the stimulation of IL-6. The amino acids sequence RGD in IGFBP-1 can bonds to the fibronectine-integrine a5b1 receptor, affecting the migration and cellular adhesion. The IGFBP-1 deficienty results in defects in the DNA synthesis, diminishing of the expression of proteins that regulate the cellular cycle and the liver necrosis after hepatectomy, indicating a possible mitogenic role in the liver regeneration 12. The substance of hepatic stimulation (HSS), a peptidic growth factor that is specific in the liver, is related to the protection and proliferation of hepatocytes through the repairing of DNA synthesis and self-fosforilation of the residues of tiroxine of EGFR receptor 13. In the inhibition phase, the signals for the termination of growth or the stop signals (stop signals) are responsible for the decrease of DNA synthesis. The most known of these is the TGFb, secreted by hepatocytes and plaquetes 14. In mice, the TGFb increased expression results in the absence of regeneration and development of hepatocarcinoma 15. As a member of the superfamily TGF, it sgnals through TGFR receptor, antogonizes with TGF-a, a growth factor that competes for the same TGFR receptors. The control between these signals occurs by the earlier secretion of TGFa and later secrection of TGFb 16. Another mitosis inhibitor is IL-1 that expresses itself in the two phases of the regeneration, first as a proliferative stimulation and in the terminal phase of this process. The end of the hepatocyte regenerative process is not well defined, such as the initial and proliferative phases, possibly this latter initiates through the stimulation of hepatocyte growth factors 17
Proteolitic Modulation: Extra Cellular Matrix
The liver is made up by 60% of parechymatous cells (hepatocytes) and non-parenchymatous cells (Kupffer's cells, colangiocytes, epithelial, endothelial and oval cells). Basically, the liver parenchyme structure is organized into hexagonal lobes that contain a central vein surrounded by six portal spaces (Figure 2). The portal space consists of a vein, a hepatic artery and a biliar duct that drain spaces between the hepatocytes and is surrounded by endothelial cells termed sinusoides. The Kupffer's cells are found in the sinusoides and the stellat cells are located in the perisinusoidal spaces (Figure 3). The liver extra cellular matrix (MEC) works as a foundation for the cells and occupy less than 3% of the liver area. Collagen is the main protein of the extra cellular matrix, with collagen type I, III and V located in the portal area and central vein, and collagen type IV together with laminine and entactino-nidogen making up the basal membrane 18. Other molecules include glicoproteins (laminine, fibronectine, tenascine, nidogen and SPARC) and proteinoglicans (heparam, dermatam, condritine sulphate, perlacam, hialuronic acid and decorim). The MEC continuous network around the cells make it possible to store and transmit signals through bonds and liberations of growth factors, hormones, enzymes and cytokines. The process of restoration of the hepatic mass involves a series of proteolitic mechanisms located in MEC, which allows a rapid, but controlled response through the sending of signals for the initiation and stop of the regenerative process. Among these extra cellular factors, proteinases, serines and metalproteins (MMPs) exist as a complex proteolitic cascade that requires a specific mechanism of activation or as latent enzymes, inhibited by specific inhibitory proteases (TIMPs) and a 2 microglobulines 19. The proteolitic process is a mechanism to modify pre-existent signals without the need of a new synthesis. The proteolitic cleavage that occurs in the pericellular interface (MEC) may function both as an activator or inhibitor of the cellular signaling processes. It may acts directly or modifying the bonding proteins. Some important signaling pathways in the regeneration process have been clarified, recent studies have used mice deficient of tissue inhibitors of MMPs, type TIMP 1 or 3 20. The proteolitic regulation of the signalization through TNFa is well elucidated and it is known that the TNFa-converting enzyme, TACE/ADAN 17 can be inhibited by TIMP-3, increasing the TNFa action. TACE/ADAM 17 covers TNFR1/2 on the cellular surface, blocking the TNFa action21.
Another important element in the regenerative process, HGF, is secreted in the form of a pro-enzyme, Pro-HGF. This must be metabolized in MEC by MMPs, among them UPA (activator of plasminogen type uroquinase) 22. EGF and TNFa act through receptors of the ERBB family. EGF is synthesized as a pro-HB-EGF, has a bonding heparinoid, HB, and requires proteolitic cleavage in MEC so that the fosforilation tyrosine quinase of EGF receptor occurs 23. TNFa exists in the form of a transmebranic precursor that is discharged from the surface by proteolisis action. TACE/ADAN 17 participates in this process, being crucial for the TNFa activation 24. VEFG is composed of isoforms with heparinoid domains and HSPGs (proteinoglicans / heparam sulphates). MMP-9 and plasmine are capable to increase free VEFG levels, promoting induced angiogenesis during the regeneration process 25. FGF acts through a tirosine quinase recptor, the FGFR1-4. This receptor is of perlacam type, FGF activates these receptors through perlacam bonding elements, that limit the FGF diffusion, protecting it from the proteolitic action. MMP1 and 3 act by the liberation of FGF from the perlacam complex inhibiting the action of this element 26. In studies with hematopoetic stem cells, it has been shown that quimioquine SDF-1, a potent chemiotatic secreted by damaged hepatocytes and its receptor, the CXCR4, participates in the process of mobilization of stem cells of the bone marrow and the migration of these cells to the damaged liver tissue 27. In fibrosis and hepatic cirrhosis, there is the association with the increase of TIMPs expression and reduction of the expression of MMPs 28. The processes of chronic hepatitis are associated with metalproteinases 29. The extra cellular proteolisis may show an important role in the development of hepatocarcinomas. The identification and definition of the metaloproteinase function in the liver regenerative process must be a new strategy for the development of new clinical treatments of hepatic diseases.
Conflict of interest: none
Financial source: FAPESP, CNPq.
- 1. Fausto N. Liver regeneration. J Hepatol. 2002; 32 :19. 1477-87.
- 2. Kountouas J, Boura P,Lygidakis NJ. Liver regeneration after hepatectomy. Hepatogastroenterology. 2001; 48:556-62.
- 3. Pahl MK. Activators and target genes of REL/ nf- kappa B transcription factors. Oncogene. 1999; 18:6853-66.
- 4. Haga S, Terui K, Zhang H. Q. STA3 protects against Fas-induced liver injury by redox-dependent and independent mechanisms. J. Clin. Invest. 2003; 112:989-98.
- 5. Yamada Y, Kirillova I, Peschon JJ, Fausto N. Initiation of liver growth by tumor necrosis factor: deficient liver regeneration in mice lacking type I tumor necrosis receptor factor receptor. Proc. Natl. Acad. Sci. 1997; 94, 1441-6.
- 6. Zeini M, Hortelano S, Traves PG. Assessment of dual regulatory role of NO in liver regeneration after partial hepatectomy: Protection against apoptosis and retardation of hepatocytes proliferation. FASEB J. 2005; 19:995-7.
- 7. Michalopoulos GK, De Frances MC. Liver regeneration. Science. 1997; 276:60-6.
- 8. Kollet O, Shivtiel S, Chen YQ, Suriawinata J, Thung SN, Dabeva MD, Kahn L, Samira S, Dar A. HGF,SDF-1 and MMP-9 are involved in stress induced human CD34+ stem cell recruitment to the liver. J Clin. Invest. 2003; 112, 160-9.
- 9. Borowiak M, Garratt AN, Wustefeld T, Strehle M, Trautwein C, Birchmeier C. Met provides essential signals for liver regeneration. Proc Natl.Sci. 2004; 101:10608-13.
- 10. Webber EM, Wu JC, Wang L, Merlino G, Fausto N. Overexpression of transforming growth factor-alpha causes liver enlargement and increased hepatocytes proliferation in transgenic mice. Am. J. Pathol. 1994; 145:398-408.
- 11. Mohammed FF, Khokha R. Thinking outside the cell: proteases regulate hepatocyte divison. Trends in Cell Biol. 2005; 15:555-63.
- 12. Leu JI, Crissey MA, Craig LE, Taub R. Impaired hepatocytes DNA synthetic response posthepatectomy in insulin-like growth factor binding protein 1-deficient mice with defects in C/EBP beta and mitogen- activated protein kinase/extracellular signal-regulated kinase regulation. Mol Cell Biol. 2003; 23:1251-9.
- 13. An W, Liu XJ. Lei TG, Dai J, Du GG. Growth induction of hepatic stimulator substance in hepatocytes throught its regulation on EGF receptors. Cell Res. 1999; 9:37-42.
- 14. Nihikawa Y. Wang M. Carr BI. Changes in TGF-beta receptor of rat hepatocytes during primary culture and liver regeneration: Increased expression of TGF- beta receptors associated with increased sensitivity to TGF- beta mediated growth inhibition. J. Cell Physiol. 1998; 176: 612-7.
- 15. Bottinger EP, Factor VM, Tsang ML, Weatherbee JA, Kopp JB, Qian SW, Wakefield LM, Roberts AB, Thorgeirsson SS, Sporn MB. The recombinant proregion of transforming growth factor beta1 inibits active transforming growth factor beta 1 in transgenic mice. Proc Natl Acad Sci. 1996; 93: 5877-82.
- 16. Kobayashi T, Niimi S, Hashimoto O, Hayakawa T. Expression of inhibin betaA, betaB and follistatin mRNAs in carbon tetrachloride induced rat liver regeneration model. Biol Pharm Bull. 2000; 23 755-7.
- 17. Boulton R, Woodman A, Calnan D, Selden C, Tam F, Hodgson H. Nonparenchymal cells from regenerating rat liver generate interleukin-1 alpha and1 beta: a mechanism of negative regulation of hepatocytes proliferation. Hepatology. 1997; 26: 49-58.
- 18. Bedossa P, Paradis V. Liver extracellular matrix in helth and disease. J Pathol. 2003; 200: 504-15.
- 19. Rao J. Molecular mechanisms of glioma invasiveness: The role of proteases. Nat Rev Cancer. 2003; 3: 489-501.
- 20. Mohammed FF, Pennington CJ, Kassiri Z, Rubin JS, Soloway PD, Ruther U, Edwards DR, Khokha R. Metalloproteinases inhibitor TIMP-1 affects hepatocyte cell cycle via HGF activation in murine liver regeneration. Hepatology. 2005; 41:857-67.
- 21. Peschon JJ, Slack JL, Raddy P, Stocking KL, Sunnarborg SW, Lee DC, Russell WE, Castner DJ, Johnson RS, Fitzner JN, Boyce RW, Nelson N, Kozlosky CJ, Wolfson MF, Rauch CT, Cerreti DP, Paxton RJ, March CJ, Black RA. An essential role for ectodomain shedding in mammalian development. Science. 1998; 282: 1281-4.
- 22. Lee SL. Activation of hepatocyte growth factor and urokinase/plasminogen activator by matriptase, an epithelial membrane serine protease. J Biol Chem. 2000; 36: 720-5.
- 23. Kiso S, Kawata S, Tamura S, Inui Y, Yoshida Y, Saway Y, Umeki S, Ito N, Yamada A, Miyagawa J, Higashiyama S, Iwawaki T, Saito M, Taniguchi N, Matsuzawa Y, Kohno K. Liver regeneration in heparin-binding EGF-like growth factor transgenic mice after partial hepatectomy. Gastroenterology. 2003; 124: 701-7.
- 24. Lee DC, Sunnarborg SW, Hinkle CL, Myers TJ, Stevenson MY, Russell WE, Castner BJ, Gerhart MJ, Paxton RJ, Black RA, Chang A, Jackson LF. ACE/ADAM17 processing of EGFR ligands indicates a role as a physiological convertase. Ann NY Acad Sci. 2003; 995: 22-38.
- 25. LeCouter J, Moritz DR, Li B, Phillips GL, Liang XH, Gerber HP, Hillan KJ, Ferrara N. Angiogenesis-independent andothelial protection of liver: role of VEGFR-1. Science. 2003; 299: 890-3.
- 26. Ornitz DM. FGFs, heparan sulfate and FGFRs: complex interations essential for development. Bioessays. 2000; 22: 108-12.
- 27. Dalakas E, Newsome PN, Harrison DJ, Plevris JN. Hematopoietic stem cell trafficking in liver injury. FASEB J. 2005;19: 1225-31.
- 28. Arthur MJ. Fibrogenesis II. Metalloproteinases and their inhibitors in liver fibrosis. Am J.Physiol Gastrointest Liver Physiol. 2000; 279: 245-9.
- 29. Taub H. Cytokines and liver diseases. Can J Gastroenterology. 2001; 15; 661-8.
- 30. Matin DC, Fowlkes JL, Babic B, Khokha R. Insulin-like growth factor II signaling in neoplastic proliferation is blocked by transgenic expression of the metalloproteinase inhibitor TIMP-1. J Cell Biol. 1999, 146:881-92.
Publication Dates
-
Publication in this collection
20 Sept 2006 -
Date of issue
2006