Abstracts
Use of genetic markers can aid on the identification of animals with highest breeding values in beef cattle. The PIT1 gene codes for the Pit-1 transcription factor is essential for the activation of prolactin, growth hormone and PIT1 genes. This research is an investigation of the effect of PIT1-HinfI polymorphism on growth traits of 509 Canchim animals, from two lineages, GG1 and GG2. PIT1 genotypes were identified through PCR-RFLP. Genotype effect on phenotypic values for birth weight (BW), standardized weaning weight (W240), weight at 12 months of age (W365), and the average daily weight gain from birth to weaning (AGBW), and from weaning to 12 months of age (AGW12) were analyzed by the least squares method. Effects of the interaction between the animal's genetic group and PIT1 genotype for W240, AGBW and AGW12 were observed (P < 0.05). Differences between means of HinfI (-/-), HinfI (+/+) and HinfI (+/-) genotypes for W240 and AGBW were observed in GG2 (P < 0.05), revealing superiority of (-/-) genotype for those traits. Means for genotypes (+/+) and (+/-) for W240 and AGBW, did not differ from one another, suggesting a dominance effect of the HinfI (+) allele. The HinfI (-) allele had a favorable effect on W240 and AGBW in GG2, when present in homozygosis. The difference between PIT1 behavior observed in the two genetic groups may suggest the action of a quantitative trait locus linked to PIT1, segregating only in GG2 population.
RFLP; bovine; pituitary factor 1; candidate gene
O uso de marcadores genéticos pode favorecer a identificação de animais geneticamente superiores em bovino de corte. O gene PIT1 codifica para o fator de transcrição Pit-1, crítico para ativar a expressão dos genes da prolactina, hormônio de crescimento e PIT1. Esta pesquisa investiga o efeito do polimorfismo HinfI-PIT1 sobre características de crescimento em 509 animais da raça Canchim, pertencentes à duas linhagens, GG1 e GG2. Os genótipos do gene PIT1 foram identificados pela técnica PCR-RFLP. Os efeitos dos genótipos sobre valores fenotípicos para pesos ao nascimento (PN), pesos padronizados à desmama (P240) e ao ano (P365), ganhos médios diários de peso do nascimento à desmama (GMND), e da desmama aos 12 meses de idade (GMD12) foram analisados pelo método dos quadrados mínimos. Foram observados efeitos de interação entre grupo genético do animal e genótipo de PIT1 sobre P240, GMND e GMD12 (P < 0.05). Diferenças entre médias dos genótipos HinfI (-/-), HinfI (+/+) e HinfI (+/-) para P240 e GMND foram observadas em GG2 (P < 0,05), revelando superioridade do genótipo HinfI (-/-). As médias dos genótipos (+/+) e (+/-) para P240 e GMND, não diferiram, sugerindo efeito de dominância do alelo HinfI (+). O alelo HinfI (-) mostrou-se favorável sobre P240 e GMND em GG2, quando em homozigose. A diferença de comportamento de PIT1 observada nos dois grupos genéticos pode sugerir a ação de loco de característica quantitativa ligado ao gene PIT1, segregando apenas em GG2.
RFLP; bovinos; fator pituitário 1; gene candidato
ANIMAL SCIENCE AND PASTURES
Association of PIT1 genotypes with growth traits in Canchim cattle
Associação de genótipos PIT1 com características de crescimento em gado Canchim
Sônia Mara CarrijoI; Maurício Mello de AlencarII; Fábio Luiz Buranelo ToralIII; Luciana Correia de Almeida RegitanoII, * * Corresponding author < luciana@cppse.embrapa.br>
IUFSCar - Programa de Pós-Graduação em Genética e Evolução, C.P. 676 - 13565-905 - São Carlos, SP - Brasil
IIEmbrapa Pecuária Sudeste, C.P. 339 - 13560-970 - São Carlos, SP - Brasil
IIIUFMT - Campus Universitário de Rondonópolis, Rod. MT 270, km 6 - 78735-901 - Rondonópolis, MT - Brasil
ABSTRACT
Use of genetic markers can aid on the identification of animals with highest breeding values in beef cattle. The PIT1 gene codes for the Pit-1 transcription factor is essential for the activation of prolactin, growth hormone and PIT1 genes. This research is an investigation of the effect of PIT1-HinfI polymorphism on growth traits of 509 Canchim animals, from two lineages, GG1 and GG2. PIT1 genotypes were identified through PCR-RFLP. Genotype effect on phenotypic values for birth weight (BW), standardized weaning weight (W240), weight at 12 months of age (W365), and the average daily weight gain from birth to weaning (AGBW), and from weaning to 12 months of age (AGW12) were analyzed by the least squares method. Effects of the interaction between the animal's genetic group and PIT1 genotype for W240, AGBW and AGW12 were observed (P < 0.05). Differences between means of HinfI (/), HinfI (+/+) and HinfI (+/) genotypes for W240 and AGBW were observed in GG2 (P < 0.05), revealing superiority of (/) genotype for those traits. Means for genotypes (+/+) and (+/) for W240 and AGBW, did not differ from one another, suggesting a dominance effect of the HinfI (+) allele. The HinfI () allele had a favorable effect on W240 and AGBW in GG2, when present in homozygosis. The difference between PIT1 behavior observed in the two genetic groups may suggest the action of a quantitative trait locus linked to PIT1, segregating only in GG2 population.
Key words: RFLP, bovine, pituitary factor 1, candidate gene
RESUMO
O uso de marcadores genéticos pode favorecer a identificação de animais geneticamente superiores em bovino de corte. O gene PIT1 codifica para o fator de transcrição Pit-1, crítico para ativar a expressão dos genes da prolactina, hormônio de crescimento e PIT1. Esta pesquisa investiga o efeito do polimorfismo HinfI-PIT1 sobre características de crescimento em 509 animais da raça Canchim, pertencentes à duas linhagens, GG1 e GG2. Os genótipos do gene PIT1 foram identificados pela técnica PCR-RFLP. Os efeitos dos genótipos sobre valores fenotípicos para pesos ao nascimento (PN), pesos padronizados à desmama (P240) e ao ano (P365), ganhos médios diários de peso do nascimento à desmama (GMND), e da desmama aos 12 meses de idade (GMD12) foram analisados pelo método dos quadrados mínimos. Foram observados efeitos de interação entre grupo genético do animal e genótipo de PIT1 sobre P240, GMND e GMD12 (P < 0.05). Diferenças entre médias dos genótipos HinfI (/), HinfI (+/+) e HinfI (+/) para P240 e GMND foram observadas em GG2 (P < 0,05), revelando superioridade do genótipo HinfI (/). As médias dos genótipos (+/+) e (+/) para P240 e GMND, não diferiram, sugerindo efeito de dominância do alelo HinfI (+). O alelo HinfI () mostrou-se favorável sobre P240 e GMND em GG2, quando em homozigose. A diferença de comportamento de PIT1 observada nos dois grupos genéticos pode sugerir a ação de loco de característica quantitativa ligado ao gene PIT1, segregando apenas em GG2.
Palavras-chave: RFLP, bovinos, fator pituitário 1, gene candidato
INTRODUCTION
Most economic traits are conditioned by polygenes and subject to environmental variations, which hamper the identification of the best genotypes. Thus, genetic progress in populations depends on statistical methodologies to reveal how the properties observed in populations are influenced by properties of the genes involved and by non-genetic circumstances that might affect a metric trait (Falconer & Mackay, 1996).
Polymorphism detection in genes related to production traits and the identification of the allele which results in a phenotype of interest can allow for marker assisted selection (MAS). Among the genes involved in mammalian growth, PIT1 is considered as a candidate. PIT1 was mapped to Bos taurus chromosome 1 (Moody et al., 1995), and belongs to a group of genes that code for proteins involved in animal development. The specific pituitary transcription factor Pit-1, coded by the PIT1 gene, is required for the initiation of the expression of the growth hormone releasing factor gene (Lin et al., 1992) and is involved in the maintenance of the expression of the gene coding for the b subunit of thyroid stimulating hormone (Lin et al., 1994). It is a critical factor for the activation of the expression of prolactin and growth hormone genes, participating in the activation of PIT1 itself (Parmentier et al., 1999).
A HinfI polymorphism is located within exon 6 and characterized by a silent mutation on HinfI () allele (Dierkes et al., 1998), and their effects were associated with growth traits in cattle (Conde et al., 2001; Silva et al., 2006). The objective of this study was to analyze the distribution of the Restriction Fragment Length Polymorphism (RFLP) PIT1-HinfI and to investigate the influence of allelic variants on traits related to growth in a herd of Canchim cattle.
MATERIAL AND METHODS
Experimental animals
The sample included 509 Canchim animals, resulted from crosses between Bos taurus and Bos indicus (Zebu) breeds, born between 1998 and 2000, fed on pasture with mineral supplements. Animals were separated in two genetic groups. Genetic group 1 (GG1), consisting of 232 Canchim animals with average of 5/8 Charolais and 3/8 of Zebu (Nelore, Guzerá and Indubrasil) (Alencar, 1988). Genetic group 2 (GG2), consisting of 277 animals, with an average of 21/32 Charolais and 11/32 Nelore breeds (Barbosa, 2000). The phenotypic traits considered in the analyses were birth weight (BW), weaning weight (WW), weight at one year (W12), the average daily weight gain from birth to weaning (AGBW), and from weaning to 12 months of age (AGW12).
DNA isolation
DNA samples belonged to the bovine DNA collection, obtained from leukocytes by a salting out procedure (Regitano, 2001).
PCR-RFLP analysis
Polimerase Chain Reaction (PCR) with the PIT1 specific primers 5'-CAATGAGAAAGTTGGTGC-3' and 5'-TCTGCATTCGAGATGCTC-3' was analyzed according to Moody et al. (1995). Amplified products were digested with the restriction enzyme HinfI and fragments were separated in 3% agarose gel, stained with 0.45 µg mL-1 ethidium bromide. The DNA size standard fX174/HaeIII was used for fragment size analysis.
Statistical analysis
The allelic frequencies of the candidate gene PIT1 were estimated in each genetic group by direct count and consisted of the ratio between the number of alleles detected in the population and the number of chromosomes analyzed. The chi-square test was used to compare their proportions in the two genetic groups (GG1 and GG2) according to Snedecor & Cochran (1967).
The effect of the candidate gene PIT1 was analyzed for BW, WW and W12, and for the average daily weight gain from birth to weaning (AGBW) and from weaning to 12 months of age (AGW12). WW and W12 values were standardized for 240 (W240) and 365 (W365) days of age, respectively. Before the analyses, BW, WW and W12 data for the total sample were investigated in comparison with normal distribution by the Shapiro-Wilk's statistics using the SAS univariate procedure, and for homogeneity of variance using Levene's test, using the GLM procedure (SAS, 2000).
The mean phenotypic values attributed to each genotype class were compared by variance analysis using the least squares method, following the GLM procedure (SAS, 2000).
The linear model used was:
Yijkl = µ + GGi + GFj + GPk + GG*PIT1l + eijkl
where Yijkl represents the phenotypic value of the individual for the trait; m, the overall population mean; GGi, the effect of the ith genetic group; GFj, the effect of the jth group of fixed effects (contemporary group, sex and maternal age as a linear and quadratic effect); GPk, the fixed effect of the kth genotype of the candidate gene PIT1; GG*PIT1l, the effect of genetic group x PIT1 genotype interaction; and eijkl, the residual random effect.
Scheffé's test was used to determine the differences between the mean values of the three genotypes at the PIT1 locus (Chew, 1976). To investigate the type of gene action at this locus, the F test was used to determine the contrast between the mean of the two homozygous and the mean of the heterozygous genotypes. Average effect of allele substitution, genetic values, genotypic values and dominance deviations were estimated according to Falconer & Mackay (1996).
RESULTS AND DISCUSSION
Allele frequencies
The amplification of the PIT1 gene fragment resulted in a single product of 1.301 kb. Treatment of this product with the restriction enzyme HinfI revealed the HinfI (+) allele with fragments of 260, 617, 379 and 45 base pairs (bp) and the HinfI () allele with fragments of 260, 617 and 424 bp. The HinfI () allele was the less frequent in the two genetic groups (Table 1). The predominance of the HinfI (+) allele was also detected in different European cattle breeds (Moody et al., 1995; Sabour et al., 1996; Klauzinska et al., 2000; Oprzadek et al., 2003), in Gir and crossbred dairy cattle (Mattos, 2000), and in Gir × Holstein crosses (Silva et al., 2006).
Test for the comparison of allele proportions in GG1 and GG2, revealed that the HinfI () allele frequency was higher in GG2 than in GG1 (P < 0.01). Distribution differences of allele frequencies between different populations may indicate genetic differences in the base populations. Genetic group GG2 originated from genetically superior animals in terms of production traits, since animals from the Nelore, Charolais and Canchim breeds used to produce this population were approximately 40 years of selection apart from the ones used to produce GG1. Another factor that may be considered as having contributed to allelic frequency differences between the two populations is genetic drift.
PIT1 effects on production traits
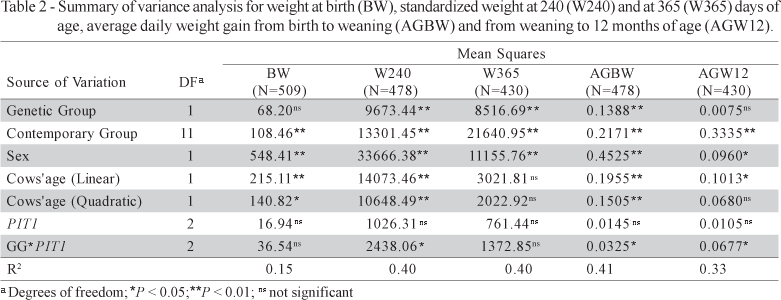
Shapiro-Wilk's statistics indicated that the total population represents a population with normal distribution. Levene's tests for the assessment of homogeneity of variance indicated that the variances observed for the different treatments did not differ from one another. Variance analysis for the total population revealed interaction (P < 0.05) between genetic group and PIT1 genotype for W240, AGBW and AGW12, demonstrating a different PIT1 behavior in the two genetic groups (Table 2).
PIT1 genotype means were different for W240 and AGBW only in GG2 (Table 3). Genotype HinfI (/) was superior to HinfI (+/+) and HinfI (+/) by 24.37 and 24.26 kg, respectively. The HinfI (/) was the most favorable genotype in terms of productivity, since it was related to higher weaning weight values without a concomitant increase in birth weight, which is a desirable condition in beef cattle breeding. Homozygous animals HinfI (/), gained 90.8 and 89.3 g day-1 more than HinfI (+/+) and HinfI (+/), respectively, from birth to weaning.
The effect of PIT1 on W240 and AGBW as detected in GG2 may be the result of its influence on the expression of growth-hormone (GH) and prolactin (PRL) genes, which are equally important for growth from birth to weaning. Studies with swine showed correlation between pituitary GH and PIT1b mRNA levels and effect of PIT1 genotypes on the plasma levels of GH and PRL (Sun et al., 1998, 2002). Therefore, the effects of PIT1 on the W240 and AGBW traits, observed in GG2, ought to be carefully analyzed since, at this stage, calf development is subject to major influences of their mothers' genetic values concerning the maternal abilities, particularly in relation to milk capacity, which was not possible to take into consideration in these analyses. HinfI (/) genotype at the PIT1 gene was associated with higher milk, fat and protein production and with fat percentage in crossbred dairy cattle (Mattos, 2000).
The polymorphism at this locus lies at exon 6 of the PIT1 gene, in the coding regions for the Pit-1 pituitary transcription factor. However, the mutation does not alter the amino acid in the peptide chain of Pit-1 protein. Thus, the differences in W240 and AGBW means for the genotypes of PIT1 in GG2 could be related to other functional mutations in the same gene that would be in linkage disequilibrium with the HinfI-PIT1 polymorphism in GG2 but not in GG1 or would not be present in GG1.
PIT1 encodes proteins that bind to regulatory elements. Forms resulting from alternative splicing can produce proteins with altered properties. In mouse cell cultures, the Pit-1b protein activates the GH promoter more strongly than the promoters of the PRL (Konzak & Moore, 1992), PIT1 (Theill et al., 1992) or b-subunit of thyroid stimulating hormone (TSHb) gene (Haugen et al., 1994), but is an unstable protein (Konzak & Moore, 1992). Results regarding GH activation in swine, revealed a higher influence of Pit-1a mRNA than of PIT1-b on GH expression (Sun et al., 2002). Pit-1a and Pit-1T proteins interact to activate TSHb transcription (Haugen et al., 1993), and this interaction is necessary and specific for the TSHb promoter (Haugen et al., 1994).
Alternative forms of Pit-1, such as Pit-1D4, observed in swine, appear at times of highest PRL expression (Day & Day, 1994), and the Pit-1D3 form expressed in bacteria, was unable to recognize the GH-promoting elements, whereas Pit-1a was active (Yu et al., 2001). There is a strong similarity between the PIT1 genes of swine, cattle and rodents, with approximately 90 to 95% identity between PIT1cDNA and the sequence of its protein deduced in swine, humans, cattle, sheep and rodents (Day & Day, 1994).
The different results observed in GG1 and GG2, concerning the effects of PIT1 on the productive performance of animals, hint at the presence of a quantitative trait locus (QTL) strongly linked to PIT1 in linkage disequilibrium only in GG2, and not the direct effect of this polymorphism on production. This hypothesis, however, could not be tested because of existing constraints due to sample size. Quantitative trait loci may present different effect among populations due to the epistatic interactions between the QTL and the genome (Pomp, 1994). The different behavior of PIT1 in the two genetic groups in this study could be the interaction between the effects of the PIT1 gene and the effects of the target regulatory sequences.
The absence of differences between HinfI (+/+) and HinfI (+/) genotypes, for W240 and AGBW, suggest that the HinfI (+) allele is expressed in a dominant manner in relation to the HinfI () allele. Recessive nature of the allele HinfI () may suggest that the product of this allele has less affinity for the region promoting the growth hormone gene or other genes of the somatotrophic axis on which PIT1 may act, but results in a better pattern of expression. Studies on the affinity of allele specific forms of Pit-1 with the promoters involved in animal development might further elucidate this question.
Higher frequencies of the HinfI (+) allele and of the (+/+) genotype of the PIT1 gene, despite having no positive effect on production, suggest that PIT1 could be linked to a locus with some adaptive advantage located at a distance that is small but enough to generate recombination between them. Alternatively, they may reflect the evolutionary history of this mutation if the HinfI (+) allele is considered as the ancestral allele that underwent a rare case of mutation. This hypothesis seems more appropriate, since the HinfI (+) allele is prevalent in most bovine breeds (Moody et al., 1995; Mattos, 2000).
Allele substitution effect
The estimated population mean for W240 trait in GG2, as a function of allele frequencies, was 219.27 kg. The estimated mean effects of the HinfI () and HinfI (+) alleles were 4.84 and 1.79 kg, respectively, and the mean effect of the allele substitution was 6.63 kg. HinfI () allele was responsible for 2.21% of the mean W240 value, and that the replacement of a HinfI (+) allele with a HinfI () allele may lead to an increase of 6.63 kg on W240 in the animals in this population.
Estimated population mean as a function of allele frequencies for AGBW in GG2, was 771 g day-1. Estimated means of the effects of HinfI () and HinfI (+) alleles were 18.42 and 6.81 g day-1, respectively, and the mean for the effect of allele substitution was 25.23 g day-1. In terms of traits mean, the HinfI () allele may contribute 2.38% of the AGBW mean and that the replacement of one allele HinfI (+) with a HinfI () may lead to an increase of 25.23 g day-1 from birth to weaning in this group of animals.
PIT1 HinfI () allele was related to higher body weight at 7 months of age in double-muscled Belgian Blue bulls (Renaville et al., 1997) and higher weights at 60, 250, 365 days of age and on weight gain from born to 60 days of age in Gir × Holstein crosses (Silva et al., 2006). This polymorphism was associated with weaning weight in Nelore cattle (Conde et al., 2001). On the other hand, no associations of the PIT1 genotypes with meat production traits were found in Piemontese cattle (Di Stasio et al., 2002) or with growth and carcass traits in Angus beef cattle (Zhao et al., 2004). This genetic marker seems to have different effects in different populations, which is in agreement with the results presented here.
Concerning to W240, the genetic values for W240 of homozygous HinfI (/) animals were superior by 6.64 kg compared to heterozygous and by 13.27 kg compared to HinfI (+/+) animals. As for the AGBW trait, HinfI (/) homozygous individuals presented genetic values 25.2 and 50.4 g day-1 greater than the values of heterozygous and of (+/+), respectively (Table 4). HinfI () allele was favorably expressed when present in homozygosis and may be responsible for the superiority of the phenotypic values of weaning weight and of average daily weight gains from birth to weaning observed in this group of animals.
Genotypic value of the HinfI (/) genotype, 22.55 kg, proved to be superior to the remaining genotypes for the W240 trait in GG2. Part of this value is attributed to dominance deviation, whose value was equal to 12.86 kg (Table 4). Similarly, for the AGBW trait, the portion of the genotypic value, 83.5 g day-1, attributed to dominance deviation was 46.7 g day-1. Deviations due to dominance proved to be more important than the genetic values for HinfI (/) and HinfI (+/), implying in an advantage in the use of selection in favor of the HinfI () allele to increase weaning weight and average daily weight gain from birth to weaning. Since the favorable HinfI () allele is recessive to HinfI (+), crossing in order to increase weaning weight and average daily weight gain from birth to weaning would be disadvantageous with respect to this locus.
CONCLUSIONS
Selection in favor of the HinfI () allele of the PIT1 gene may lead to increased weaning weight, without increasing birth weight. However, the difference in PIT1 behavior observed between the two genetic groups denotes the need to test the effects of this polymorphism on different populations before using it in marker-assisted selection. It also indicates the need for a better understanding of the genetic mechanisms involved in the physiology of this pituitary transcription factor in the process of growth in cattle.
ACKNOWLEDGMENTS
To Embrapa and CNPq (fellowships to Maurício Mello de Alencar and Luciana Correia de Almeida Regitano).
Received May 29, 2006
Accepted November 08, 2007
- ALENCAR, M.M. Bovino-raça Canchim: origem e desenvolvimento. Brasília: EMBRAPA-DPU, 1988. 102p.
- BARBOSA, P.F. O Canchim na Embrapa Pecuária Sudeste. In: CONVENÇÃO NACIONAL DA RAÇA CANCHIM, 4., São Carlos, 2000. Anais São Carlos: Embrapa Pecuária Sudeste; São Paulo: ABCCAN, 2000. p.55-68.
- CHEW, V. Comparing treatment means: a compendium. HortScience, v.11, p.348-357, 1976.
- CONDE, S.B.; GUIMARÃES, S.E.F.; EUCLYDES, R.F.; SILVA, L.O.C.; EUCLYDES, F.K. Marcadores moleculares associados com crescimento em bovinos. Revista Brasileira de Reprodução Animal, v.25, p.485-486, 2001.
- DAY, R.N.; DAY, K.H. An alternatively spliced form of Pit-1 represses prolactin gene expression. Molecular Endocrinology, v.8, p.374-381, 1994.
- DIERKES, B.; KRIEGESMANN, B.; BAUMGARTNER, B.G.; BRENIG, B. Partial genomic structure of the bovine PIT1 gene and characterization of a HinfI transition polymorphism in exon 6. Animal Genetics, v.29, p.405, 1998.
- DI STASIO, L.; SARATORE, S.; ALBERA, A. lack of association of GH1 and Pou1F1 gene variants with meat production in Piemontese cattle. Animal Genetics, v.33, p.61-64, 2002.
- FALCONER, D.S.; MACKAY, T.F.C. Introduction to quantitative genetics 4 ed. Edinburgh: Longman, 1996. 464p.
- HAUGEN, B.R.; WOOD, W.M.; GORDON, D.F.; RIDGWAY, E.C. A thyrotrope-specific variant of Pit-1 transactivates the thyrotropin b promoter. The Journal of Biological Chemistry, v.268, p.20818-20824, 1993.
- HAUGEN, B.R.; GORDON, D.F.; NELSON, A.R.; WOOD, W.M.; RIDGWAY, E.C. The combination of Pit-1 and Pit-1T have a synergistic stimulatory effect on the thyrotropin b-subunit promoter but not the growth hormone or prolactin promoters. Molecular Endocrinology, v.8, p.1574-1582, 1994.
- KLAUZINSKA, M.; ZWIERZCHOWSKI, L.; SIADKOWSKA, E.; SZYMANOWSKA, M.; GROCHOWSKA, R.; ZURKOWSKI, M. Comparison of selected gene polymorphisms in Polish Red and Polish Black-and-White cattle. Animal Science Papers and Reports, v.18, p.107-116, 2000.
- KONZAK, K.E.; MOORE, D.D. Functional isoforms of Pit-1 generated by alternative messenger RNA splicing. Molecular Endocrinology, v.6, p.241-247, 1992.
- LIN, C.; LIN, S.C.; CHANG, C.P.; ROSENFELD, M.G. Pit-1-dependent expression of the receptor for growth hormone releasing factor mediates pituitary cell growth. Nature, v.360, p.765-768, 1992.
- LIN, S.C.; LI, S.; DROLET, D.W.; ROSENFELD, M.G. Pituitary ontogeny of the snell dwarf mouse reveals pit-1-independent and pit-1-dependent origins of the thyrotrope. Development, v.120, p.515-522, 1994.
- MATTOS, K.K. Polimorfismos de DNA nos genes BGH e PIT1 em populações zebuínas e mestiças com aptidão leiteira e sua associação com características de produção. São Carlos: UFSCar, 2000. 92p. Tese (Doutorado).
- MOODY, D.E.; POMP, D.; BARENDSE, W. Restriction fragment length polymorphism in amplification products of the bovine PIT1 gene and assignment of PIT1 to bovine chromosome 1. Animal Genetics, v.26, p.45-47, 1995.
- OPRZADEK, J.; FLISIKOWSKI, K.; ZWIERCHOWSKA, L.; DYMNICKI, E.; Polymorphisms at loci of leptin (LEP), PIT1 and STAT5A and their association with growth, feed conversion and carcass quality in Black-and-White bulls. Animal Science Reports, v.2, p.135-145, 2003.
- PARMENTIER, I.; PORTETELLE, D.; GENGLER, N.; PRANDI, A.; BERTOZZI, C.; VLEURICK, L.; GILSON, R.; RENAVILLE, R. Candidate gene markers associated with somatotropic axis and milk selection. Domestic Animal Endocrinology, v.17, p.139-148, 1999.
- POMP, D. Biotechnology and beef cattle improvement: myths and realities. In: RESEARCH SYMPOSIUM AND ANNUAL MEETING, 26., Des Moines, 1994. Proceedings Des Moines: Beef Improvement Fed., 1994. p.236-241.
- REGITANO, L.C.A. Extração de DNA para aplicação em reação em cadeia da polimerase (PCR). In: REGITANO, L.C.A.; COUTINHO, L.L. (Ed.) Biologia molecular aplicada à produção animal Brasília: Embrapa Informação Tecnológica, 2001. p.179-186.
- RENAVILLE, R.; GENGLER, N.; PARMENTIER, I.; MORTIAUX, F.; MASSART, S.; BERTOZZI, C.; BURNY, A.; PORTETELLE, D. Pit-1 gene Hinf I RFLP and growth traits in double-muscled Belgian Blue cattle. Journal of Animal Science, v.75, p.146, 1997. Supplement 1.
- SABOUR, M.P.; LIN, C.Y.; LEE, A.J.; MacLLISTER, A.J. Association between milk protein variants and genetics values of Canadian Holstein bulls for milk yield traits. Journal of Dairy Science, v.79, p.1050-1056, 1996.
- SAS Institute. SAS/Stat: user´s guide; statistics. Cary: SAS Institute, 2000. 3884p.
- SILVA, M.V.G.B.; MARTINEZ, M.L.; MACHADO, M.A.; NASCIMENTO, C.S.; CAMPOS, A.L.; GUIMARÃES, M.F.M.; AZEVEDO, A.L.S.; MOITA, A.K.F.; LUI, J.F. Genes do eixo somatotrófico e características de crescimento numa população F2 de bovinos. Pesquisa Agropecuária Brasileira, v.4, p.981-986, 2006.
- SNEDECOR, G.W.; COCHRAN, W.G. Statistical methods 6 ed. Ames: Iowa State University Press, 1967. 593p.
- SUN, H.S.; ANDERSON, L.L.; TUGGLE, C.K.; KLINDT, J. Quantitative measurement of PIT1, GH and PRL mRNA and circulating hormone levels in pig families segregating PIT1 genotypes. In: INTERNATIONAL CONFERENCE ON ANIMAL GENETICS, 26., Auckland, 1998. Proceedings Auckland: International Society of Animal Genetics, 1998. p.85.
- SUN, H.S.; ANDERSON, L.L.; YU, T.P.; KIM, K.S.; KLINDT, J.; TUGGLE, C.K. Neonatal Meishan pigs show POU1F1 genotype effects on plasma GH and PRL concentration. Animal Reproduction Science, v.69, p.223-237, 2002.
- THEILL, L.E.; HATTORI, K.; LAZZARO, D.; CASTRILLO, J.L.; KARIN, M. Differential splicing of the GHF1 primary transcript gives rise to two functionally distinct homeodomain proteins. The European Molecular Biology Organization Journal, v.11, p.2261-2269, 1992.
- YU, T.P.; SUN, H.S.; WAHLS, S.; SANCHEZ-SERRANO, I.; ROTHSCHILD, M.F.; TUGGLE, C.K. Cloning of the full length pig PIT1 (POU1F1) CDNA and a novel alternative PIT1 transcript, and functional studies of their encoded proteins. Animal Biotechnology, v.12, p.1-19, 2001.
- ZHAO, Q.; DAVIS, M.E.; HINES, H.C. Associations of polymorphisms in the Pit-1 gene with growth and carcass traits in Angus beef cattle. Journal of Animal Science, v.82, p.2229-2233, 2004.
Publication Dates
-
Publication in this collection
31 Mar 2008 -
Date of issue
Apr 2008
History
-
Received
29 May 2006 -
Accepted
08 Nov 2007