Abstracts
The occurence of Listeria species in refrigerated chicken carcasses was evaluated, comparing the conventional culture methodology of FDA, modified by the introduction of a secondary enrichment step prior plating, and the ClearviewTM rapid method (Oxoid, UK Ltd). Forty-eight refrigerated whole chicken carcasses purchased from supermarkets in Florianópolis, SC, Brazil, were analysed. Listeria species occured in 21 (43.7%) samples. Using the Clearview method, 17 (35.4%) samples were positive for Listeria species. Of these isolates, 11 (23%) were L. monocytogenes, 4 (8.3%) L. innocua, 1 (2.1%) L. welshimeri and 1 (2.1%) L. seeligeri. Using the conventional culture methodology of FDA (with modifications), 14 (29.2%) samples were positive for Listeria species. Among these 7 (14.6%) were L. monocytogenes, 6 (12.5%) L. innocua and 1 (2.1%) L. seeligeri. With the Clearview rapid method plus API Listeria for identification, results were confirmed to Listeria species level within 115-139 h. Using the conventional culture method of FDA (with modifications) plus API Listeria, results were confirmed within 120-160 h. However, the Clearview method could indicate the presence of Listeria organisms in only 43 h. Results given by the methods were in moderate concordance and the differences between them were not significant (C.I. = 95%).
Listeria; chicken carcasses; ClearviewTM
Avaliou-se a ocorrência de Listeria spp em carcaças refrigeradas de frango, comparando-se a metodologia convencional recomendada pelo FDA, modificada pela introdução de uma segunda etapa de enriquecimento antes do plaqueamento, e o método rápido ClearviewTM (Oxoid, UK, Ltd). Foram analisadas 48 carcaças de frango de diferentes marcas e supermercados de Florianópolis, Brasil. Listeria spp foi encontrada em 21 (43,7%) amostras. Através do método Clearview encontrou-se 17 (35,4%) amostras positivas para Listeria spp, das quais 11 (23%) eram L. monocytogenes, 4 (8.3%) L. innocua, 1 (2.1%) L. welshimeri e 1 (2.1%) L. seeligeri. Através do método convencional modificado obteve-se um total de 14 (29.2%) amostras positivas para Listeria spp, das quais 7 (14.6%) eram L. monocytogenes, 6 (12.5%) L. innocua e 1 (2.1%) L. seeligeri. Com o método Clearview + API Listeria, obteve-se resultados confirmados à nível de espécie em 115-139 h, e com o método convencional modificado + API Listeria os resultados foram obtidos em 120-160 h. No entanto, o método Clearview pode indicar a presença de Listeria spp em apenas 43 h. Os resultados obtidos pelos métodos utilizados mostraram-se moderadamente concordantes e não apresentaram diferença significativa num intervalo de confiança de 95%.
Listeria spp; carcaças de frango; ClearviewTM
DETECTION OF LISTERIA SPECIES IN REFRIGERATED CHICKEN CARCASSES USING CLEARVIEWTM AND A MODIFIED CONVENTIONAL CULTURE METHOD
Márcia R. Pelisser1; Sandra D.C. Mendes1; Alastair D. Sutherland2; Cleide R.V. Batista1* * Corresponding author. Mailing address: Universidade Federal de Santa Catarina, Departamento de Ciência e Tecnologia de Alimentos, Rod. Admar Gonzaga, 1346, Itacorubi. 88034-001, Florianópolis, SC, Brasil.Tel.: (+5548) 334-4888. Fax: (+5548) 331-9943. E-mail: cbatista@mbox1.ufsc.br
1Departamento de Ciência e Tecnologia de Alimentos, Universidade Federal de Santa Catarina, Florianópolis, SC, Brasil. 2School of Biological and Biomedical Sciences, Glasgow Caledonian University, Glasgow, Scotland.
Submitted: August 31, 2000; Returned to authors for corrections: March 16, 2001; Approved: June 20, 2001
ABSTRACT
The occurence of Listeria species in refrigerated chicken carcasses was evaluated, comparing the conventional culture methodology of FDA, modified by the introduction of a secondary enrichment step prior plating, and the ClearviewTM rapid method (Oxoid, UK Ltd). Forty-eight refrigerated whole chicken carcasses purchased from supermarkets in Florianópolis, SC, Brazil, were analysed. Listeria species occured in 21 (43.7%) samples. Using the Clearview method, 17 (35.4%) samples were positive for Listeria species. Of these isolates, 11 (23%) were L. monocytogenes, 4 (8.3%) L. innocua, 1 (2.1%) L. welshimeri and 1 (2.1%) L. seeligeri. Using the conventional culture methodology of FDA (with modifications), 14 (29.2%) samples were positive for Listeria species. Among these 7 (14.6%) were L. monocytogenes, 6 (12.5%) L. innocua and 1 (2.1%) L. seeligeri. With the Clearview rapid method plus API Listeria for identification, results were confirmed to Listeria species level within 115-139 h. Using the conventional culture method of FDA (with modifications) plus API Listeria, results were confirmed within 120-160 h. However, the Clearview method could indicate the presence of Listeria organisms in only 43 h. Results given by the methods were in moderate concordance and the differences between them were not significant (C.I. = 95%).
Key words: Listeria, chicken carcasses, ClearviewTM
INTRODUCTION
Listeria monocytogenes has been recognised as a pathogen for more than sixty years. However, it has only been identified as a foodborne pathogen after the 1980s, as a result of many human outbreaks of listeriosis (7,15).
L. monocytogenes is widely spread in nature, being commonly found in natural food products and food processing environment as a biofilm which has the capacity to multiply at refrigeration temperatures (10). The organism has been isolated from soil, silage, vegetation, faecal material, water and domestic and industrial effluents (9). Men, animals and the environment are natural reservoirs for this organism. It has been isolated from a variety of animals: more than forty species of mammals and a minimum of 17 different species of birds, including domestic chicken and turkey (4).
Many foods have been implicated in outbreaks of listeriosis including raw and pasteurized milk, cheese, raw and cooked meat products from different animals, vegetables, fish products and ready to eat foods (15). The contamination of both frozen and refrigerated chicken is high in comparison to other foods (13). According to Uboldi Eiroa (16) and Uyttendaele et al. (17), this high incidence is alarming and increases the risk for cross-contamination between raw and cooked food during preparation. It is therefore important to devise a good quality control programme for this bacterium when processing chicken products.
Recent foodborne outbreaks of listeriosis stress the need to develop effective methods for the detection and identification of L. monocytogenes. Particularly, it is important to develop sensitive and rapid methods since conventional methods are time-consuming and laborious.
This study investigated the occurrence of Listeria species in refrigerated whole chicken carcasses sold in supermarkets in Florianópolis, SC, Brazil and evaluated the ClearviewTM rapid method (Oxoid, UK Ltd.) (11) in comparison to the modified FDA conventional culture methods (5,6) for detection of Listeria species. ClearviewTM rapid method is based on the immunoassay technology for the detection of Listeria flagella antigen in cultured samples using monoclonal antibodies. Flagellar protein is extracted from Listeria cells and added to a membrane to which a line of anti-flagellin antibody is bound. If bound flagellin protein is present, it is detected by the addition of a second soluble, anti-flagellin antibody, blue latex complex. The development of a blue line indicates the presence of Listeria species flagellin protein (11).
MATERIALS AND METHODS
Collection, preparation of samples for analysis and testing
Forty-eight refrigerated whole chicken carcasses were randomly collected from six supermarkets in Florianópolis, SC, Brazil, from May to July 1999. Samples were transported to the laboratory in a cold box filled with ice and subjected to microbiological analysis within one hour of collection.
The carcasses were removed aseptically from their original packaging and repackaged in sterile plastic sample bags. Buffered peptone water (BPW) (300 ml) was added to each bag and shaken fifty times to wash the carcass evenly. The BPW was then poured off into a sterile measuring cylinder.
Conventional culture method of isolating Listeria species was modified by the introduction of secondary enrichment step prior plating. A 25 ml aliquot of BPW was added to 225 ml of Listeria Enrichment Broth LEB (Oxoid, UK). The samples were incubated for 4 h at 30ºC, them selective supplements (SR 140E, Oxoid, UK) were added, and samples were incubated a further 20 h. This step helps the recovery of stressed Listeria cells from the food samples (5). Following incubation, 0.1 ml of LEB was transferred to 10 ml Fraser Broth (FB) and incubated at 30ºC for 24 to 40 h. A loopfull of the broth was plated on PALCAM agar (Oxoid, UK) and incubated at 35ºC for up to 48 h. Three typical single colonies were streaked onto Tryptone Soya Yeast Extract Agar (TSYEA) (Oxoid, UK. Ltd.), incubated at 35ºC for 24 h, and submitted to biochemical identification to Listeria species.
ClearviewTM method also employs two enrichment steps to selectively grow Listeria organisms. 25 ml of BPW was added to 225 ml of FB and incubated at 30ºC for 21 h. Following incubation, 0.1 ml of FB was transferred to 10 ml LEB and incubated at 30ºC for 21 h. An aliquot (2 ml) of LEB was heated at 80ºC for 20 min in a water bath to release flagellin protein. The aliquot was then cooled to room temperature and 135 ml inoculated onto the ClearviewTM immunoassay strip as directed by the manufacturer. The results were read after 20 min incubation at room temperature. Positive samples were identified to Listeria species level by plating a loopfull of unheated LEB onto PALCAM agar (Oxoid Ltd. UK) which was incubated at 35ºC for up to 48 h. Typical single colonies were streaked onto Tryptone Soya Yeast Extract Agar (TSYEA), incubated at 35ºC for 24 h, and submitted to biochemical identification.
Identification of isolated colonies to the species level from both the modified FDA conventional culture and the Clearview methods was done using the API Listeria identification strips (10300, bioMérieux, Marcy-l'Étoile, France).
Evaluation of methodologies
The sensitivity of a method is related to its capacity to avoid false negative results, while specificity is related to its ability to not produce false positive results. In this study, a sample was considered true positive when presumptive positive results in both methods were confirmed as Listeria species by the API Listeria identification strips. A sample was considered a true negative when it was negative by the two methods or when it was presumptive positive by one method, but failed to confirm the presence of Listeria species. A result was considered a false positive when it was positive in the kit but was not confirmed as Listeria. A sample was considered as a false negative when it showed a negative result by one method but was a confirmed positive in the other.
Statistical analysis of data
Two non-parametric statistical tests were applied to the data. The first test used SAS software (Microsoft, USA) and determined the coefficient of concordance between the two methods with a confidence interval (CI) of 95% and a a of 5%. The second test correlated the results obtained by the two methods using Statgraf software (Microsoft, USA) with a CI of 95%.
RESULTS
Samples of chicken carcasses were considered positive if Listeria species were isolated by either method studied and the isolates then identified to the species level using API-Listeria strips. Combining the results of the two methods, it was found that 21 out of 48 (43.7%) refrigerated chicken carcass samples were Listeria positive.
The total number of samples positive for Listeria species using the modified FDA conventional culture method was 14 (29.2%). Four (8.4%) of these samples were negative by the Clearview rapid method (Fig. 1). The total number of samples positive for Listeria species using the Clearview rapid method was 17 (35.4%) and seven (14.6%) of them were negative by the conventional culture method (Fig. 1). The number of samples found positive by both methods was 10 (20.8%).
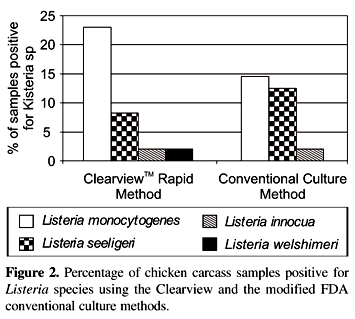
Among 17 samples positive for Listeria species by the Clearview method, 11 (23%) were L. monocytogenes, 4 (8.3%) L. innocua, 1 (2.1%) L. welshimeri and 1 (2.1%) L. seeligeri. Of the 14 samples positive for Listeria species by the conventional culture method; 7 (14.6%) were L. monocytogenes, 6 (12.5%) L. innocua and 1 (2.1%) L. seeligeri (Fig. 2). None of the samples yielded more than one Listeria species.
The rapid ClearviewTM method identified samples positive for Listeria in 43 h. By the modified FDA conventional culture method, presumptive Listeria colonies were detected after 72 to 112 h, which were confirmed species level 48 h later (120 to 160 h).
Results for presence or absence of Listeria species observed in both methods presented a concordance coefficient of 48%. This was considered a moderate concordance, but significant for a 5% (C.I. 95%). On comparing positive and negative results, the paired correlation test showed that the two assay methods were not significantly different at a significance level of 5%.
DISCUSSION
The incidence of Listeria species in chicken carcasses was similar to that reported previously (1,19) using conventional culture methods of isolation. There was a variation in incidence ranging from 2 to 94% for the isolation of Listeria species from chicken depending upon the country of origin and the method employed for isolation (18).
The four false negative samples found by the ClearviewTM rapid method could perhaps be ascribed to insufficient antigen being present for the immunological assay if there was only a small amount of growth in the prior enrichment culture step. According to Roberts (14), the Clearview TM rapid method requires a minimum level of about 5 x 104 to 1 x 105 bacteria/ml in the LEB enrichment step for the assay to record a positive result. It is suggested that the count could be assessed photometrically or confirmed retrospectively by plate counting LEB intended for immunoassay. The same author found no false positives and only one false negative (a chicken sample) were found using the ClearviewTM method on 995 food samples. The seven false negative samples found by the modified FDA conventional culture method could possibly have been due to the presence of other bacteria which inhibited the growth and therefore the isolation of Listeria species. These false negative samples were positive in the ClearviewTM rapid method. This may be because the more inhibitory FB (having more acriflavine hydrochloride) is used before LEB in the Clearview protocol which may have discouraged growth of other bacterial species.
The presence of Listeria species in 43.7% refrigerated chicken carcasses in Florianópolis, Brazil, suggests that there is a significant contamination problem in the processing chain. L. monocytogenes was the most commonly isolated species (23% and 14.6% of Listeria species isolated by ClearviewTM rapid and modified FDA conventional culture methods, respectively). With an increased consumption of chicken in this region this occurence poses a significant threat for an increase in foodborne listeriosis, if chicken is undercooked.
Compared to conventional culture methods, Clearview method is substantially quicker in detecting the presence of Listeria species in samples. By the ClearviewTM rapid method. Samples can be considered negative for Listeria organisms after only 43 h. Thus, the method is convenient for screening Listeria negative food samples.
RESUMO
Detecção de Listeria spp em carcaças refrigeradas de frangos empregando Clearview e um método convencional de cultura modificado
Avaliou-se a ocorrência de Listeria spp em carcaças refrigeradas de frango, comparando-se a metodologia convencional recomendada pelo FDA, modificada pela introdução de uma segunda etapa de enriquecimento antes do plaqueamento, e o método rápido ClearviewTM (Oxoid, UK, Ltd). Foram analisadas 48 carcaças de frango de diferentes marcas e supermercados de Florianópolis, Brasil. Listeria spp foi encontrada em 21 (43,7%) amostras. Através do método Clearview encontrou-se 17 (35,4%) amostras positivas para Listeria spp, das quais 11 (23%) eram L. monocytogenes, 4 (8.3%) L. innocua, 1 (2.1%) L. welshimeri e 1 (2.1%) L. seeligeri. Através do método convencional modificado obteve-se um total de 14 (29.2%) amostras positivas para Listeria spp, das quais 7 (14.6%) eram L. monocytogenes, 6 (12.5%) L. innocua e 1 (2.1%) L. seeligeri. Com o método Clearview + API Listeria, obteve-se resultados confirmados à nível de espécie em 115-139 h, e com o método convencional modificado + API Listeria os resultados foram obtidos em 120-160 h. No entanto, o método Clearview pode indicar a presença de Listeria spp em apenas 43 h. Os resultados obtidos pelos métodos utilizados mostraram-se moderadamente concordantes e não apresentaram diferença significativa num intervalo de confiança de 95%.
Palavras chave:Listeria spp, carcaças de frango, ClearviewTM
- 1. Arnold, G.J.; Coble, J. Incidence of Listeria species in foods in NSW, Food Australia, 47(2):71-75, 1995.
- 2. Beumer, R.R.; Brinkman, E.; Rombouts, F.M. Enzyme-linked immunoassays for the detection of Salmonella spp.: a comparison with other methods. Int. J. Food Microbiol, 12:363374, 1991.
- 3. Casolari, C.F.; Mensione, A.; Messi, G.; Kuaglia, P. Characterization of Listeria monocytogenes strains detected in meat and meat products. Ig. Mod, 101(2):193215, 1994.
- 4. Donnelly, C.; Brackett, R.E.; Doores, S. Listeria In Vanderzant, C., Splittstoesser, D.F. (eds.), Compendium of methods for the microbiological examination of foods, 3. Ed. American Public Health Association Washington, 1992, pp.637-663.
- 5. Hitchins, A.D. In: FDA Bacteriological Analytical Manual 8. Ed., AOAC Int. Gaithersburg, USA, 1995, pp.10.01-10.13.
- 6. Lovett, J. Listeria isolation. In: United States. Department of Health Education and Welfare, Food and Drug Administration. Bacteriological Analytical Manual 6. Ed. supl. 9., Washington, 1987, p.29.1.
- 7. Lovett, J. Listeria monocytogenes In Doyle, M.P. (ed), Foodborne Bacterial Pathogens Marcel Dekker Inc., N.Y., 1989.
- 8. MacDonald, F.; Sutherland, A.D. Important differences in the generation times of Listeria monocytogenes and List. Innocua in two Listeria enrichment broths. J. Dairy Res 61:433-436, 1994.
- 9. McGlaughlin, J. Listeria monocytogenes, recent advances in the taxonomy and epidemiology of listeriosis in humans. J. Appl. Bacteriol, 63:1-11, 1987.
- 10. Muriama, P.M. Bacteriocins for control of Listeria spp. in food. J. Food Prot, 59:54-63, 1996.
-
11Oxoid UK. Ltd. Listeria Rapid Test FT401. Technical Information, Hampshire, N17, 1997.
- 12. Petran, R.L.; Swanson, K.M.J. Simultaneous growth of Listeria monocytogenes and Listeria innocua J. Food Prot, 56(7):616-618, 1993.
- 13. Pini, P.N.; Gilbert, R.J. The occurence in the U.K. of Listeria species in raw chickens and soft cheeses. Int. J. Food Prot, 56(7):616-618, 1993.
- 14. Roberts, P. An improved cultural/immunoassay for the detection of Listeria species in foods and environmental samples. Microb. Eur, 2(5):18-21, 1994.
- 15. Ryser, E.T.; Marth, E.H. Listeria, Listeriosis, and Food Safety University of Wisconsin, New York, 1991, 632p.
- 16. Uboldi-Eiroa, M.N. Listeria monocytogenes Características, ocorrência e desenvolvimento em alimentos. Col. Inst.Tec. Alim, Campinas. 20(1):13-22, 1990.
- 17. Uyttendale, M.R.; Neyts. K.D.; Lips, R.M.; Debevere, J.M. Incidence of Listeria monocytogenes in poultry and poultry products obtained from Belgian and French abbatoirs. Food Microb 14(4):339-345, 1997.
- 18. Waldroup, A.L. Contamination of raw poultry with pathogens. World Poultry Sci. J, 52:7-25, 1996.
- 19. Wong, H.C.; Chao, W.L.; Lee, S.J. Incidence and characterization of Listeria monocytogenes in foods available in Taiwan. Appl. Environ. Microbiol, 56:3101-3104, 1990.
Publication Dates
-
Publication in this collection
12 Dec 2001 -
Date of issue
June 2001
History
-
Reviewed
16 Mar 2001 -
Received
31 Aug 2000 -
Accepted
20 June 2001