ABSTRACT
Pimelodus yuma (formerly Pimelodus blochii) is a freshwater fish, endemic to the Colombian Magdalena-Cauca and Caribbean basins that experiences habitat disturbances resulting from anthropogenic activities. Due to the lack of information about the population genetics of this species, this study developed 14 species-specific microsatellite loci to assess the genetic diversity and population structure of samples from the lower section of the Cauca River. The studied species showed genetic diversity levels higher than the average values reported for Neotropical Siluriformes and significant inbreeding levels as was described for some congeners. Furthermore, P. yuma comprises two coexisting genetic groups that exhibit gene flow along the lower section of the Cauca River. This information constitutes a baseline for future monitoring of the genetic diversity and population structure in an anthropic influenced sector of the Magdalena-Cauca basin.
Keywords:
Freshwater fish; Genetic diversity; Gene flow; Molecular marker; Next generation sequencing
RESUMEN
Pimelodus yuma (anteriormente Pimelodus blochii) es un pez dulceacuícola endémico de las cuencas colombianas Magdalena-Cauca y Caribe que experimenta alteraciones del hábitat como resultado de actividades antropogénicas. Debido a la falta de información sobre la genética poblacional de esta especie, este estudio desarrolló 14 loci microsatélites especie-específicos para evaluar la diversidad genética y la estructura poblacional de muestras de la sección baja del río Cauca. La especie estudiada mostró niveles de diversidad genética más altos que los valores promedio reportados para Siluriformes neotropicales y niveles de endogamia significativos como se describió para algunos congéneres. Además, P. yuma comprende dos grupos genéticos coexistentes que exhiben flujo de genes a lo largo de la sección baja del río Cauca. Esta información constituye una línea base para futuros monitoreos de la diversidad genética y la estructura poblacional en un sector de influencia antrópica de la cuenca Magdalena-Cauca.
Palabras clave:
Diversidad genética; Flujo genético; Marcadores moleculares; Peces de agua dulce; Secuenciación de próxima generación
INTRODUCTION
Seasonally migratory patterns may influence the gene flow or genetic structure in the natural populations of fishes of the family Pimelodidae. Since the gene flow is only possible if the fishes successfully reproduce once they have arrived at their new site (Freeland, 2020Freeland JR. Molecular Ecology. England: John Wiley & Sons; 2020.), both the migrations and reproductive cycles explain the gene flow in Pimelodus maculatus Lacepède, 1803 in the upper section of the Uruguay River, Tibagi River basin, or Tietê River (Almeida et al., 2001Almeida FS, Fungaro MHP, Sodré LMK. RAPD and isoenzyme analysis of genetic variability in three allied species of catfish (Siluriformes: Pimelodidae) from the Tibagi River, Brazil. J Zool. 2001; 253(1):113-20. https://doi.org/10.1017/S0952836901000103
https://doi.org/10.1017/S095283690100010...
; Almeida et al., 2003; Ribolli et al., 2012Ribolli J, Melo CMR, Zaniboni-Filho E. Genetic characterization of the neotropical catfish Pimelodus maculatus (Pimelodidae, Siluriformes) in the Upper Uruguay River. Genet Mol Biol . 2012; 35(4):761-69. doi: https://doi.org/10.1590/S1415-47572012005000060
https://doi.org/10.1590/S1415-4757201200...
) and in Pseudoplatystoma corruscans (Spix & Agassiz, 1829) in the São Francisco River (Dantas et al., 2013Dantas HL, dos Santos MA, Oliveira KKC, Severi W, Diniz FM, Coimbra MR. Genetic diversity of captive and wild threatened Catfish Pseudoplatystoma corruscans in the São Francisco River. Rev Fish Sci. 2013; 21(3-4):237-46. https://doi.org/10.1080/10641262.2013.800787
https://doi.org/10.1080/10641262.2013.80...
) and the Paraguay basins in Brazil (Prado et al., 2017Prado FD, Fernandez-Cebrián R, Foresti F, Oliveira C, Martínez P, Porto-Foresti F. Genetic structure and evidence of anthropogenic effects on wild populations of two neotropical catfishes: Baselines for conservation. J Fish Biol . 2017; 92(1):55-72. https://doi.org/10.1111/jfb.13486
https://doi.org/10.1111/jfb.13486...
).
Despite the lack of spatial genetic structure, discrete genetic stocks that cohabitate the same river area during the migratory period has been observed in Pseudoplatystoma reticulatum Eigenmann & Eigenmann, 1889 in Paraná-Paraguay basin (Prado et al., 2017Prado FD, Fernandez-Cebrián R, Foresti F, Oliveira C, Martínez P, Porto-Foresti F. Genetic structure and evidence of anthropogenic effects on wild populations of two neotropical catfishes: Baselines for conservation. J Fish Biol . 2017; 92(1):55-72. https://doi.org/10.1111/jfb.13486
https://doi.org/10.1111/jfb.13486...
) and Pimelodus grosskopfii Steindachner, 1879 in the Cauca River (Restrepo-Escobar, Márquez, acceptedRestrepo-Escobar N, Yepes-Acevedo AJ, Márquez EJ. Population genetics of three threatened catfish species in heterogeneous environments of the Cauca River, Colombia. Neotrop Ichthyol . Forthcoming 2021.). Additionally, some species in the genera Pimelodus, Pseudoplatystoma and Steindachneridion exhibit genetic structure explained by short-distance migration range, isolation by distance, geographical accidents, disconnections among rivers, or homing behavior (Sekine et al., 2002Sekine ES, Prioli AJ, Prioli SMAP, Júlio Júnior HF. Genetic differentiation among populations of Pseudoplatystoma corruscans (Agassiz, 1829) (Osteichthyes, Pimelodidae) isolated by the Guaíra Falls in the Paraná River. Acta Sci Nol Sci. 2002; 24(2):507-12. Available from: http://repositorio.uem.br:8080/jspui/bitstream/1/5222/1/375.pdf
http://repositorio.uem.br:8080/jspui/bit...
; Almeida et al., 2003Almeida FS, Sodré LMK, Contel EPB. Population structure analysis of Pimelodus maculatus (Pisces, Siluriformes) from the Tietê and Paranapanema Rivers (Brazil). Genet Mol Biol . 2003; 26(3):301-05. https://doi.org/10.1590/S1415-47572003000300014
https://doi.org/10.1590/S1415-4757200300...
; Ramella et al., 2006Ramella MS, Kroth MA, Meurer S, Oliveira AP, Zaniboni E, Maisonnave AC. Genetic variability in four fish species (Pimelodus maculatus, Prochilodus lineatus, Salminus brasiliensis and Steindachneridion scripta) from Uruguay River Basin. Braz Arch Biol Technol. 2006; 49(4):589-98. https://doi.org/10.1590/S1516-89132006000500008
https://doi.org/10.1590/S1516-8913200600...
; Pereira et al., 2009Pereira LHG, Foresti F, Oliveira C. Genetic structure of the migratory catfish Pseudoplatystoma corruscans (Siluriformes: Pimelodidae) suggests homing behaviour. Ecol Freshw Fish. 2009; 18(2):215-25. https://doi.org/10.1111/j.1600-0633.2008.00338.x
https://doi.org/10.1111/j.1600-0633.2008...
; Abreu et al., 2009Abreu MM, Pereira LHG, Barretto Vila V, Foresti F, Oliveira C. Genetic variability of two populations of Pseudoplatystoma reticulatum from the Upper Paraguay River Basin. Genet Mol Biol. 2009; 32(4):868-73. https://doi.org/10.1590/S1415-47572009005000075; Carvalho et al., 2012Carvalho DC, Oliveira DAA, Beheregaray LB, Torres RA. Hidden genetic diversity and distinct evolutionarily significant units in a commercially important Neotropical apex predator, the catfish Pseudoplatystoma corruscans. Conserv Genet. 2012; 13(6):1671-75. https://doi.org/10.1007/s10592-012-0402-6
https://doi.org/10.1007/s10592-012-0402-...
; Telles et al., 2014Telles MPC, Collevatti RG, Braga RS, Guedes LBS, Castro TG, Costa MC, Silva Júnior NJ, Barthem RB, Diniz Filho JAF. Geographical genetics of Pseudoplatystoma punctifer (Castelnau, 1855) (Siluriformes, Pimelodidae) in the Amazon Basin. Genet Mol Res. 2014; 13(2):3656-66. https://doi.org/10.4238/2014.may.9.8
https://doi.org/10.4238/2014.may.9.8...
; Fonseca et al., 2017Fonseca FS, Domingues RR, Hallerman EM, Hilsdorf AWS. Genetic diversity of an imperiled Neotropical catfish and recommendations for its restoration. Front Genet. 2017; 8:196. https://doi.org/10.3389/fgene.2017.00196
https://doi.org/10.3389/fgene.2017.00196...
; Prado et al., 2017Prado FD, Fernandez-Cebrián R, Foresti F, Oliveira C, Martínez P, Porto-Foresti F. Genetic structure and evidence of anthropogenic effects on wild populations of two neotropical catfishes: Baselines for conservation. J Fish Biol . 2017; 92(1):55-72. https://doi.org/10.1111/jfb.13486
https://doi.org/10.1111/jfb.13486...
).
Pimelodus yumaVilla-Navarro & Acero, 2017Villa-Navarro FA, Acero A, Cala Cala P. Taxonomic review of Trans-Andean species of Pimelodus (Siluriformes: Pimelodidae), with the descriptions of two new species. Zootaxa. 2017; 4299(3):337-60. https://doi.org/10.11646/zootaxa.4299.3.2
https://doi.org/10.11646/zootaxa.4299.3....
is one of the 36 valid species of the genus Pimelodus Lacepède, 1803 (Fricke et al., 2020Fricke R, Eschmeyer WN, Van der Laan R. Eschmeyer’s catalog of fishes: genera, species, references [Internet]. San Francisco: California Academy of Science; 2020. Available from: http://researcharchive.calacademy.org/research/ichthyology/catalog/fishcatmain.asp
http://researcharchive.calacademy.org/re...
) distributed in Cauca, Magdalena and Sinú River drainages in Colombia (Villa-Navarro et al., 2017Villa-Navarro FA, Acero A, Cala Cala P. Taxonomic review of Trans-Andean species of Pimelodus (Siluriformes: Pimelodidae), with the descriptions of two new species. Zootaxa. 2017; 4299(3):337-60. https://doi.org/10.11646/zootaxa.4299.3.2
https://doi.org/10.11646/zootaxa.4299.3....
). This species and their congener Pimelodus crypticus Villa-Navarro & Cala, 2017 were both considered as Pimelodus blochii Valenciennes, 1840 before 2017 (Villa-Navarro et al., 2017Villa-Navarro FA, Acero A, Cala Cala P. Taxonomic review of Trans-Andean species of Pimelodus (Siluriformes: Pimelodidae), with the descriptions of two new species. Zootaxa. 2017; 4299(3):337-60. https://doi.org/10.11646/zootaxa.4299.3.2
https://doi.org/10.11646/zootaxa.4299.3....
). Currently, there are no reports of any risk category for P. yuma in the Colombian red list of threatened freshwater fishes (Mojica et al., 2012Mojica JI, Usma-Oviedo JS, Álvarez-León R, Lasso CA, editors. Libro Rojo de peces dulceacuicolas de Colombia (2012). Bogotá, D.C. Colombia: Instituto de Investigación de los Recursos Biológicos Alexander von Humboldt, Instituto de Ciencias Naturales de la Universidad Nacional de Colombia, WWF Colombia, Universidad de Manizales; 2012.) or in the red list of threatened species of the International Union for Conservation of Nature, IUCN. Nevertheless, there is concern about the real status of the species considering the available information of fisheries in the Magdalena-Cauca basin.
Based on the bioeconomic model analysis, P. yuma is considered overexploited (Gutiérrez et al., 2011Gutiérrez F, Barreto C, Mancilla P. Diagnóstico de la pesquería en la cuenca del Magdalena-Cauca. In: Lasso CA, Gutiérrez F, Morales-Betancourt MA, Agudelo E, Ramírez-Gil H, Ajiaco-Martínez RE, editors. II. Pesquerías continentales de Colombia: cuencas del Magdalena-Cauca, Sinú, Canalete, Atrato, Orinoco, Amazonas y vertiente del Pacífico. Bogotá, D.C: Serie Editorial Recursos Hidrobiológicos y Pesqueros Continentales de Colombia, Instituto de Investigación de los Recursos Biológicos Alexander von Humboldt; 2011. p.35-73.). Additionally, fisheries and the fish production modeling data show a decline in number of captures from 17,969 tons in 1975 to 8,370 tons in 2016, indicating the high fishing pressure exerted over this catfish (Barreto Reyes, 2017Barreto Reyes CG. Producción pesquera de la Cuenca del río Magdalena: desembarcos y estimación ecosistémica. Bogotá: The Nature Conservancy, Mac Arthur Foundation, AUNAP; 2017. Available from: http://sepec.aunap.gov.co/Home/VerPdf/63
http://sepec.aunap.gov.co/Home/VerPdf/63...
). Additionally, other factors that compromise this species are the pollution of water due to disposal materials, anthropogenic disturbance of hydrodynamics of the river (hydropower stations), deforestation, and introduction of alien fish species (Jiménez-Segura et al., 2016Jiménez-Segura LF, Galvis-Vergara G, Cala-Cala P, García-Alzate CA, López-Casas S, Ríos-Pulgarín MI et al. Freshwater fish faunas, habitats and conservation challenges in the Caribbean river basins of north-western South America. J Fish Biol. 2016; 89(1):65-101. https://doi.org/10.1111/jfb.13018
https://doi.org/10.1111/jfb.13018...
; Tognelli et al., 2016Tognelli MF, Lasso CA, Bota-Sierra CA, Jiménez-segura LF, Cox NA, editors. Estado de conservación y distribución de la biodiversidad de agua dulce en los Andes tropicales. Gland, Suiza, Cambridge, UK y Arlington, USA: UICN; 2016.). Nonetheless, studies of population genetics of P. yuma remain absent, although they could provide information to estimate the degree of genetic vulnerability of this species.
The use of microsatellite loci permits a high-sensitivity evaluation of the population genetic diversity due to their polymorphism levels and wide distribution in the genome (Triantafyllidis et al., 2002Triantafyllidis A, Krieg F, Cottin C, Abatzopoulos TJ, Triantaphyllidis C, Guyomard R. Genetic structure and phylogeography of European catfish (Silurus glanis) populations. Mol Ecol . 2002; 11(6):1039-55. https://doi.org/10.1046/j.1365-294X.2002.01501.x
https://doi.org/10.1046/j.1365-294X.2002...
). Additional to these advantages, considering the high mutation rates of microsatellite loci, they are also useful to examine recent events (Chistiakov et al., 2006Chistiakov DA, Hellemans B, Volckaert F. Microsatellites and their genomic distribution, evolution, function and applications: A review with special reference to fish genetics. Aquaculture. 2006; 255(1-4):1-29. https://doi.org/10.1016/j.aquaculture.2005.11.031
https://doi.org/10.1016/j.aquaculture.20...
). Despite of the available microsatellite loci for three species within the genus (Paiva, Kalapothakis, 2008Paiva ALB, Kalapothakis E. Isolation and characterization of microsatellite loci in Pimelodus maculatus (Siluriformes: Pimelodidae). Mol Ecol Resour. 2008; 8(5):1078-80. https://doi.org/10.1111/j.1755-0998.2008.02160.x
https://doi.org/10.1111/j.1755-0998.2008...
; Restrepo-Escobar, Márquez, 2020Restrepo-Escobar, Márquez, Yepes- Acevedo AJ, Márquez EJ. Microsatellite loci development for three catfish species from northwestern South America. Neotrop Ichthyol. 2020; 18(1): e190079.https://doi.org/10.1590/1982-0224-2019-0079
https://doi.org/10.1590/1982-0224-2019-0...
; Savada et al., 2020Savada CS, Lima MCC, Orsi ML, Almeida FS. Isolation and characterization of microsatellite loci in Pimelodus microstoma (Siluriformes: Pimelodidae). J Appl Ichthyol. 2020; 36:825-29. https://doi.org/10.1111/jai.14104
https://doi.org/10.1111/jai.14104...
) and some others within Pimelodidae (Batista et al., 2010Batista JS, Farias IP, Formiga-Aquino K, Sousa ACB, Alves-Gomes JA. DNA microsatellite markers for “dourada” (Brachyplatystoma rousseauxii, Siluriformes: Pimelodidae), a migratory catfish of utmost importance for fisheries in the Amazon: development, characterization and inter-specific amplification. Conserv Genet Resour. 2010; 2(1):5-10. https://doi.org/10.1007/s12686-009-9117-5
https://doi.org/10.1007/s12686-009-9117-...
; Carvalho, Beheregaray, 2011Carvalho DC, Beheregaray LB. Rapid development of microsatellites for the endangered Neotropical catfish Conorhynchus conirostris using a modest amount of 454 shot-gun pyrosequencing. Conserv Genet Resour . 2011; 3(2):373-75. https://doi.org/10.1007/s12686-010-9365-4
https://doi.org/10.1007/s12686-010-9365-...
; Saulo-Machado et al., 2011Saulo-Machado AC, Formiga KM, Ortiz MF, Sousa ACB, Alves-Gomes JA, Batista JS. Polymorphic microsatellite DNA markers for the Amazonian catfish Pseudoplatystoma punctifer (Siluriformes: Pimelodidae). Conserv Genet Resour . 2011; 3(2):307-10. https://doi.org/10.1007/s12686-010-9349-4
https://doi.org/10.1007/s12686-010-9349-...
; Souza et al., 2012Souza CA, Hashimoto DT, Pereira LHG, Oliveira C, Foresti F, Porto-Foresti F. Development and characterization of microsatellite loci in Phractocephalus hemioliopterus (Siluriformes: Pimelodidae) and their cross-species amplification in six related species. Conserv Genet Resour . 2012; 4(2):499-501. https://doi.org/10.1007/s12686-011-9584-3
https://doi.org/10.1007/s12686-011-9584-...
; Prado et al., 2014Prado FD, Pardo BG, Guerra-Varela J, Senhorini JA, Martínez P, Foresti F, Porto-Foresti F. Development and characterization of 16 microsatellites for the Neotropical catfish Pseudoplatystoma reticulatum and cross species analysis. Conserv Genet Resour . 2014; 6(3):679-81. https://doi.org/10.1007/s12686-014-0180-1
https://doi.org/10.1007/s12686-014-0180-...
), their use for studying the population genetics of P. yuma may be problematic. The limitations of cross-amplification of microsatellite loci, related to unsuccessful amplification in phylogenetically distant species (Barbará et al., 2007Barbará T, Palma-Silva C, Paggi GM, Bered F, Fay MF, Lexer C. Cross-species transfer of nuclear microsatellite markers: Potential and limitations. Mol Ecol. 2007; 16(18):3759-67. https://doi.org/10.1111/j.1365-294X.2007.03439.x
https://doi.org/10.1111/j.1365-294X.2007...
), include low levels of polymorphism, presence of null alleles (Rutkowski et al., 2009Rutkowski R, Sielezniew M, Szostak A. Contrasting levels of polymorphism in cross-amplified microsatellites in two endangered xerothermophilous, obligatorily myrmecophilous, butterflies of the genus Phengaris (Maculinea) (Lepidoptera: Lycaenidae). Eur J Entomol. 2009; 106(4):457-69. https://doi.org/10.14411/eje.2009.058
https://doi.org/10.14411/eje.2009.058...
), allele size homoplasy (Estoup et al., 2002Estoup A, Jarne P, Cornuet JM. Homoplasy and mutation model at microsatellite loci and their consequences for population genetics analysis. Mol Ecol . 2002; 11(9):1591-604. https://doi.org/10.1046/j.1365-294X.2002.01576.x
https://doi.org/10.1046/j.1365-294X.2002...
), and inability to evaluate properly orthologous loci (Yue et al., 2010Yue GH, Kovacs B, Orban L. A new problem with cross-species amplification of microsatellites: Generation of non-homologous products. Dongwuxue Yanjiu. 2010; 31(2):131-40. doi: https://doi.org/10.3724/SP.J.1141.2010.02131
https://doi.org/10.3724/SP.J.1141.2010.0...
).
Since Pimelodus yuma was catalogued as a medium-distance migrant (100-500 km) (Usma et al., 2013Usma Oviedo JS, Villa-Navarro F, Lasso CA, Castro F, Zúñiga-Upegui PT, Cipamocha CA, Ortega-Lara A, Ajiaco RE, Ramírez-Gil H, Jiménez LF, Maldonado-Ocampo J, Muñoz JA, Suárez JT. Peces dulceacuícolas Migratorios de Colombia. In: Zapata LA, Usma JS, editors. Guía de las Especies Migratorias de la Biodiversidad en Colombia. Peces. Vol. 2. Bogotá: Ministerio de Ambiente y Desarrollo Sostenible, WWF-Colombia; 2013. p.215-441.) and the sampling sites in this study are separated by less than 260 km with a geography lacking slopes, rapids, cascades, the existence of gene flow in P. yuma was hypothesized for three sites of the lower sections of the Cauca River. Furthermore, given the anthropogenic intervention in the Magdalena-Cauca basin and the decrease of captures, it is expected a loss in genetic diversity. To test these hypotheses, we developed a set of species-specific microsatellite loci that allow the study of the population genetics for the endemic Colombian catfish.
MATERIAL AND METHODS
This study analyzed 138 muscle tissues of Pimelodus yuma from the main channel and floodplain lakes in the lower sections of the Cauca River. Although a sampling effort using gillnet, and cast nets was made along the eight sections of the Cauca River described by Landínez-García, Márquez (2016Landínez-García RM, Márquez EJ. Development and characterization of 24 polymorphic microsatellite loci for the freshwater fish Ichthyoelephas longirostris (Characiformes: Prochilodontidae). PeerJ. 2016; 4:e2419. https://doi.org/10.7717/peerj.2419
https://doi.org/10.7717/peerj.2419...
), this species was found only in the area that corresponds to three sampling sections (S4/5, S6, and S7/8) (Fig. 1, S1). The S4/5 section consists of sites distributed along the main channel of the river and floodplain lakes, whereas S6 and S7/8 encompass floodplain lakes in the lower section of the Cauca River. The analyzed sections were selected according to the availability of samples and geographical distance. All the samples were collected by Integral S.A. from 2011 to 2014, before the construction of the Hydroelectric Ituango Project. The samples, preserved in ethanol 96%, were provided to the laboratory by Integral S.A. through two scientific cooperation agreements (September 19th, 2013; Grant CT-2013).
| Studied sampling sites of Pimelodus yuma along the lower sections (S4-S8) of the Cauca River. The pentagons indicate sampling sites in floodplain lakes and the stars indicate sites along the main channel of the river.
We followed the methodology described by Landínez-García, Marquez (2018Landínez-García RM, Marquez EJ. Microsatellite loci development and population genetics in Neotropical fish Curimata mivartii (Characiformes: Curimatidae). PeerJ. 2018; 6:e5959. https://doi.org/10.7717/peerj.5959
https://doi.org/10.7717/peerj.5959...
) to develop the species-specific microsatellite loci for P. yuma. The DNA isolation was performed in one individual of P. yuma from S8, a site of the lower section of the Cauca River. The 454 GS FLX (+) (ROCHE) technology was used for pyrosequencing the previously obtained genomic library. Sequences reads were analyzed with the PRINSEQ-LITE v0.20.4 to assess the quality of the reads and to eliminate sequences with less than 100 bp in length. Then, PAL_FINDER v0.02.03 (Castoe et al., 2010Castoe TA, Poole AW, Gu W, Jason de Koning AP, Daza JM, Smith EN, Pollock DD. Rapid identification of thousands of copperhead snake (Agkistrodon contortrix) microsatellite loci from modest amounts of 454 shotgun genome sequence. Mol Ecol Resour. 2010; 10(2):341-47. https://doi.org/10.1111/j.1755-0998.2009.02750.x
https://doi.org/10.1111/j.1755-0998.2009...
) was used to identify and extract microsatellite loci with perfect di-, tri-, tetra-, penta- and hexa- nucleotide motifs. To design the primers, the flanking sequences of the microsatellite loci were analyzed using PRIMER3 v2.0 (Rozen, Skaletsky, 2000Rozen S, Skaletsky H. Primer3 on the World Wide Web for general users and for biologist programmers. In: Misener S, Krawetz S, editors. Bioinformatics methods and protocols. New Jersey: Humana Press; 2000. p.365-386). An electronic PCR (primer-BLAST; available in https://ncbiinsights.ncbi.nlm.nih.gov/2017/06/28/e-pcr-is-retiring-use-primer-blast/) was carried out to evaluate the correct alignment of the selected primers.
Forty-one microsatellite loci were selected to evaluate their amplification capacity. To assess PCR conditions (Landínez-García, Márquez, 2016Landínez-García RM, Márquez EJ. Development and characterization of 24 polymorphic microsatellite loci for the freshwater fish Ichthyoelephas longirostris (Characiformes: Prochilodontidae). PeerJ. 2016; 4:e2419. https://doi.org/10.7717/peerj.2419
https://doi.org/10.7717/peerj.2419...
), DNA was extracted with the GeneJET Genomic DNA Purification Kit (ThermoScientific) following the manufacturer’s instructions. The amplification capacity of the loci was tested using two random samples and the products of the amplifications were separated through electrophoresis in 10% polyacrylamide gel setting 100 volts during 45 min in a Mini-PROTEAN Tetra Cell (Bio-Rad) and visualized with silver stain. Then, 28 loci that showed amplification capacity were tested using 12 random DNA samples from the three sections analyzed. Later, 22 loci were selected due to their level of polymorphism and band resolution within 100 and 350 bp in size. A set of 14 microsatellite loci that showed well-defined peaks and absence of stutter bands were considered for population genetic analysis.
The optimal conditions and concentrations for amplification reactions were as proposed by Landínez-García, Marquez (2018Landínez-García RM, Marquez EJ. Microsatellite loci development and population genetics in Neotropical fish Curimata mivartii (Characiformes: Curimatidae). PeerJ. 2018; 6:e5959. https://doi.org/10.7717/peerj.5959
https://doi.org/10.7717/peerj.5959...
) with some modifications described below. In this three-primer strategy, 0.5 pmol/μL was used for each of the forward primer labeled in the 5’ end with each one the adapters (Blacket et al., 2012Blacket MJ, Robin C, Good RT, Lee SF, Miller AD. Universal primers for fluorescent labelling of PCR fragments - an efficient and cost-effective approach to genotyping by fluorescence. Mol Ecol Resour. 2012; 12(3):456-63. https://doi.org/10.1111/j.1755-0998.2011.03104.x
https://doi.org/10.1111/j.1755-0998.2011...
), 1.0 pmol/μL of each reverse primer and 10-12 ng/μL of template DNA. Furthermore, the PCR amplifications were carried out with an initial denaturalization at 94°C for 3 min, followed by 45 cycles consisting of a denaturalization step of 90°C for 22 s and an annealing step of 56°C for 18 s without an extension step or final elongation. The products were separated by electrophoresis in an automatic sequencer ABI 3500 HD (Applied Biosystems), using the GeneScan™ 600 LIZ™ dye size standard (Applied Biosystems), and the allelic sizes were edited using GENEMAPPER ID-X v1.5. Potential genotyping errors were detected using Micro-Checker v.2.2.3 software (Van Oosterhout et al., 2004Van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P. MICRO-CHECKER: Software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes. 2004; 4(3):535-38. https://doi.org/10.1111/j.1471-8286.2004.00684.x).
The polymorphic information content (PIC) was calculated with CERVUS v3.0.7 (Marshall et al., 1998Marshall TC, Slate J, Kruuk LEB, Pemberton JM. Statistical confidence for likelihood-based paternity inference in natural populations. Mol Ecol . 1998; 7(5):639-55. https://doi.org/10.1046/j.1365-294x.1998.00374.x
https://doi.org/10.1046/j.1365-294x.1998...
) for each locus. The software ARLEQUIN v3.5.2.2 (Excoffier, Lischer, 2010Excoffier L, Lischer HEL. Arlequin suite ver 3.5: A new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour. 2010; 10(3):564-67. https://doi.org/10.1111/j.1755-0998.2010.02847.x
https://doi.org/10.1111/j.1755-0998.2010...
) was used to test Hardy-Weinberg (HW) and linkage equilibrium for all the microsatellite loci. Additionally, in multiple comparisons, the statistical significance was adjusted with the sequential Bonferroni correction (Holm, 1978Holm S. A simple sequentially rejective multiple test procedure. Scand Stat Theory Appl. 1979; 6(2): 65-70. Available from: https://www.jstor.org/stable/4615733
https://www.jstor.org/stable/4615733...
; Rice, 1989Rice W. Analyzing tables of statistical tests. Evolution. 1989; 43(1):223-25. http://dx.doi.org/10.1111/j.1558-5646.1989.tb04220.x
http://dx.doi.org/10.1111/j.1558-5646.19...
). The genetic diversity estimators, such as allelic diversity, expected (HE), observed (HO) heterozygosities, and inbreeding coefficient (FIS), were calculated for each locus and across loci with ARLEQUIN v3.5 (Excoffier, Lischer, 2010Excoffier L, Lischer HEL. Arlequin suite ver 3.5: A new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour. 2010; 10(3):564-67. https://doi.org/10.1111/j.1755-0998.2010.02847.x
https://doi.org/10.1111/j.1755-0998.2010...
).
Evidences of geographical genetic differentiation among populations were examined with the standardized statistics F’ST (Meirmans, 2006Meirmans PG. Using the AMOVA framework to estimate a tandardized genetic differentiation measure. Evolution. 2006; 60(11):2399-2402. https://doi.org/10.1554/05-631.1), Jost’s D’EST (Meirmans, Hedrick, 2011Meirmans PG, Hedrick PW. Assessing population structure: FST and related measures. Mol Ecol Resour. 2011; 11(1):5-18. https://doi.org/10.1111/j.1755-0998.2010.02927.x
https://doi.org/10.1111/j.1755-0998.2010...
), and an analysis of molecular variance (AMOVA) (Meirmans, 2006Meirmans PG. Using the AMOVA framework to estimate a tandardized genetic differentiation measure. Evolution. 2006; 60(11):2399-2402. https://doi.org/10.1554/05-631.1) included in GENALEX v6.503 (Peakall, Smouse, 2012Peakall R, Smouse PE. GenALEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics. 2012; 28(19):2537-39. https://doi.org/10.1093/bioinformatics/bts460
https://doi.org/10.1093/bioinformatics/b...
). Additionally, a discriminant analysis of principal components was carried out with using the R-package ADEGENET (Jombart, 2008Jombart T. Adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics. 2008; 24(11):1403-05. https://doi.org/10.1093/bioinformatics/btn129
https://doi.org/10.1093/bioinformatics/b...
) considering 138 individuals and 18 retained principal components.
The above calculations were complemented with a Bayesian analysis in STRUCTURE v2.3.4 (Pritchard et al., 2000Pritchard JK, Stephens M, Donnelly P. Inference of population structure using multilocus genotype data. Genetics 2000; 155(2):945-59. https://doi.org/10.1111/j.1471-8286.2007.01758.x
https://doi.org/10.1111/j.1471-8286.2007...
; Falush et al., 2002Falush D, Matthew S, Pritchard KJ. Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics. 2003; 164(4): 1567-87. Available from: https://www.genetics.org/content/164/4/1567
https://www.genetics.org/content/164/4/1...
, 2007Falush D, Stephens M, Pritchard JK. Inference of population structure using multilocus genotype data: Dominant markers and null alleles. Mol Ecol Notes. 2007; 7(4):574-78. https://doi.org/10.1111/j.1471-8286.2007.01758.x
https://doi.org/10.1111/j.1471-8286.2007...
; Hubisz et al., 2009Hubisz MJ, Falush D, Stephens M, Pritchard JK. Inferring weak population structure with the assistance of sample group information. Mol Ecol Resour. 2009; 9(5):1322-32. https://doi.org/10.1111/j.1755-0998.2009.02591.x
https://doi.org/10.1111/j.1755-0998.2009...
). This analysis was performed with 200,000 Monte Carlo Markov Chain (MCMC) steps, 20,000 iterations as burn-in and setting the parameters admixture and non-admixture models, as well as correlated frequencies. The analysis was repeated 20 times per each simulated genetic group (K), which range between 1 and 6 groups. Finally, the web-based software STRUCTURESELECTOR (Li, Liu, 2018Li YL, Liu JX. StructureSelector: A web-based software to select and visualize the optimal number of clusters using multiple methods. Mol Ecol Resour. 2018; 18(1):176-77. https://doi.org/10.1111/1755-0998.12719
https://doi.org/10.1111/1755-0998.12719...
) was used to calculate the best estimated K, the estimators MEDMEANK, MAXMEANK, MEDMEDK, MAXMEDK (Puechmaille, 2016Puechmaille SJ. The program STRUCTURE does not reliably recover the correct population structure when sampling is uneven: Subsampling and new estimators alleviate the problem. Mol Ecol Resour. 2016; 16(3):608-27. https://doi.org/10.1111/1755-0998.12512
https://doi.org/10.1111/1755-0998.12512...
), and heuristic scores (Raj et al., 2014Raj A, Stephens M, Pritchard JK. fastSTRUCTURE: Variational Inference of Population Structure in Large SNP Data Sets. Genetics. 2014; 197(2):573-89. https://doi.org/10.1534/genetics.114.164350
https://doi.org/10.1534/genetics.114.164...
). The summarized results of the runs were plotted in a co-ancestry histogram of all the individuals using STRUCTURESELECTOR and the integrated software CLUMPAK (Kopelman et al., 2015Kopelman NM, Mayzel J, Jakobsson M, Rosenberg NA, Mayrose I. Clumpak: a program for identifying clustering modes and packaging population structure inferences across K. Mol Ecol Resour. 2015; 15(5):1179-91. https://doi.org/10.1111/1755-0998.12387
https://doi.org/10.1111/1755-0998.12387...
). Finally, based on coancestry coefficients obtained with STRUCTURE and CLUMPP v1.1.1 (Jakobsson, Rosenberg, 2007Jakobsson M, Rosenberg NA. CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics. 2007; 23(14):1801-06. https://doi.org/10.1093/bioinformatics/btm233
https://doi.org/10.1093/bioinformatics/b...
), the individuals were rearranged by genetic stock and then were accordingly analyzed.
RESULTS
A total of 14 microsatellite loci were characterized in this study (Tab. 1); such loci did not show evidence of large allele dropout or stutter bands. Although two loci seemed to exhibit null alleles after Bonferroni correction, the departures from Hardy-Weinberg and heterozygote deficit, also observed in other loci in greater sample size, indicate a possible Wahlund effect (Tab. 2) as discussed below. The number of alleles per locus ranged between 8 and 23 with an average value of 13.071 alleles/locus. The allele sizes were in the expected range (100-350 bp) with a minimum size of 114 bp and a maximum of 347 bp. The polymorphic information content (PIC) showed values ranging from 0.508 to 0.937. Additionally, the observed and expected heterozygosities presented average values of 0.670 and 0.849, respectively. Five of the 14 developed microsatellite loci were excluded from the further genetic diversity analysis pending exploration in a greater sample size (Pyu6, Pyu7, Pyu11, Pyu22, and Pyu32).
| Sequences and characteristics of 14 polymorphic microsatellite loci selected for Pimelodus yuma. Na: number of alleles per locus; Ra: allele size range; HO and HE: observed and expected heterozygosities, respectively; P: statistical significance for the tests of Hardy-Weinberg equilibrium; PIC: polymorphic information content.
| Genetic diversity of Pimelodus yuma in three sections (S4/5, S6, and S7/8) of the Cauca River in Colombia. Na: number of alleles per locus; HO and HE: observed and expected heterozygosities, respectively; FIS: inbreeding coefficient; P: statistical significance for the Hardy-Weinberg test, values in bold indicate significance after Bonferroni correction.
The diversity estimators (Tab. 2) showed the lowest average expected heterozygosity in S4/5 (0.832) and the highest in S7/8 (0.873). The observed heterozygosity exhibited the lowest value in S6 (0.703) and the highest in S4/5 (0.744). Additionally, S4/5 (12.556 alleles/locus) showed the lowest average number of alleles per locus while S6 showed the highest average number (15.889 alleles/locus). Only two of the nine loci were departed from HWE in the three evaluated sites and the other were in the HWE in at least one site. Additionally, the across loci values of the inbreeding coefficients for all analyzed sites showed deficit of heterozygosity (0.109 < FIS < 0.186). These levels of genetic diversity and inbreeding were similar to those obtained for the genetic stocks (Stock1 and Stock2) supported by the genetic structure analysis.
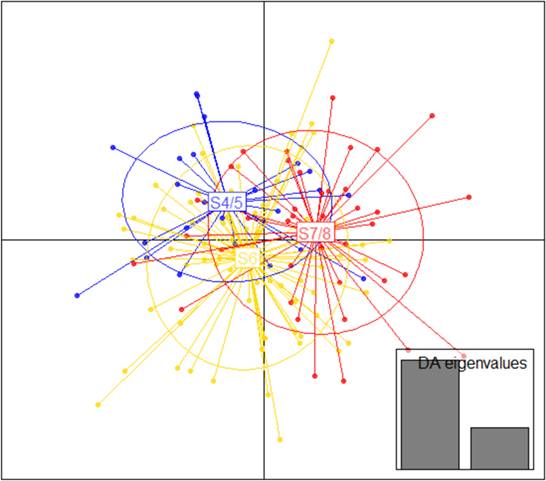
Using the pairwise (S4/5-S6, S4/5-S7/8, S6-S7/8) standardized estimators F’ST (-0.014, 0.019, -0.004) and DEST (0.007, 0.011, 0.005) there was not statistical significance among the individuals of the different analyzed sites. Similarly, the AMOVA (F’ST = -0.003; P= 0.605) and the DAPC (Fig. 2) also showed that there were no differences among the three sections of the river. Hence, the individuals from the three sampling sites were genetically similar.
The analysis in STRUCTURE showed two genetic stocks that coexist in the studied area (Fig. 3, S2; DeltaK = 2; MEDMEANK, MAXMEANK, MEDMEDK = 2; Mean LnP(K) = -5944.415 and -5930.270, for admixture and non-admixture models, respectively), without evidence of geographical structure. The analysis of diversity of the two stocks found in the study area evidenced that even without a geographical arrangement of the individuals, the heterozygosity deficit (0.000; 0.000) and FIS (0.157; 0.159) were statistically significant for both genetic stocks.
| Discriminant analysis of principal components for nine microsatellite loci and 138 individuals of Pimelodus yuma in three sections (S4/5, S6 and S7/8) of the Cauca River.
| STRUCTURE results for Pimelodus yuma showing K= 2 genetic stocks in three sections (S4/5, S6 and S7/8) of the Cauca River.
DISCUSSION
This study developed species-specific microsatellite loci to tests the hypothesis that Pimelodus yuma exhibits gene flow in three sites without geographical barriers, separated by less than 260 km in the middle and lower sections of the Cauca River. Among 41 microsatellite loci examined, 14 are considered highly informative showing values of PIC above the range proposed by Botstein et al. (1980Botstein D, White RL, Skolnick M, Davis RW. Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet. 1980; 32(3):314-31. Available from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1686077/
https://www.ncbi.nlm.nih.gov/pmc/article...
). These loci exhibited values of average number alleles per locus (13.071) and expected heterozygosities (0.849) lower than those reported in the congener P. grosskopfii (Restrepo-Escobar, Márquez, 2020Restrepo-Escobar, Márquez, Yepes- Acevedo AJ, Márquez EJ. Microsatellite loci development for three catfish species from northwestern South America. Neotrop Ichthyol. 2020; 18(1): e190079.https://doi.org/10.1590/1982-0224-2019-0079
https://doi.org/10.1590/1982-0224-2019-0...
), but higher than those reported in P. microstoma Steindachner, 1877 (Savada et al., 2020Savada CS, Lima MCC, Orsi ML, Almeida FS. Isolation and characterization of microsatellite loci in Pimelodus microstoma (Siluriformes: Pimelodidae). J Appl Ichthyol. 2020; 36:825-29. https://doi.org/10.1111/jai.14104
https://doi.org/10.1111/jai.14104...
) and were similar to those reported in P. maculatus (Paiva, Kalapothakis, 2008Paiva ALB, Kalapothakis E. Isolation and characterization of microsatellite loci in Pimelodus maculatus (Siluriformes: Pimelodidae). Mol Ecol Resour. 2008; 8(5):1078-80. https://doi.org/10.1111/j.1755-0998.2008.02160.x
https://doi.org/10.1111/j.1755-0998.2008...
). Furthermore, these values were higher than those reported in Brachyplatystoma rousseauxii (Castelnau, 1855) (Batista et al., 2010Batista JS, Farias IP, Formiga-Aquino K, Sousa ACB, Alves-Gomes JA. DNA microsatellite markers for “dourada” (Brachyplatystoma rousseauxii, Siluriformes: Pimelodidae), a migratory catfish of utmost importance for fisheries in the Amazon: development, characterization and inter-specific amplification. Conserv Genet Resour. 2010; 2(1):5-10. https://doi.org/10.1007/s12686-009-9117-5
https://doi.org/10.1007/s12686-009-9117-...
), Conorhynchus conirostris (Valenciennes, 1840) (Carvalho, Beheregaray, 2011Carvalho DC, Beheregaray LB. Rapid development of microsatellites for the endangered Neotropical catfish Conorhynchus conirostris using a modest amount of 454 shot-gun pyrosequencing. Conserv Genet Resour . 2011; 3(2):373-75. https://doi.org/10.1007/s12686-010-9365-4
https://doi.org/10.1007/s12686-010-9365-...
), Pseudoplatystoma reticulatum (Prado et al., 2014Prado FD, Pardo BG, Guerra-Varela J, Senhorini JA, Martínez P, Foresti F, Porto-Foresti F. Development and characterization of 16 microsatellites for the Neotropical catfish Pseudoplatystoma reticulatum and cross species analysis. Conserv Genet Resour . 2014; 6(3):679-81. https://doi.org/10.1007/s12686-014-0180-1
https://doi.org/10.1007/s12686-014-0180-...
), Pseudoplatystoma punctifer (Castelnau, 1855) (Saulo-Machado et al., 2011Saulo-Machado AC, Formiga KM, Ortiz MF, Sousa ACB, Alves-Gomes JA, Batista JS. Polymorphic microsatellite DNA markers for the Amazonian catfish Pseudoplatystoma punctifer (Siluriformes: Pimelodidae). Conserv Genet Resour . 2011; 3(2):307-10. https://doi.org/10.1007/s12686-010-9349-4
https://doi.org/10.1007/s12686-010-9349-...
), and Phractocephalus hemioliopterus (Bloch & Schneider, 1801) (Souza et al., 2012Souza CA, Hashimoto DT, Pereira LHG, Oliveira C, Foresti F, Porto-Foresti F. Development and characterization of microsatellite loci in Phractocephalus hemioliopterus (Siluriformes: Pimelodidae) and their cross-species amplification in six related species. Conserv Genet Resour . 2012; 4(2):499-501. https://doi.org/10.1007/s12686-011-9584-3
https://doi.org/10.1007/s12686-011-9584-...
).
The genetic analysis of 138 samples of P. yuma showed high genetic diversity comparing values of alleles per locus (17.444) and expected heterozygosity (0.861) to those described in Neotropical Siluriformes (Na: 7.450, HE: 0.609; Hilsdorf, Hallerman, 2017Hilsdorf AWS, Hallerman EM. Genetic resources of Neotropical fishes. Cham: Springer International Publishing AG; 2017.). Additionally, P. yuma showed similar values to those reported for the congeners P. grosskopfii (Restrepo-Escobar, Márquez, 2020Restrepo-Escobar, Márquez, Yepes- Acevedo AJ, Márquez EJ. Microsatellite loci development for three catfish species from northwestern South America. Neotrop Ichthyol. 2020; 18(1): e190079.https://doi.org/10.1590/1982-0224-2019-0079
https://doi.org/10.1590/1982-0224-2019-0...
) and P. maculatus (Ribolli et al., 2012Ribolli J, Melo CMR, Zaniboni-Filho E. Genetic characterization of the neotropical catfish Pimelodus maculatus (Pimelodidae, Siluriformes) in the Upper Uruguay River. Genet Mol Biol . 2012; 35(4):761-69. doi: https://doi.org/10.1590/S1415-47572012005000060
https://doi.org/10.1590/S1415-4757201200...
). In contrast, P. yuma showed higher diversity than the reported for some members of the family Pimelodidae such as Brachyplatystoma rousseauxii (Carvajal-Vallejos et al., 2014Carvajal-Vallejos FM, Duponchelle F, Desmarais E, Cerqueira F, Querouil S, Nuñez J et al. Genetic structure in the Amazonian catfish Brachyplatystoma rousseauxii: Influence of life history strategies. Genetica. 2014; 142(4):323-36. https://doi.org/10.1007/s10709-014-9777-2
https://doi.org/10.1007/s10709-014-9777-...
), P. corruscans (Vaini et al., 2016Vaini JO, Crispim BA, Santos Silva DB, Benites C, Russo MR, Grisolia AB. Genetic variability of pure Pseudoplatystoma corruscans and Pseudoplatystoma reticulatum individuals in the Paraná and Paraguay River basins. Fish Sci. 2016; 82(4):605-11. https://doi.org/10.1007/s12562-016-0999-3
https://doi.org/10.1007/s12562-016-0999-...
), and P. reticulatum (Prado et al., 2017Prado FD, Fernandez-Cebrián R, Foresti F, Oliveira C, Martínez P, Porto-Foresti F. Genetic structure and evidence of anthropogenic effects on wild populations of two neotropical catfishes: Baselines for conservation. J Fish Biol . 2017; 92(1):55-72. https://doi.org/10.1111/jfb.13486
https://doi.org/10.1111/jfb.13486...
). A high genetic diversity is a desirable characteristic for species, since it helps them to adapt to eventual adverse events (Li et al., 2017Li L, Lin H, Tang W, Liu D, Bao B, Yang J. Population genetic structure in wild and aquaculture populations of Hemibarbus maculates inferred from microsatellites markers. Aquac Fish. 2017; 2(2):78-83. https://doi.org/10.1016/j.aaf.2017.03.004
https://doi.org/10.1016/j.aaf.2017.03.00...
).
Despite the high genetic diversity, P. yuma exhibited significant deficit of observed heterozygosity that could be explained by several non-excluding alternatives. The presence of null alleles is invoked as a technical cause of the deficit of observed heterozygosity although the development of species-specific loci minimizes this possibility. As discussed below, P. yuma represents two genetic stocks that cohabitate the studied area, which may explain the deficit of observed heterozygosity (Wahlund effect). However, since the deficit of observed heterozygosity remained significant in the analysis by genetic stocks (not only by sampling site), other biological causes such as inbreeding may also explain this result considering the reduction in fishery production (Barreto Reyes, 2017Barreto Reyes CG. Producción pesquera de la Cuenca del río Magdalena: desembarcos y estimación ecosistémica. Bogotá: The Nature Conservancy, Mac Arthur Foundation, AUNAP; 2017. Available from: http://sepec.aunap.gov.co/Home/VerPdf/63
http://sepec.aunap.gov.co/Home/VerPdf/63...
), and the overexploitation exerted over the species since 2010 (Gutiérrez et al., 2011Gutiérrez F, Barreto C, Mancilla P. Diagnóstico de la pesquería en la cuenca del Magdalena-Cauca. In: Lasso CA, Gutiérrez F, Morales-Betancourt MA, Agudelo E, Ramírez-Gil H, Ajiaco-Martínez RE, editors. II. Pesquerías continentales de Colombia: cuencas del Magdalena-Cauca, Sinú, Canalete, Atrato, Orinoco, Amazonas y vertiente del Pacífico. Bogotá, D.C: Serie Editorial Recursos Hidrobiológicos y Pesqueros Continentales de Colombia, Instituto de Investigación de los Recursos Biológicos Alexander von Humboldt; 2011. p.35-73.). Besides the fishing pressure, degradation of the Magdalena-Cauca basin caused by environmental, economic and demographic factors, also contribute to the observed decline in the number of catches of this fish species (Jiménez-Segura et al., 2016Jiménez-Segura LF, Galvis-Vergara G, Cala-Cala P, García-Alzate CA, López-Casas S, Ríos-Pulgarín MI et al. Freshwater fish faunas, habitats and conservation challenges in the Caribbean river basins of north-western South America. J Fish Biol. 2016; 89(1):65-101. https://doi.org/10.1111/jfb.13018
https://doi.org/10.1111/jfb.13018...
). Some Neotropical catfishes such as P. grosskopfii (Restrepo-Escobar et al., 2020Restrepo-Escobar, Márquez, Yepes- Acevedo AJ, Márquez EJ. Microsatellite loci development for three catfish species from northwestern South America. Neotrop Ichthyol. 2020; 18(1): e190079.https://doi.org/10.1590/1982-0224-2019-0079
https://doi.org/10.1590/1982-0224-2019-0...
), P. maculatus (Ribolli et al., 2012Ribolli J, Melo CMR, Zaniboni-Filho E. Genetic characterization of the neotropical catfish Pimelodus maculatus (Pimelodidae, Siluriformes) in the Upper Uruguay River. Genet Mol Biol . 2012; 35(4):761-69. doi: https://doi.org/10.1590/S1415-47572012005000060
https://doi.org/10.1590/S1415-4757201200...
), and Pseudoplatystoma reticulatum (Abreu et al., 2009Abreu MM, Pereira LHG, Barretto Vila V, Foresti F, Oliveira C. Genetic variability of two populations of Pseudoplatystoma reticulatum from the Upper Paraguay River Basin. Genet Mol Biol. 2009; 32(4):868-73. https://doi.org/10.1590/S1415-47572009005000075), have also shown evidences of inbreeding. Following Franklin (1980Franklin IR. Evolutionary change in small populations. In: Soulé ME, Wilcox BA, editors. Conservation biology: an evolutionary-ecological perspective. Sunderland: Sinauer Associates; 1980. p.135-49.) and Soulé (1980Soulé ME. Thresholds for survival: Maintaining fitness and evolutionary potential. In: Soulé ME, Wilcox BA, editors. Conservation biology: an evolutionary-ecological perspective . Sunderland: Sinauer Associates ; 1980. p.151-69.), P. yuma requires a rigorous management of the populations since values above 10% indicate possible adverse effects over these populations.
According to the a priori expectation, this study showed evidences that the populations of P. yuma are not genetically structured in the middle and lower sections of the Cauca River, which is in line with the medium-range migration capacity of the species (100-500 km) and the absence of geographical barriers that facilitate the gene flow. This outcome was also found in some congeners like P. grosskopfii from the middle and lower sections of the Cauca River (Restrepo-Escobar et al., in pressRestrepo-Escobar N, Yepes-Acevedo AJ, Márquez EJ. Population genetics of three threatened catfish species in heterogeneous environments of the Cauca River, Colombia. Neotrop Ichthyol . Forthcoming 2021.) and P. maculatus from upper section of the Uruguay River (Ribolli et al., 2012Ribolli J, Melo CMR, Zaniboni-Filho E. Genetic characterization of the neotropical catfish Pimelodus maculatus (Pimelodidae, Siluriformes) in the Upper Uruguay River. Genet Mol Biol . 2012; 35(4):761-69. doi: https://doi.org/10.1590/S1415-47572012005000060
https://doi.org/10.1590/S1415-4757201200...
); although this species also showed genetic structure due to the multiple cascades in the rivers Uruguay and Paranapanema (Almeida et al., 2003Almeida FS, Sodré LMK, Contel EPB. Population structure analysis of Pimelodus maculatus (Pisces, Siluriformes) from the Tietê and Paranapanema Rivers (Brazil). Genet Mol Biol . 2003; 26(3):301-05. https://doi.org/10.1590/S1415-47572003000300014
https://doi.org/10.1590/S1415-4757200300...
; Ramella et al., 2006Ramella MS, Kroth MA, Meurer S, Oliveira AP, Zaniboni E, Maisonnave AC. Genetic variability in four fish species (Pimelodus maculatus, Prochilodus lineatus, Salminus brasiliensis and Steindachneridion scripta) from Uruguay River Basin. Braz Arch Biol Technol. 2006; 49(4):589-98. https://doi.org/10.1590/S1516-89132006000500008
https://doi.org/10.1590/S1516-8913200600...
). In contrast to the study of P. grosskopfii (Restrepo-Escobar et al., in pressRestrepo-Escobar N, Yepes-Acevedo AJ, Márquez EJ. Population genetics of three threatened catfish species in heterogeneous environments of the Cauca River, Colombia. Neotrop Ichthyol . Forthcoming 2021.), this work did not find individuals of P. yuma upstream from one of the slopes of the river, suggesting that this fish species is not capable to overcome this steeped geography.
Although P. yuma did not show genetic differences between the evaluated sections, it comprises two genetic stocks that cohabit. A similar genetic structure was found in its congener P. grosskopffi in the same area (Restrepo-Escobar, Márquez, 2020Restrepo-Escobar, Márquez, Yepes- Acevedo AJ, Márquez EJ. Microsatellite loci development for three catfish species from northwestern South America. Neotrop Ichthyol. 2020; 18(1): e190079.https://doi.org/10.1590/1982-0224-2019-0079
https://doi.org/10.1590/1982-0224-2019-0...
). The existence of those two genetic stocks might be related to spatial or seasonal reproductive isolation. Nevertheless, it remains to explore if the reproductive isolation by season can explain these differences since it was demonstrated that P. yuma (as P. blochii) from a floodplain lake in the medium section of the Magdalena River basin only reproduces during rainfall peaks, despite maintaining a constant condition factor at different environmental conditions (Lopez-Casas, Jimenez-Segura, 2007Lopez Casas S, Jimenez Segura LF. Reproducción y hábitos alimenticios del Nicuro, Pimelodus blochii (Valenciennes, 1840) (Pisces: Pimelodidae), en la ciénaga de cachimbero, Río Magdalena, Colombia. Actual Biol. 2007; 29(87):199-207. Available from: http://www.scielo.org.co/pdf/acbi/v29n87/v29n87a4.pdf
http://www.scielo.org.co/pdf/acbi/v29n87...
). Although our study was not designed to test the hypothesis of temporal genetic structure, the comparisons among available samples in the rain and drought periods per year (S3) did not show any tendency to seasonal or annual genetic structure.
This study led to stablish that the populations of P. yuma from the lower sections of the Cauca River do not exhibit spatial genetic structure, although there are two biological populations along the study area. Likewise, even though this species is affected by different anthropogenic activities, it was found that P. yuma shows a high genetic diversity, but with a deficit in the observed heterozygosity. The first 14 developed polymorphic microsatellite loci for this species are recommended for further studies of genetic diversity on population of P. yuma. Finally, these results are the baseline for future studies aiming to monitor the genetic diversity and structure of the populations of this endemic Colombian fish species in order to generate appropriate management plans.
ACKNOWLEDGMENTS
The authors thank to the Centro Nacional de Secuenciación Genómica, Universidad de Antioquia (Medellín, Colombia) for assistance in bioinformatic analysis and to José David Rangel-Medrano for helping with experiments. This work was supported by Empresas Públicas de Medellín and Universidad Nacional de Colombia - Sede Medellín through two scientific projects: “Variabilidad genotípica y fenotípica poblaciones de especies de peces reófilas presentes en el área de influencia del proyecto hidroeléctrico Ituango - CT-2013-002443-R1” and “Variabilidad genética de un banco de peces de los sectores medio y bajo del río Cauca - CT-2019-000661”.
REFERENCES
- Abreu MM, Pereira LHG, Barretto Vila V, Foresti F, Oliveira C. Genetic variability of two populations of Pseudoplatystoma reticulatum from the Upper Paraguay River Basin. Genet Mol Biol. 2009; 32(4):868-73. https://doi.org/10.1590/S1415-47572009005000075
- Almeida FS, Sodré LMK, Contel EPB. Population structure analysis of Pimelodus maculatus (Pisces, Siluriformes) from the Tietê and Paranapanema Rivers (Brazil). Genet Mol Biol . 2003; 26(3):301-05. https://doi.org/10.1590/S1415-47572003000300014
» https://doi.org/10.1590/S1415-47572003000300014 - Almeida FS, Fungaro MHP, Sodré LMK. RAPD and isoenzyme analysis of genetic variability in three allied species of catfish (Siluriformes: Pimelodidae) from the Tibagi River, Brazil. J Zool. 2001; 253(1):113-20. https://doi.org/10.1017/S0952836901000103
» https://doi.org/10.1017/S0952836901000103 - Barbará T, Palma-Silva C, Paggi GM, Bered F, Fay MF, Lexer C. Cross-species transfer of nuclear microsatellite markers: Potential and limitations. Mol Ecol. 2007; 16(18):3759-67. https://doi.org/10.1111/j.1365-294X.2007.03439.x
» https://doi.org/10.1111/j.1365-294X.2007.03439.x - Barreto Reyes CG. Producción pesquera de la Cuenca del río Magdalena: desembarcos y estimación ecosistémica. Bogotá: The Nature Conservancy, Mac Arthur Foundation, AUNAP; 2017. Available from: http://sepec.aunap.gov.co/Home/VerPdf/63
» http://sepec.aunap.gov.co/Home/VerPdf/63 - Batista JS, Farias IP, Formiga-Aquino K, Sousa ACB, Alves-Gomes JA. DNA microsatellite markers for “dourada” (Brachyplatystoma rousseauxii, Siluriformes: Pimelodidae), a migratory catfish of utmost importance for fisheries in the Amazon: development, characterization and inter-specific amplification. Conserv Genet Resour. 2010; 2(1):5-10. https://doi.org/10.1007/s12686-009-9117-5
» https://doi.org/10.1007/s12686-009-9117-5 - Blacket MJ, Robin C, Good RT, Lee SF, Miller AD. Universal primers for fluorescent labelling of PCR fragments - an efficient and cost-effective approach to genotyping by fluorescence. Mol Ecol Resour. 2012; 12(3):456-63. https://doi.org/10.1111/j.1755-0998.2011.03104.x
» https://doi.org/10.1111/j.1755-0998.2011.03104.x - Botstein D, White RL, Skolnick M, Davis RW. Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet. 1980; 32(3):314-31. Available from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1686077/
» https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1686077/ - Carvajal-Vallejos FM, Duponchelle F, Desmarais E, Cerqueira F, Querouil S, Nuñez J et al Genetic structure in the Amazonian catfish Brachyplatystoma rousseauxii: Influence of life history strategies. Genetica. 2014; 142(4):323-36. https://doi.org/10.1007/s10709-014-9777-2
» https://doi.org/10.1007/s10709-014-9777-2 - Carvalho DC, Beheregaray LB. Rapid development of microsatellites for the endangered Neotropical catfish Conorhynchus conirostris using a modest amount of 454 shot-gun pyrosequencing. Conserv Genet Resour . 2011; 3(2):373-75. https://doi.org/10.1007/s12686-010-9365-4
» https://doi.org/10.1007/s12686-010-9365-4 - Carvalho DC, Oliveira DAA, Beheregaray LB, Torres RA. Hidden genetic diversity and distinct evolutionarily significant units in a commercially important Neotropical apex predator, the catfish Pseudoplatystoma corruscans Conserv Genet. 2012; 13(6):1671-75. https://doi.org/10.1007/s10592-012-0402-6
» https://doi.org/10.1007/s10592-012-0402-6 - Castoe TA, Poole AW, Gu W, Jason de Koning AP, Daza JM, Smith EN, Pollock DD. Rapid identification of thousands of copperhead snake (Agkistrodon contortrix) microsatellite loci from modest amounts of 454 shotgun genome sequence. Mol Ecol Resour. 2010; 10(2):341-47. https://doi.org/10.1111/j.1755-0998.2009.02750.x
» https://doi.org/10.1111/j.1755-0998.2009.02750.x - Chistiakov DA, Hellemans B, Volckaert F. Microsatellites and their genomic distribution, evolution, function and applications: A review with special reference to fish genetics. Aquaculture. 2006; 255(1-4):1-29. https://doi.org/10.1016/j.aquaculture.2005.11.031
» https://doi.org/10.1016/j.aquaculture.2005.11.031 - Dantas HL, dos Santos MA, Oliveira KKC, Severi W, Diniz FM, Coimbra MR. Genetic diversity of captive and wild threatened Catfish Pseudoplatystoma corruscans in the São Francisco River. Rev Fish Sci. 2013; 21(3-4):237-46. https://doi.org/10.1080/10641262.2013.800787
» https://doi.org/10.1080/10641262.2013.800787 - Estoup A, Jarne P, Cornuet JM. Homoplasy and mutation model at microsatellite loci and their consequences for population genetics analysis. Mol Ecol . 2002; 11(9):1591-604. https://doi.org/10.1046/j.1365-294X.2002.01576.x
» https://doi.org/10.1046/j.1365-294X.2002.01576.x - Excoffier L, Lischer HEL. Arlequin suite ver 3.5: A new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour. 2010; 10(3):564-67. https://doi.org/10.1111/j.1755-0998.2010.02847.x
» https://doi.org/10.1111/j.1755-0998.2010.02847.x - Falush D, Matthew S, Pritchard KJ. Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics. 2003; 164(4): 1567-87. Available from: https://www.genetics.org/content/164/4/1567
» https://www.genetics.org/content/164/4/1567 - Falush D, Stephens M, Pritchard JK. Inference of population structure using multilocus genotype data: Dominant markers and null alleles. Mol Ecol Notes. 2007; 7(4):574-78. https://doi.org/10.1111/j.1471-8286.2007.01758.x
» https://doi.org/10.1111/j.1471-8286.2007.01758.x - Fonseca FS, Domingues RR, Hallerman EM, Hilsdorf AWS. Genetic diversity of an imperiled Neotropical catfish and recommendations for its restoration. Front Genet. 2017; 8:196. https://doi.org/10.3389/fgene.2017.00196
» https://doi.org/10.3389/fgene.2017.00196 - Franklin IR. Evolutionary change in small populations. In: Soulé ME, Wilcox BA, editors. Conservation biology: an evolutionary-ecological perspective. Sunderland: Sinauer Associates; 1980. p.135-49.
- Freeland JR. Molecular Ecology. England: John Wiley & Sons; 2020.
- Fricke R, Eschmeyer WN, Van der Laan R. Eschmeyer’s catalog of fishes: genera, species, references [Internet]. San Francisco: California Academy of Science; 2020. Available from: http://researcharchive.calacademy.org/research/ichthyology/catalog/fishcatmain.asp
» http://researcharchive.calacademy.org/research/ichthyology/catalog/fishcatmain.asp - Gutiérrez F, Barreto C, Mancilla P. Diagnóstico de la pesquería en la cuenca del Magdalena-Cauca. In: Lasso CA, Gutiérrez F, Morales-Betancourt MA, Agudelo E, Ramírez-Gil H, Ajiaco-Martínez RE, editors. II. Pesquerías continentales de Colombia: cuencas del Magdalena-Cauca, Sinú, Canalete, Atrato, Orinoco, Amazonas y vertiente del Pacífico. Bogotá, D.C: Serie Editorial Recursos Hidrobiológicos y Pesqueros Continentales de Colombia, Instituto de Investigación de los Recursos Biológicos Alexander von Humboldt; 2011. p.35-73.
- Hilsdorf AWS, Hallerman EM. Genetic resources of Neotropical fishes. Cham: Springer International Publishing AG; 2017.
- Holm S. A simple sequentially rejective multiple test procedure. Scand Stat Theory Appl. 1979; 6(2): 65-70. Available from: https://www.jstor.org/stable/4615733
» https://www.jstor.org/stable/4615733 - Hubisz MJ, Falush D, Stephens M, Pritchard JK. Inferring weak population structure with the assistance of sample group information. Mol Ecol Resour. 2009; 9(5):1322-32. https://doi.org/10.1111/j.1755-0998.2009.02591.x
» https://doi.org/10.1111/j.1755-0998.2009.02591.x - Jakobsson M, Rosenberg NA. CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics. 2007; 23(14):1801-06. https://doi.org/10.1093/bioinformatics/btm233
» https://doi.org/10.1093/bioinformatics/btm233 - Jiménez-Segura LF, Galvis-Vergara G, Cala-Cala P, García-Alzate CA, López-Casas S, Ríos-Pulgarín MI et al Freshwater fish faunas, habitats and conservation challenges in the Caribbean river basins of north-western South America. J Fish Biol. 2016; 89(1):65-101. https://doi.org/10.1111/jfb.13018
» https://doi.org/10.1111/jfb.13018 - Jombart T. Adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics. 2008; 24(11):1403-05. https://doi.org/10.1093/bioinformatics/btn129
» https://doi.org/10.1093/bioinformatics/btn129 - Kopelman NM, Mayzel J, Jakobsson M, Rosenberg NA, Mayrose I. Clumpak: a program for identifying clustering modes and packaging population structure inferences across K. Mol Ecol Resour. 2015; 15(5):1179-91. https://doi.org/10.1111/1755-0998.12387
» https://doi.org/10.1111/1755-0998.12387 - Landínez-García RM, Márquez EJ. Development and characterization of 24 polymorphic microsatellite loci for the freshwater fish Ichthyoelephas longirostris (Characiformes: Prochilodontidae). PeerJ. 2016; 4:e2419. https://doi.org/10.7717/peerj.2419
» https://doi.org/10.7717/peerj.2419 - Landínez-García RM, Marquez EJ. Microsatellite loci development and population genetics in Neotropical fish Curimata mivartii (Characiformes: Curimatidae). PeerJ. 2018; 6:e5959. https://doi.org/10.7717/peerj.5959
» https://doi.org/10.7717/peerj.5959 - Li L, Lin H, Tang W, Liu D, Bao B, Yang J. Population genetic structure in wild and aquaculture populations of Hemibarbus maculates inferred from microsatellites markers. Aquac Fish. 2017; 2(2):78-83. https://doi.org/10.1016/j.aaf.2017.03.004
» https://doi.org/10.1016/j.aaf.2017.03.004 - Li YL, Liu JX. StructureSelector: A web-based software to select and visualize the optimal number of clusters using multiple methods. Mol Ecol Resour. 2018; 18(1):176-77. https://doi.org/10.1111/1755-0998.12719
» https://doi.org/10.1111/1755-0998.12719 - Lopez Casas S, Jimenez Segura LF. Reproducción y hábitos alimenticios del Nicuro, Pimelodus blochii (Valenciennes, 1840) (Pisces: Pimelodidae), en la ciénaga de cachimbero, Río Magdalena, Colombia. Actual Biol. 2007; 29(87):199-207. Available from: http://www.scielo.org.co/pdf/acbi/v29n87/v29n87a4.pdf
» http://www.scielo.org.co/pdf/acbi/v29n87/v29n87a4.pdf - Marshall TC, Slate J, Kruuk LEB, Pemberton JM. Statistical confidence for likelihood-based paternity inference in natural populations. Mol Ecol . 1998; 7(5):639-55. https://doi.org/10.1046/j.1365-294x.1998.00374.x
» https://doi.org/10.1046/j.1365-294x.1998.00374.x - Meirmans PG. Using the AMOVA framework to estimate a tandardized genetic differentiation measure. Evolution. 2006; 60(11):2399-2402. https://doi.org/10.1554/05-631.1
- Meirmans PG, Hedrick PW. Assessing population structure: FST and related measures. Mol Ecol Resour. 2011; 11(1):5-18. https://doi.org/10.1111/j.1755-0998.2010.02927.x
» https://doi.org/10.1111/j.1755-0998.2010.02927.x - Mojica JI, Usma-Oviedo JS, Álvarez-León R, Lasso CA, editors. Libro Rojo de peces dulceacuicolas de Colombia (2012). Bogotá, D.C. Colombia: Instituto de Investigación de los Recursos Biológicos Alexander von Humboldt, Instituto de Ciencias Naturales de la Universidad Nacional de Colombia, WWF Colombia, Universidad de Manizales; 2012.
- Paiva ALB, Kalapothakis E. Isolation and characterization of microsatellite loci in Pimelodus maculatus (Siluriformes: Pimelodidae). Mol Ecol Resour. 2008; 8(5):1078-80. https://doi.org/10.1111/j.1755-0998.2008.02160.x
» https://doi.org/10.1111/j.1755-0998.2008.02160.x - Peakall R, Smouse PE. GenALEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics. 2012; 28(19):2537-39. https://doi.org/10.1093/bioinformatics/bts460
» https://doi.org/10.1093/bioinformatics/bts460 - Pereira LHG, Foresti F, Oliveira C. Genetic structure of the migratory catfish Pseudoplatystoma corruscans (Siluriformes: Pimelodidae) suggests homing behaviour. Ecol Freshw Fish. 2009; 18(2):215-25. https://doi.org/10.1111/j.1600-0633.2008.00338.x
» https://doi.org/10.1111/j.1600-0633.2008.00338.x - Prado FD, Fernandez-Cebrián R, Foresti F, Oliveira C, Martínez P, Porto-Foresti F. Genetic structure and evidence of anthropogenic effects on wild populations of two neotropical catfishes: Baselines for conservation. J Fish Biol . 2017; 92(1):55-72. https://doi.org/10.1111/jfb.13486
» https://doi.org/10.1111/jfb.13486 - Prado FD, Pardo BG, Guerra-Varela J, Senhorini JA, Martínez P, Foresti F, Porto-Foresti F. Development and characterization of 16 microsatellites for the Neotropical catfish Pseudoplatystoma reticulatum and cross species analysis. Conserv Genet Resour . 2014; 6(3):679-81. https://doi.org/10.1007/s12686-014-0180-1
» https://doi.org/10.1007/s12686-014-0180-1 - Pritchard JK, Stephens M, Donnelly P. Inference of population structure using multilocus genotype data. Genetics 2000; 155(2):945-59. https://doi.org/10.1111/j.1471-8286.2007.01758.x
» https://doi.org/10.1111/j.1471-8286.2007.01758.x - Puechmaille SJ. The program STRUCTURE does not reliably recover the correct population structure when sampling is uneven: Subsampling and new estimators alleviate the problem. Mol Ecol Resour. 2016; 16(3):608-27. https://doi.org/10.1111/1755-0998.12512
» https://doi.org/10.1111/1755-0998.12512 - Raj A, Stephens M, Pritchard JK. fastSTRUCTURE: Variational Inference of Population Structure in Large SNP Data Sets. Genetics. 2014; 197(2):573-89. https://doi.org/10.1534/genetics.114.164350
» https://doi.org/10.1534/genetics.114.164350 - Ramella MS, Kroth MA, Meurer S, Oliveira AP, Zaniboni E, Maisonnave AC. Genetic variability in four fish species (Pimelodus maculatus, Prochilodus lineatus, Salminus brasiliensis and Steindachneridion scripta) from Uruguay River Basin. Braz Arch Biol Technol. 2006; 49(4):589-98. https://doi.org/10.1590/S1516-89132006000500008
» https://doi.org/10.1590/S1516-89132006000500008 - Restrepo-Escobar, Márquez, Yepes- Acevedo AJ, Márquez EJ. Microsatellite loci development for three catfish species from northwestern South America. Neotrop Ichthyol. 2020; 18(1): e190079.https://doi.org/10.1590/1982-0224-2019-0079
» https://doi.org/10.1590/1982-0224-2019-0079 - Restrepo-Escobar N, Yepes-Acevedo AJ, Márquez EJ. Population genetics of three threatened catfish species in heterogeneous environments of the Cauca River, Colombia. Neotrop Ichthyol . Forthcoming 2021.
- Ribolli J, Melo CMR, Zaniboni-Filho E. Genetic characterization of the neotropical catfish Pimelodus maculatus (Pimelodidae, Siluriformes) in the Upper Uruguay River. Genet Mol Biol . 2012; 35(4):761-69. doi: https://doi.org/10.1590/S1415-47572012005000060
» https://doi.org/10.1590/S1415-47572012005000060 - Rice W. Analyzing tables of statistical tests. Evolution. 1989; 43(1):223-25. http://dx.doi.org/10.1111/j.1558-5646.1989.tb04220.x
» http://dx.doi.org/10.1111/j.1558-5646.1989.tb04220.x - Rozen S, Skaletsky H. Primer3 on the World Wide Web for general users and for biologist programmers. In: Misener S, Krawetz S, editors. Bioinformatics methods and protocols. New Jersey: Humana Press; 2000. p.365-386
- Rutkowski R, Sielezniew M, Szostak A. Contrasting levels of polymorphism in cross-amplified microsatellites in two endangered xerothermophilous, obligatorily myrmecophilous, butterflies of the genus Phengaris (Maculinea) (Lepidoptera: Lycaenidae). Eur J Entomol. 2009; 106(4):457-69. https://doi.org/10.14411/eje.2009.058
» https://doi.org/10.14411/eje.2009.058 - Saulo-Machado AC, Formiga KM, Ortiz MF, Sousa ACB, Alves-Gomes JA, Batista JS. Polymorphic microsatellite DNA markers for the Amazonian catfish Pseudoplatystoma punctifer (Siluriformes: Pimelodidae). Conserv Genet Resour . 2011; 3(2):307-10. https://doi.org/10.1007/s12686-010-9349-4
» https://doi.org/10.1007/s12686-010-9349-4 - Savada CS, Lima MCC, Orsi ML, Almeida FS. Isolation and characterization of microsatellite loci in Pimelodus microstoma (Siluriformes: Pimelodidae). J Appl Ichthyol. 2020; 36:825-29. https://doi.org/10.1111/jai.14104
» https://doi.org/10.1111/jai.14104 - Sekine ES, Prioli AJ, Prioli SMAP, Júlio Júnior HF. Genetic differentiation among populations of Pseudoplatystoma corruscans (Agassiz, 1829) (Osteichthyes, Pimelodidae) isolated by the Guaíra Falls in the Paraná River. Acta Sci Nol Sci. 2002; 24(2):507-12. Available from: http://repositorio.uem.br:8080/jspui/bitstream/1/5222/1/375.pdf
» http://repositorio.uem.br:8080/jspui/bitstream/1/5222/1/375.pdf - Soulé ME. Thresholds for survival: Maintaining fitness and evolutionary potential. In: Soulé ME, Wilcox BA, editors. Conservation biology: an evolutionary-ecological perspective . Sunderland: Sinauer Associates ; 1980. p.151-69.
- Souza CA, Hashimoto DT, Pereira LHG, Oliveira C, Foresti F, Porto-Foresti F. Development and characterization of microsatellite loci in Phractocephalus hemioliopterus (Siluriformes: Pimelodidae) and their cross-species amplification in six related species. Conserv Genet Resour . 2012; 4(2):499-501. https://doi.org/10.1007/s12686-011-9584-3
» https://doi.org/10.1007/s12686-011-9584-3 - Telles MPC, Collevatti RG, Braga RS, Guedes LBS, Castro TG, Costa MC, Silva Júnior NJ, Barthem RB, Diniz Filho JAF. Geographical genetics of Pseudoplatystoma punctifer (Castelnau, 1855) (Siluriformes, Pimelodidae) in the Amazon Basin. Genet Mol Res. 2014; 13(2):3656-66. https://doi.org/10.4238/2014.may.9.8
» https://doi.org/10.4238/2014.may.9.8 - Tognelli MF, Lasso CA, Bota-Sierra CA, Jiménez-segura LF, Cox NA, editors. Estado de conservación y distribución de la biodiversidad de agua dulce en los Andes tropicales. Gland, Suiza, Cambridge, UK y Arlington, USA: UICN; 2016.
- Triantafyllidis A, Krieg F, Cottin C, Abatzopoulos TJ, Triantaphyllidis C, Guyomard R. Genetic structure and phylogeography of European catfish (Silurus glanis) populations. Mol Ecol . 2002; 11(6):1039-55. https://doi.org/10.1046/j.1365-294X.2002.01501.x
» https://doi.org/10.1046/j.1365-294X.2002.01501.x - Usma Oviedo JS, Villa-Navarro F, Lasso CA, Castro F, Zúñiga-Upegui PT, Cipamocha CA, Ortega-Lara A, Ajiaco RE, Ramírez-Gil H, Jiménez LF, Maldonado-Ocampo J, Muñoz JA, Suárez JT. Peces dulceacuícolas Migratorios de Colombia. In: Zapata LA, Usma JS, editors. Guía de las Especies Migratorias de la Biodiversidad en Colombia. Peces. Vol. 2. Bogotá: Ministerio de Ambiente y Desarrollo Sostenible, WWF-Colombia; 2013. p.215-441.
- Vaini JO, Crispim BA, Santos Silva DB, Benites C, Russo MR, Grisolia AB. Genetic variability of pure Pseudoplatystoma corruscans and Pseudoplatystoma reticulatum individuals in the Paraná and Paraguay River basins. Fish Sci. 2016; 82(4):605-11. https://doi.org/10.1007/s12562-016-0999-3
» https://doi.org/10.1007/s12562-016-0999-3 - Van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P. MICRO-CHECKER: Software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes. 2004; 4(3):535-38. https://doi.org/10.1111/j.1471-8286.2004.00684.x
- Villa-Navarro FA, Acero A, Cala Cala P. Taxonomic review of Trans-Andean species of Pimelodus (Siluriformes: Pimelodidae), with the descriptions of two new species. Zootaxa. 2017; 4299(3):337-60. https://doi.org/10.11646/zootaxa.4299.3.2
» https://doi.org/10.11646/zootaxa.4299.3.2 - Yue GH, Kovacs B, Orban L. A new problem with cross-species amplification of microsatellites: Generation of non-homologous products. Dongwuxue Yanjiu. 2010; 31(2):131-40. doi: https://doi.org/10.3724/SP.J.1141.2010.02131
» https://doi.org/10.3724/SP.J.1141.2010.02131
ADDITIONAL NOTES
-
HOW TO CITE THIS ARTICLE
Joya CD, Landínez-García RM, Márquez EJ. Development of microsatellite loci and population genetics of the catfish Pimelodus yuma (Siluriformes: Pimelodidae). Neotrop Ichthyol. 2021; 19(1):e200114. https://doi.org/10.1590/1982-0224-2020-0114
Edited by
Data availability
Data citations
Fricke R, Eschmeyer WN, Van der Laan R. Eschmeyer’s catalog of fishes: genera, species, references [Internet]. San Francisco: California Academy of Science; 2020. Available from: http://researcharchive.calacademy.org/research/ichthyology/catalog/fishcatmain.asp
Publication Dates
-
Publication in this collection
08 Mar 2021 -
Date of issue
2021
History
-
Received
10 Oct 2020 -
Accepted
16 Dec 2020